Abstract
In patients with breast cancer, primary chemotherapy often fails due to survival of chemoresistant breast cancer stem cells (BCSCs) which results in recurrence and metastasis of the tumor. However, the factors determining the chemoresistance of BCSCs have remained to be investigated. Here, we profiled a series of differentially expressed microRNAs (miRNAs) between parental adherent breast cancer cells and BCSC-mimicking mammosphere-derived cancer cells, and identified hsa-miR-27a as a negative regulator for survival and chemoresistance of BCSCs. In the mammosphere, we found that the expression of hsa-miR-27a was downregulated, and ectopic overexpression of hsa-miR-27a reduced both number and size of mammospheres. In addition, overexpression of hsa-miR-27a sensitized breast cancer cells to anticancer drugs by downregulation of genes essential for detoxification of reactive oxygen species (ROS) and impairment of autophagy. Therefore, enhancing the hsa-miR-27a signaling pathway can be a potential therapeutic modality for breast cancer.
Similar content being viewed by others
Introduction
Breast cancer is a leading cause of cancer-associated deaths for women world-wide. Although recent improvements of regimens for the conventional chemotherapy for breast cancer gradually reduce the rate of relapse, a significant proportion of patients suffer from unexpected relapse and metastasis even after years or decades of periods with no detectable tumors [1, 2]. Current hypotheses suggest that these recurrence and metastases arise from residual disseminated tumor cells, which disseminate from primary lesions, undergoing an extended period of dormancy in the stroma of metastasized organs [3,4,5,6,7,8,9]. Importantly, these dormant tumor cells seem to have a special drug resistance machinery which protects them from chemotherapy, and accumulating evidences suggest that the dormant disseminated breast cancer cells should be equated with breast cancer stem cells (BCSCs), characterized by self-renewal and differentiation into multiple cell types of breast cancer [10].
The molecular mechanism by which BCSCs tolerate cytotoxic chemotherapy still remains enigmatic; however, their ability to maintain intracellular reactive oxygen species (ROS) levels lower than their matured progeny cells may be implicated in the survival [11]. Intracellular ROS plays pivotal roles in cellular signaling and homeostasis including inflammation, apoptosis, DNA damage, and posttranslational modification thereby its level is strictly regulated by a complexed antioxidative defense system including the Keap1-Nrf2 pathway [12,13,14]. Under normal condition, ROS is produced as a byproduct of biochemical reactions, such as oxidative phosphorylation at mitochondria and oxidative folding of proteins in endoplasmic reticulum. Once excessive amount of ROS is produced, cytosolic Keap1 is inactivated by oxidation of its cysteine residues and releases a transcription factor Nrf2. After translocation into the nucleus, Nrf2 induces expression of a series of antioxidant genes including glutathione (GSH) reductase [15,16,17] which produces endogenous antioxidant, reduced GSH.
Another possible mechanism explaining the special chemoresistance of BCSCs is higher level of basal autophagic flux [3, 18, 19]. Autophagy is a cellular process which degrades organelles and proteins in response to various stimuli, such as nutrient starvation, and autophagy augments supply of amino acids to BCSCs which survive for years in dormancy. Importantly, not only playing roles in supplying ingredients, but autophagy also regulates cell death in both promoting and inhibiting fashions [18]. In breast cancer, autophagy has been reported to confer protection against chemotherapy. Adriamycin-resistant MCF-7/ADM cells maintain a high basal level of autophagy and inhibition of autophagy counteracts their resistance to the drug [19]. Moreover, survival of dormant CSCs in secondary sites is dependent on autophagy and autophagy helps their subsequent growth as macrometastases [3, 20, 21].
The master regulator gene(s) which confers these diverse phenotypes to BCSCs remains unidentified. Although conventional oncogenes, such as KRAS and EGFR, play roles in both BCSCs and their progeny cells, their activation cannot fully explain the complexed phenotypes [22]. Recently, evidences showing implication of miRNAs in oncogenesis have been emerged [23]. miRNAs are short 20–22 nucleotide RNA molecules that are negative regulators of gene expression in a variety of eukaryotic organisms [24], and pro-oncogenic “onco-miR” and antioncogenic “anti-onco-miR” can modulates multiple genes by interacting 3′UTR of target genes [23, 25, 26], thus miRNA might explain the multiple features of BCSCs [27, 28].
