Abstract
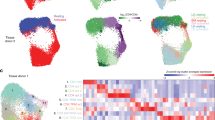
The somatic translocation t(8;21)(q22;q22)/RUNX1-RUNX1T1 is one of the most frequent rearrangements found in children with standard-risk acute myeloid leukemia (AML). Despite the favorable prognostic role of this aberration, we recently observed a higher than expected frequency of relapse. Here, we employed an integrated high-throughput approach aimed at identifying new biological features predicting relapse among 34 t(8;21)-rearranged patients. We found that the DNA methylation status of patients who suffered from relapse was peculiarly different from that of children maintaining complete remission. The epigenetic signature, made up of 337 differentially methylated regions, was then integrated with gene and protein expression profiles, leading to a network, where cell-to-cell adhesion and cell-motility pathways were found to be aberrantly activated in relapsed patients. We identified most of these factors as RUNX1-RUNX1T1 targets, with Ras Homolog Family Member (RHOB) overexpression being the core of this network. We documented how RHOB re-organized the actin cytoskeleton through its downstream ROCK–LIMK–COFILIN axis: this increases blast adhesion by stress fiber formation, and reduces mitochondrial apoptotic cell death after chemotherapy treatment. Altogether, our data show an epigenetic heterogeneity within t(8;21)-rearranged AML patients at diagnosis able to influence the program of the chimeric transcript, promoting blast re-emergence and progression to relapse.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Rubnitz JE, Inaba H, Dahl G, Ribeiro RC, Bowman WP, Taub J, et al. Minimal residual disease-directed therapy for childhood acute myeloid leukaemia: results of the AML02 multicentre trial. Lancet Oncol. 2010;11:543–52.
Abrahamsson J, Forestier E, Heldrup J, Jahnukainen K, Jónsson OG, Lausen B, et al. Response-guided induction therapy in pediatric acute myeloid leukemia with excellent remission rate. J Clin Oncol. 2011;29:310–5.
Gibson BES, Webb DKH, Howman AJ, De Graaf SSN, Harrison CJ, Wheatley K. Results of a randomized trial in children with acute myeloid leukaemia: medical research council AML12 trial. Br J Haematol. 2011;155:366–76.
Creutzig U, Zimmermann M, Bourquin J-P, Dworzak MN, Fleischhack G, Graf N, et al. Randomized trial comparing liposomal daunorubicin with idarubicin as induction for pediatric acute myeloid leukemia: results from study AML-BFM 2004. Blood. 2013;122:37–43.
Pession A, Masetti R, Rizzari C, Putti MC, Casale F, Fagioli F, et al. Results of the AIEOP AML 2002/01 multicenter prospective trial for the treatment of children with acute myeloid leukemia. Blood. 2013;122:170–8.
Zwaan CM, Kolb EA, Reinhardt D, Abrahamsson J, Adachi S, Aplenc R, et al. Collaborative efforts driving progress in pediatric acute myeloid leukemia. J Clin Oncol. 2015;33:2949–62.
De Rooij JDE, Zwaan CM, van den Heuvel-Eibrink M. Pediatric AML: from biology to clinical management. J Clin Med. 2015;4:127–49.
Cheson BD. Revised recommendations of the international working group for diagnosis, standardization of response criteria, treatment outcomes, and reporting standards for therapeutic trials in acute myeloid leukemia. J Clin Oncol. 2003;21:4642–9.
Rubnitz JE, Pounds S, Cao X, Jenkins L, Dahl G, Bowman WP, et al. Treatment outcome in older patients with childhood acute myeloid leukemia. Cancer. 2012;118:6253–9.
Nguyen S, Leblanc T, Fenaux P, Witz F, Blaise D, Pigneux A, et al. A white blood cell index as the main prognostic factor in t(8;21) acute myeloid leukemia (AML): a survey of 161 cases from the French AML intergroup. Blood. 2002;99:3517–23.
Chang M, Raimondi SC, Ravindranath Y, Carroll AJ, Camitta B, Gresik MV, et al. Prognostic factors in children and adolescents with acute myeloid leukemia (excluding children with Down syndrome and acute promyelocytic leukemia): univariate and recursive partitioning analysis of patients treated on Pediatric Oncology Group (POG) study. Leukemia. 2000;14:1201–7.
Witte T, Plass C, Gerhauser C. Pan-cancer patterns of DNA methylation. Genome Med. 2014;6:66.
Weisenberger DJ. Characterizing DNA methylation alterations from The Cancer Genome Atlas. J Clin Invest. 2014;124:17–23.
Huether R, Dong L, Chen X, Wu G, Parker M, Wei L, et al. The landscape of somatic mutations in epigenetic regulators across 1,000 paediatric cancer genomes. Nat Commun. 2014;5:619–22.
Paganin M, Pigazzi M, Bresolin S, Masetti R, Fagioli F, Chiaretti S, et al. DNA methyltransferase 3a hot-spot locus is not mutated in pediatric patients affected by acute myeloid or T-cell acute lymphoblastic leukemia: an Italian study. Haematologica. 2011;96:1886–7.
Pigazzi M, Ferrari G, Masetti R, Falini B, Martinolli F, Basso G, et al. Low prevalence of IDH1 gene mutation in childhood AML in Italy. Leukemia. 2011;25:173–4.
Accordi B, Espina V, Giordan M, VanMeter A, Milani G, Galla L, et al. Functional protein network activation mapping reveals new potential molecular drug targets for poor prognosis pediatric BCP-ALL. PLoS ONE. 2010;5:e13552.
Goeman JJ, van de Geer SA, de Kort F, van Houwelingen HC. A global test for groups of genes: testing association with a clinical outcome. Bioinformatics. 2003;20:93–9.
Daskalos A, Nikolaidis G, Xinarianos G, Savvari P, Cassidy A, Zakopoulou R, et al. Hypomethylation of retrotransposable elements correlates with genomic instability in non-small cell lung cancer. Int J Cancer. 2009;124:81–7.
Lee E, Iskow R, Yang L, Gokcumen O, Haseley P, Luquette LJ, et al. Landscape of somatic retrotransposition in human cancers. Science. 2012;337:967–71.
Feinberg AP, Tycko B. The history of cancer epigenetics. Nat Rev Cancer. 2004;4:143–53.
Choi SH, Worswick S, Byun H-M, Shear T, Soussa JC, Wolff EM, et al. Changes in DNA methylation of tandem DNA repeats are different from interspersed repeats in cancer. Int J Cancer. 2009;125:723–9.
Hur K, Cejas P, Feliu J, Moreno-Rubio J, Burgos E, Boland CR, et al. Hypomethylation of long interspersed nuclear element-1 (LINE-1) leads to activation of proto-oncogenes in human colorectal cancer metastasis. Gut. 2013;63:635–46.
Saied MH, Marzec J, Khalid S, Smith P, Down TA, Rakyan VK, et al. Genome wide analysis of acute myeloid leukemia reveal leukemia specific methylome and subtype specific hypomethylation of repeats. PLoS ONE. 2012;7:e33213.
Phillips-Cremins JE, Sauria MEG, Sanyal A, Gerasimova TI, Lajoie BR, Bell JSK, et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell. 2013;153:1281–95.
Symmons O, Uslu VV, Tsujimura T, Ruf S, Nassari S, Schwarzer W, et al. Functional and topological characteristics of mammalian regulatory domains. Genome Res. 2014;24:390–400.
Bell AC, Felsenfeld G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature. 2000;405:482–5.
Zhu H-H, Zhang X-H, Qin Y-Z, Liu D-H, Jiang H, Chen H, et al. MRD-directed risk stratification treatment may improve outcomes of t(8;21) AML in the first complete remission: results from the AML05 multicenter trial. Blood. 2013;121:4056–62.
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. STRINGv10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015;43:D447–52.
Bachas C, Schuurhuis GJ, Zwaan CM, van den Heuvel-Eibrink MM, den Boer ML, de Bont ESJM, et al. Gene expression profiles associated with pediatric relapsed AML. PLoS ONE. 2015;10:e0121730.
Ptasinska A, Assi SA, Mannari D, James SR, Williamson D, Dunne J, et al. Depletion of RUNX1/ETO in t(8;21) AML cells leads to genome-wide changes in chromatin structure and transcription factor binding. Leukemia. 2012;26:1829–41.
Figueroa ME, Lugthart S, Li Y, Erpelinck-Verschueren C, Deng X, Christos PJ, et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell. 2010;17:13–27.
Akalin A, Garrett-Bakelman FE, Kormaksson M, Busuttil J, Zhang L, Khrebtukova I, et al. Base-pair resolution DNA methylation sequencing reveals profoundly divergent epigenetic landscapes in acute myeloid leukemia. PLoS Genet. 2012;8:e1002781.