In this study, we performed miRNA profiling between BCSC-mimicking mammosphere and their parental cancer cell lines, and identified hsa-miR-27a was specifically downregulated in the BCSC model. Overexpression of hsa-miR-27a attenuated formation and survival of mammosphere, and ameliorated drug resistance to chemotherapeutic compounds through downregulation of ROS-detoxifying genes. Consistently, inhibition of hsa-miR-27a augmented both basal and induced autophagic flux in breast cancer cells which confers protection against chemotherapy. Although further in vivo studies are necessary, enhancement of hsa-miR-27a signaling could be an attractive modality for treatment of relapsed/metastasized breast cancer.
Materials and methods
Cell Lines
Breast cancer cell lines were obtained from the American Type Culture Collection (Manassas, VA, USA). MCF-7 cells were cultured in RPMI1640 medium (Life Technology, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS) (Cell Culture Technologies, Canada) and penicillin/streptomycin (Sigma, St. Louis, MO, USA). MDA-MB-231 cells were cultured in Leibovitz’s L-15 Medium (Sigma) supplemented with 10% FBS, penicillin/streptomycin, l-glutamine (Sigma) and 2 g/L of NaHCO3 (Wako Pure Chemical Industries, Osaka, Japan). All cells were grown at 37 °C in a humidified atmosphere containing 5% CO2.
miRNA microarray analysis
Total RNA (100 ng/sample) was isolated using ISOGEN (Nippon Gene, Tokyo, Japan), and then labeled and hybridized using the Human microRNA (miRNA) Microarray kit (Agilent Technologies, Santa Clara, CA, USA) according to the version 1.5 of the “Protocol for use with Agilent miRNA microarrays.” Hybridized signals were detected on a G2505B DNA microarray scanner (Agilent Technologies), and all scanned images were analyzed using the Agilent Feature Extraction software (v9.5.3.1).
miRNA transfection
The sequences of miRNA mimics used in this study were as follows: hsa-miR-27a sense (5′-AGGGCUUAGCUGCUUGUGAGCA-3′) and antisense (5′-UUCACAGUGGCUAAGUUCCGC-3′); hsa-let-7a-1 sense (5′-UGAGGUAGUAGGUUGUAUAGUU-3′) and antisense (5′-CUAUACAAUCUACUGUCUUUC-3′); nontargeted control miRNA sense (5′-AUCCGCGCGAUAGCACGUAUU-3′) and antisense (5′-UACGUACUAUCGCGCGGAUUU-3′). The miRNA mimics were transfected with HiPerFect reagent (final concentration, 100 nM; Qiagen, Hilden, Germany).
Quantitative real-time PCR assays
MiRNAs were quantified using TaqMan® miRNA Assays (Thermo Fischer Scientific Inc., USA) with some modifications [29]. Briefly, 10 ng of total RNA was reverse transcribed (RT) using the TaqMan® miRNA RT Kit. For amplifying miRNA specific stem-loop, TaqMan® primers (hsa-miR-27a, 000408; hsa-miR-29b, 000413; hsa-miR-101, 002253; hsa-miR-125a-5p, 002198; hsa-miR-193a-3p, 002250; RNU6B, 001093; Applied Biosystems) were used. Quantitative RT-PCR using FastStart Universal Probe Master Mix (Roche Diagnostics) was used for quantification of mRNA. The sequences of mRNA primers and probes were listed in Supplementary Table 1. Data were analyzed using the Light Cycler 96 Software Ver.1.1 (Roche Diagnostics GmbH). The comparative Ct method was used for relative quantitation of samples.
Cell viability
The breast cancer cells were treated with paclitaxel (Pac) or doxorubicin (DXR) at indicated concentration for 24 h. The cell viability after the treatment was determined using Cell Count Reagent SF (Nacalai Tesque, Kyoto, Japan).
Measurement of ROS
Breast cancer cells were incubated with 10 μM 2′,7′-dichlorofluorescein diacetate (DCFH-DA, Sigma) for 15 min at 37 °C, washed with phosphate-buffered saline (PBS) with 2% FBS, and subjected to flow cytometric analysis on BD FACS Aria III (BD Biosciences, San Jose, CA, USA).
FACS analysis
MCF-7 and MDA-MB-231 cells transfected with control miRNA, let-7a-1, or hsa-miR-27a. After 2 days of transfection, cells were dissociated by exposure to 0.25% trypsin/EDTA. Cells were incubated with anti-human CD44v9 antibody (3 μg/ml in 0.2% BSA in PBS, LKG-M001, Cosmo Bio, Japan) for 30 min on ice, washed with PBS with 2% FBS. Cells were incubated with phycoerythrin-conjugated antibody to rat IgG2a (2 μg/ml in 0.2% BSA in PBS, 558067, BD Pharmingen) for 30 min on ice, washed with PBS with 2% FBS. FACS was performed with BD FACS Aria III (BD Biosciences, San Jose, CA, USA). Aldehyde dehydrogenase (ALDH) activity was determined with ALDEFLUOR assay (STEMCELL Technologies, Canada).
GSH assay
Intracellular levels of GSH were determined using the GSH-Glo Glutathione Assay kit (Promega), as described previously [30].
Western blot analysis and autophagy assay
Breast cancer cells, lysed with TNE buffer containing 1 mM of PMSF (Sigma), were subjected to SDS-PAGE followed by western blot (WB) analyses. The primary antibodies used for WB were as follows: p62 (1:1000, ab207305, Abcam), beta-actin (1:5000, A1978, Sigma), and Autophagy watch kit (8486. MBL, Japan) for LC3 and tubulin. The signals were visualized with HRP-conjugated anti-mouse IgG secondary antibody (1: 50,000, NA9310, GE Healthcare) or HRP-conjugated anti-rabbit IgG secondary antibody (1:50,000, NA9340, GE Healthcare), and Amersham ECL select detection regent (GE Healthcare).
Statistical analysis
Data were analyzed using the unpaired Student’s t test, and were presented as means ± SD. P value < 0.05 was considered statistically significant and was denoted as follows: *P < 0.05; **P < 0.01; ***P < 0.001.
Results
Hsa-miR-27a expression is downregulated in mammospheres, and hsa-miR-27a overexpression suppresses BCSC properties
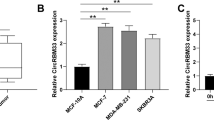
The current CSC hypothesis is based on the existence of fewer number of stem cell-like cells which can initiate self-renewal and also differentiate into multiple types of cells associated with the cancer in the special environment called niche [31]. To dissect the molecular characteristics of BCSCs using cultured cancer cell lines, we need a model which mimic the niche for BCSCs [32]. Mammosphere, a sphere-like structure consisting of three-dimensionally cultured breast cancer cells is thought to mimic the environment of niche and promote BCSC-like phenotypes [33]. First, we successfully established the mammosphere models from MCF-7, HCC-70, and HCC-1954 cell lines. Using these models, we comprehensively profiled miRNAs which differentially expressed between parental adherent cancer cells and mammosphere cells using the miRNA array (Fig. 1a). The results indicated that miR-125a-5p expression was higher in the mammospheres than in their parental adherent breast cancer cells, whereas the expressions of miR-101, miR-193a-3p, miR-27a, and miR-29b were lower in the mammospheres than in their parental adherent breast cancer cells (Fig. 1a). The results were further validated by qRT-PCR with Taqman probes (Fig. 1b). Among them, we were especially interested in miR-27a because miR-29b, miR-101, and miR-193a-3p are already established anti-oncomiRs and the miR-125a-5p can work as both pro-oncogenic and anti-oncogenic [34, 35], and although miR-27a is reported to be oncogenic for breast cancer, its expression is downregulated by formation of mammosphere.