Corces-Zimmerman MR, Hong W-J, Weissman IL, Medeiros BC, Majeti R. Preleukemic mutations in human acute myeloid leukemia affect epigenetic regulators and persist in remission. Proc Natl Acad Sci USA. 2014;111:2548–53.
Cole CB, Verdoni AM, Ketkar S, Leight ER, Russler-Germain DA, Lamprecht TL, et al. PML-RARA requires DNA methyltransferase 3A to initiate acute promyelocytic leukemia. J Clin Invest. 2015;126:85–98.
Kanduri C, Pant V, Loukinov D, Pugacheva E, Qi CF, Wolffe A, et al. Functional association of CTCF with the insulator upstream of the H19 gene is parent of origin-specific and methylation-sensitive. Curr Biol. 2000;10:853–6.
Wang H, Maurano MT, Qu H, Varley KE, Gertz J, Pauli F, et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 2012;22:1680–8.
Maurano MT, Wang H, John S, Shafer A, Canfield T, Lee K, et al. Role of DNA methylation in modulating transcription factor occupancy. Cell Rep. 2015;12:1184–95.
Schoofs T, Rohde C, Hebestreit K, Klein H-U, Göllner S, Schulze I, et al. DNA methylation changes are a late event in acute promyelocytic leukemia and coincide with loss of transcription factor binding. Blood. 2013;121:178–87.
Stadler MB, Murr R, Burger L, Ivanek R, Lienert F, Schöler A, et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature. 2011;480:490–5. https://doi.org/10.1038/nature10716.
Thurman RE, Rynes E, Humbert R, Vierstra J, Maurano MT, Haugen E, et al. The accessible chromatin landscape of the human genome. Nature. 2012;489:75–82.
Hazlehurst LA. Reduction in drug-induced DNA double-strand breaks associated with beta1 integrin-mediated adhesion correlates with drug resistance in U937 cells. Blood. 2001;98:1897–903.
Matsunaga T, Takemoto N, Sato T, Takimoto R, Tanaka I, Fujimi A, et al. Interaction between leukemic-cell VLA-4 and stromal fibronectin is a decisive factor for minimal residual disease of acute myelogenous leukemia. Nat Med. 2003;9:1158–65.
Li Z, Li L. Understanding hematopoietic stem-cell microenvironments. Trends Biochem Sci. 2006;31:589–95.
Saia M, Termanini A, Rizzi N, Mazza M, Barbieri E, Valli D, et al. AML1/ETO accelerates cell migration and impairs cell-to-cell adhesion and homing of hematopoietic stem/progenitor cells. Sci Rep. 2016;6:34957.
Hall A. Rho family GTPases. Biochem Soc Trans. 2012;40:1378–82.
Vega FM, Colomba A, Reymond N, Thomas M, Ridley AJ. RhoB regulates cell migration through altered focal adhesion dynamics. Open Biol. 2012;2:120076.
Chua BT, Volbracht C, Tan KO, Li R, Yu VC, Li P. Mitochondrial translocation of cofilin is an early step in apoptosis induction. Nat Cell Biol. 2003;5:1083–9.
Acknowledgements
This work was supported by grants from CARIPARO Istituto di Ricerca Pediatrica-Fondazione Città della Speranza and Università degli Studi di Padova and from AIRC (Associazione Italiana Ricerca sul Cancro, Special Grant “5xmille”−9962 to FL). We thank Dr. Federica Cattonaro and Dr. Davide Scaglione (IGA Udine, Italy) for RRBS service. We thank Katia Polato, Dr. Anna Leslz, Dr. AnnaMaria Di Meglio, and Dr. Emanuela Giarin for molecular genetic analysis, cytogenetic analysis, and for storing samples of children enrolled in the AML 2002/01 protocol at Hematology-Oncology Clinic at UniPD. Our thanks also to all clinicians working at the participating AIEOP centers.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Matteo Zampini and Claudia Tregnago contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zampini, M., Tregnago, C., Bisio, V. et al. Epigenetic heterogeneity affects the risk of relapse in children with t(8;21)RUNX1-RUNX1T1-rearranged AML. Leukemia 32, 1124–1134 (2018). https://doi.org/10.1038/s41375-017-0003-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-017-0003-y
This article is cited by
-
Tumor-suppressing effects of miR-381-3p in pediatric acute myeloid leukemia via ROCK1 downregulation
Functional & Integrative Genomics (2023)
-
Requirement for LIM kinases in acute myeloid leukemia
Leukemia (2020)