a Relative miRNA expression levels in adherent parental breast cancer cell lines and corresponding mammospheres, as determined using miRNA arrays. b Validation of miRNA expression levels in the parental breast cancer cell lines and corresponding mammospheres, determined by quantitative RT-PCR. The expression level of RNU6B was used as an internal control; n = 3. c Microscopic images of mammospheres established from MDA-MB-231 cells, in the presence or absence of introduction of hsa-miR-27a. The arrowheads show the mammospheres. The scale bar shows 200 μm. d The efficiency and size of mammospheres established from MDA-MB-231 cells, in the presence or absence of introduction of hsa-miR-27a; n = 3. Data are shown as mean ± SD. *P < 0.05; **P < 0.01; ***P < 0.001 (Student’s t test).
To investigate the role of hsa-miR-27a in maintenance of the BCSC properties, we examined the role of hsa-miR-27a in the mammosphere harboring more BCSC-like phenotypes. The established cancer stem cell markers for BCSC is the ratio of CD44+/CD24− [33, 36], and MDA-MB-231 cell line has been shown to have high CD44+/CD24− ratio, suggesting that MDA-MB-231 may have similar characteristics with BCSCs. MDA-MB-231 cells, transfected with hsa-miR-27a, formed 0.36-fold fewer number of mammospheres than control cells transfected without the miRNA mimic (Fig. 1c, d). In addition, the size of mammospheres derived from MDA-MB-231 cells transfected with hsa-miR-27a was smaller (0.43-fold) than the mammospheres from control cells (Fig. 1d). Next, we examined whether the introduction of hsa-miR-27a affects the ALDH activity, which is an established marker for BCSC [37], in breast cancer cells using ALDEFLUOR, a fluorescent substrate for ALDH. The rates of ALDEFLUOR-positive populations in the MCF-7 and MDA-MB-231 cells transfected with hsa-miR-27a were decreased compared with their parental control cells (MCF-7: hsa-miR-27a 4.47% vs. control 6.65%; MDA-MB-231: hsa-miR-27a 26.6% vs. control 41.6%) (Fig. 2a). Consistently, the MDA-MB-231 cells transfected with antisense-miR-27a (as-miR-27a), an antagonistic miRNA mimic against miR-27a, contained a higher proportion of ALDEFLUOR-positive cells than the cells transfected with nontargeting control cells (Fig. 2b).
a Flow cytometry of MDA-MB-231 cells transfected with miR-27a or control, stained with ALDEFLUOR dye. The ALDH-specific inhibitor (DEAB) was used to determine the background fluorescent signals and define the ALDEFLUOR-positive population. SSC side scatter. b Flow cytometry of MDA-MB-231 cells transfected with antagonizing as-miR-27a or as-control, stained with ALDEFLUOR dye. DEAB was used to determine the background fluorescent signals and define the ALDEFLUOR-positive population. SSC side scatter. c Flow cytometric analysis of MCF-7 cells transfected with control miRNA, let-7a-1 or hsa-miR-27a, immunostained with an aggressive type of breast cancer marker anti-human CD44v9 antibody.
The BCSCs population expressing variant of CD44 (CD44v) is known to aggressively disseminate and metastasize [38]. CD44v has been reported to contribute to ROS defense by promoting the synthesis of GSH through stabilization of xCT, a subunit of glutamate-cystine transporter [30], and importantly, 3′UTR of CD44 harbors two partially complementary seed sequence to hsa-miR-27a (miRDB: mirdb.org). To investigate if hsa-miR-27a could suppress the expression of CD44v, we introduced hsa-miR-27a into MCF-7 cells and MDA-MB-231 cells, and measured the rates of cells positive for CD44v. As expected, the overexpression of hsa-miR-27a inhibited the expression of CD44v in both MCF-7 cells and MDA-MB-231 cells (Fig. 2c). These findings indicate that hsa-miR-27a regulates the key features of BCSCs.
Attenuation of chemoresistance of breast cancer cells by miR-27a
Even after vigorous chemotherapy, BCSCs still survive and eventually promote relapse and metastasis in breast cancer patients [39]. If the chemoresistant feature of BCSCs can be mitigated, the outcome could be drastically improved. Therefore, we next investigated whether miR-27a attenuates the chemoresistant capacity of breast cancer cells. We introduced hsa-miR-27a into MCF-7 and MDA-MB-231 cells, and then challenged them with anticancer drugs, DXR and Pac. The microscopic images showed that hsa-miR-27a increased dead cell population and decreased the number of attached live cells after treatment with 100 nM of DXR (Fig. 3a). We also measured the viability of the cells treated with DXR (10 or 100 nM), or Pac (10 or 20 nM), and confirmed that introduction of hsa-miR-27a sensitized MCF-7 and MDA-MB-231 cells to these chemotherapeutic compounds and decreased their viability (Fig. 3b). These data indicate that hsa-miR-27a negatively regulates chemoresistance of breast cancer cells.
a Typical microscopic images of MCF-7 cells and MDA-MB-231 cells transfected with or without hsa-miR-27a. b Viability of breast cancer cell lines transfected with or without hsa-miR-27a, incubated with DXR or Pac at indicated concentrations for 3 days. The error bars show ± SD from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001 (Student’s t test).
Hsa-miR-27a impairs ROS defense by targeting CTH, xCT, and Nrf2
To dissect the molecular mechanism underlying the miR-27a-mediated chemosensitization, we next sought to find molecular targets of miR-27a. Each miRNA can interfere multiple genes by interaction with 3′UTR of mRNAs. We used the bioinformatic prediction tool, miRDB and listed potential target genes of hsa-miR-27a. Among them, we focused on a group of genes related to cellular response to ROS including cystathionine gamma-lyase (CTH), xCT cystine/glutamate transporter (SLC7A11), and Nrf2 (NFE2L2) (Fig. 4a), because activation of these genes promote chemoresistance [40]. The 3′UTRs of these genes are partially complementary to hsa-miR-27a-3p (Fig. 4b). When we introduced hsa-miR-27a mimic to MCF-7, we confirmed that hsa-miR-27a significantly downregulated the expression levels of CTH, SLC7A11, and NFE2L2 (Fig. 4c). In MDA-MB-231 cells, although we observed similar tendency to MCF-7 cells, hsa-miR-27a significantly suppressed only SLC7A11 mRNA expression (Fig. 4c).
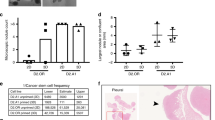
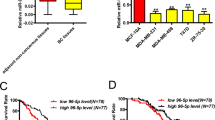
a Functional network of candidate genes targeted by hsa-miR-27a, determined by STRING (string.org). b Sequences of hsa-miR-27a and its binding sites in the 3′UTRs of SLC7A11, CTH, and NFE2L2 mRNAs. c Expression of hsa-miR-27a target genes, determined by RT-qPCR. Data were normalized against the corresponding level of β-actin mRNA. The error bars show ± SD from three independent experiments. d Basal and enhanced ROS levels in the MCF-7 and MDA-MB-231 cells, incubated with or without 0.1 mM H2O2 for 20 min, 3 days after transfection with miRNA mimics. Cellular ROS levels were determined by DCFH-DA. e Intracellular GSH/GSSG ratio 2 days after transfection with miRNA mimics. n = 3. Data are shown as mean ± SD. *P < 0.05; **P < 0.01; ***P < 0.001 (Student’s t test).
We next sought to determine whether the expression of hsa-miR-27a is functionally associated with ROS defense in breast cancer cells. To address this question, we measured the intracellular ROS levels in MCF-7 and MDA-MB-231 cells transfected with hsa-miR-27a by staining with the fluorescent ROS probe DCFH-DA. Because these potential target genes are involved in detoxification of ROS in cells and especially Nrf2 is the master regulator of cytosolic response to ROS, downregulation of these genes would promote higher ROS levels in cells. As expected, hsa-miR-27a-overexpressing cells exhibited higher DCFH-DA staining than control cells (Fig. 4d) under normal culture condition. To further examine the effect of hsa-miR-27a on defense against ROS, we burdened the cells with 0.1 mM hydrogen peroxide (H2O2). After exposure to H2O2, hsa-miR-27a-overexpressing MCF-7 cells and MDA-MB-231 cells exhibited higher DCFH-DA staining than control cells (Fig. 4d).
Cytosolic redox state is maintained as reduced condition by abundant existence of GSH [41, 42]. The methionine cycle is the major source for cysteine, a precursor of GSH in the trans-sulfuration pathway and cysteine availability is rate-limiting for GSH synthesis [43]. In the trans-sulfuration pathway, cystathionine γ-lyase (CTH/CSE) converts cystathionine into cysteine. Cystine is taken up by cells primarily through xCT, a heterodimeric protein consisting of a transporter subunit and a regulatory subunit (4F2hc, also known as CD98, encoded by the SLC3A2) [44]. In cells, cystine is reduced to cysteine and used for synthesis of GSH. Therefore, if hsa-miR-27a targets CTH and xCT (SLC7A11), production of GSH will be attenuated. To confirm this, we measured intracellular GSH level. As expected, transfection of hsa-miR-27a significantly lowered the cytosolic GSH in MCF-7 cells compared with control cells (Fig. 4e). We also observed similar tendency in MDA-MB-231 but it was not statistically significant, probably due to poor downregulation of CTH in MDA-MB-231 cells (Fig. 4c). Overall, these results suggest that expression of hsa-miR-27a increases intracellular ROS in MCF-7 cells by attenuating Nrf2 signaling and GSH synthesis pathway.
Inhibition of the expression of miR-27a induces autophagy
Another big problem of BCSCs to be conquered is that they can survive for years or decades in dormancy after chemotherapy. Still little is known about the mechanisms responsible for it; however, several studies have suggested that autophagy is implicated in this process, and autophagy supplies nutrition required for the long nap [45].
To determine whether miR-27a regulates autophagic activity of breast cancer cells, we measured autophagy flux in the breast cancer cells and examined whether it is affected by transient transfection of antagonizing as-hsa-miR-27a. Once autophagic signaling is initiated, LC3, a mammalian homolog of ATG8 is lipidated and converted to LC3-II which associates with the autophagosome membrane. The amount of LC3-II correlates with the number of autophagosomes, thus it is used for measurement of activity of autophagy. When we antagonized endogenous hsa-miR-27a by as-hsa-miR-27a, which mimicked the condition of BCSCs, LC3-II was increased in MCF-7 cells at 72 h after as-hsa-miR-27a transfection (Fig. 5a). p62 is autophagic substrate and eventually delivered to lysosomes for degradation [46]. Hence, we measured the protein levels of p62. The expression of p62 was increased in MCF-7 and MDA-MB-231 cells at 48 h, and gradually decreased by 72 h (Fig. 5b).
a Expression of LC3-II 48 and 72 h after transfection with as-hsa-miR-27a. (**P < 0.01; data are expressed relative to the corresponding value for vehicle-treated cells and are shown as means ± SD from quadruplicate experiments.) b Expression of p62 at 48 and 72 h by as-hsa-miR-27a post transfection.
Together, these data suggested that hsa-miR-27a negatively regulates autophagic flux and BCSCs with lower expression of hsa-miR-27a may benefit from nutrition supply from enhanced autophagy which is essential for dormancy for decades.
Discussion
Breast cancer is one of the most common types of cancer among women throughout the world [47]. Accumulating evidences suggest that metastatic tumor cells have already disseminated at an early stage of breast cancer [6, 48, 49]. These disseminated cells at the micrometastatic sites harbor features of CSCs, dormancy and chemoresistance [5, 6]. Therefore, for further improvement of the prognosis for breast cancer patients, we need to target BCSCs; however, currently there is no available therapy for BCSCs. To identify a therapeutic target for BCSCs, we comprehensibly profiled miRNAs differentially expressed between mammosphere and the parental cancer cells, and found that hsa-miR-27a is the master negative regulator of BCSC phenotypes. In mammosphere hsa-miR-27a was downregulated. The overexpression of hsa-miR-27a suppressed the mammosphere formation and downregulated expression of BCSC markers. Furthermore, overexpression of hsa-miR-27a chemosensitized breast cancer cells via attenuation of ROS-defense system and autophagy. Therefore, hsa-miR-27a is a possible therapeutic target for BCSCs.
BCSCs, characterized by dormancy and chemoresistance, play roles in relapse and metastasis of breast cancer. The dormant stage without detectable mitosis is one of notable phenotypes to distinguish BCSCs and breast cancer cells. When patients undergo chemotherapy, actively proliferating cells are killed by chemotherapeutic reagents but the dormant cells are chemoresistant and survive for years. To date, biomarkers including CD44+/CD24− ratio and ALDH activity are established as markers for BCSCs [36, 37], and the expression of CD44v is shown to be involved in the more aggressive metastatic phenotype [30]. As we demonstrated above, hsa-miR-27a suppressed expression of all of these markers and activity of ALDH, suggesting that hsa-miR-27a is a master negative regulator of stemness of breast cancer. Previously, hsa-miR-27a is reported as an oncomir and its expression is high in breast cancer cells, promoting cancer cell proliferation, and inhibition of hsa-miR-27a decreases the proliferation [50, 51]. The downregulation of a tumor-suppressive transcription factor FOXO by hsa-miR-27a is implicated in this phenotype, and inactivation of FOXO results in rapid turnover of cell cycles and increased number of cancer cells [50]. Therefore, hsa-miR-27a promotes cell proliferation in breast cancer cells. In this study, however, we identified novel roles of hsa-miR-27a in maintenance of features of BCSCs and hsa-miR-27a inhibits proliferation of breast cancer cells resulting in enhanced chemosensitivity and imbalance of cytosolic ROS. Which cellular component(s) determines these two controversial biological effects of hsa-miR-27a remains to be investigated, but we speculated that microenvironment in mammosphere culture converts the breast cancer cells to be more static state, and slowed cell cycle progression contribute to the discrepancy.
Detoxification of ROS is a big issue for cancer cell to survive the chemotherapy. DXR and Pac, which are widely used chemotherapeutic compounds for breast cancer, produce ROS to induce apoptosis in breast cancer cells [52, 53], and to deal with it, breast cancer cells need to sense the aberrant ROS and neutralize the ROS. Importantly, BCSCs harbor highly active antioxidative machineries and their enhanced production of GSH contributes to the survival under chemotherapy [54]. When we introduced hsa-miR-27a mimic to breast cancer cells, it downregulated not only Nrf2 but also CTH and xCT, both are essential for GSH production, and thus increased intracellular ROS level and susceptibility of breast cancer cells to DXR and Pac. The cancer cells with disturbed antioxidative response are also prone to DNA damages by radiation and thereby targeting hsa-miR-27a signaling will enhance the effect of both chemotherapy and radiotherapy.
Another determinant of survival of BCSCs is autophagy. After long dormant period, metastatic BCSCs cause relapse and it is a major cause of death of patients with breast cancer. Pharmacologic or genetic inhibition of autophagy significantly decreases cell survival and metastasis of breast cancer in animal models and in vivo preclinical models of dormancy [45]. The Inhibition of autophagy in BCSCs results in accumulation of ROS and apoptosis [45]. Therefore, maintenance of autophagic flux is critical for survival of BCSCs. In addition, the enhanced autophagy promotes chemoresistant in MCF-7 cells [19], and basal and starvation-induced autophagy flux is higher in the ALDH+ population derived from mammospheres than the bulk population [46], implying the roles of autophagy not only in the maintenance of dormancy but acquiring chemoresistance. In this study, we demonstrated that hsa-miR-27a negatively regulates autophagy flux in MCF-7 and MDA-MB-231 cells, and thus hsa-miR-27a inhibits phenotypes of BCSCs via multiple mechanisms. Why BCSCs have higher autophagic flux remains unknown. Intracellular ROS is an established trigger for autophagy but the ROS level in BCSCs is maintained lower than their progeny thus it cannot explain it, and further studies are expected.
In summary, we identified hsa-miR-27a as the master negative regulator of BCSC phenotypes through impairing ROS defense and autophagy. These our in vitro data will be further strengthened by a future study using animal models. These novel insights give us a deeper understanding of the role of miRNA in maintaining BCSC features, and providing us novel molecular targets for breast cancer.
Data availability
All datasets analyzed during the present study are available from the corresponding author on reasonable request.
References
Karrison TG, Ferguson DJ, Meier P. Dormancy of mammary carcinoma after mastectomy. J Natl Cancer Inst. 1999;91:80–5.
Klein CA. Parallel progression of primary tumours and metastases. Nat Rev Cancer. 2009;9:302–12.
Sosa MS, Bragado P, Aguirre-Ghiso JA. Mechanisms of disseminated cancer cell dormancy: an awakening field. Nat Rev Cancer. 2014;14:611–22.
Chambers AF, Groom AC, MacDonald IC. Dissemination and growth of cancer cells in metastatic sites. Nat Rev Cancer. 2002;2:563–72.
Demicheli R. Tumour dormancy: findings and hypotheses from clinical research on breast cancer. Semin Cancer Biol. 2001;11:297–306.
Barkan D, Kleinman H, Simmons JL, Asmussen H, Kamaraju AK, Hoenorhoff MJ, et al. Inhibition of metastatic outgrowth from single dormant tumor cells by targeting the cytoskeleton. Cancer Res. 2008;68:6241–50.
Gao H, Chakraborty G, Lee-Lim AP, Mo Q, Decker M, Vonica A, et al. The BMP inhibitor Coco reactivates breast cancer cells at lung metastatic sites. Cell. 2012;150:764–79.
Nguyen DX, Bos PD, Massague J. Metastasis: from dissemination to organ-specific colonization. Nat Rev Cancer. 2009;9:274–84.
Valastyan S, Weinberg RA. Tumor metastasis: molecular insights and evolving paradigms. Cell. 2011;147:275–92.
La Belle Flynn A, Calhoun BC, Sharma A, Chang JC, Almasan A, Schiemann WP. Autophagy inhibition elicits emergence from metastatic dormancy by inducing and stabilizing Pfkfb3 expression. Nat Commun. 2019;10:3668.
Diehn M, Cho RW, Lobo NA, Kalisky T, Dorie MJ, Kulp AN, et al. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature. 2009;458:780–3.
Kansanen E, Kuosmanen S, Leinonen H, Levonen A. The Keap1–Nrf2 pathway: mechanisms of activation and dysregulation in cancer. Redox Biol. 2013;1:45–9.
Edwards MR, Johnson B, Mire CE, Xu W, Shabman RS, Speller LN, et al. The Marburg virus VP24 protein interacts with Keap1 to activate the cytoprotective antioxidant response pathway. Cell Rep. 2014;6:1017–25.
Du Y, Villeneuve NF, Wang XJ, Sun Z, Chen W, Li J, et al. Oridonin confers protection against arsenic-induced toxicity through activation of the Nrf2-mediated defensive response. Environ Health Perspect. 2008;116:1154–61.
Motohashi H, Yamamoto M. Nrf2-Keap1 defines a physiologically important stress response mechanism. Trends Mol Med. 2004;10:549–57.
Baird L, Dinkova-Kostova AT. The cytoprotective role of the Keap1-Nrf2 pathway. Arch Toxicol. 2011;85:241–72.
Narasimhan M, Patel D, Vedpathak D, Rathinam M, Henderson G, Mahimainathan L. Identification of novel microRNAs in post-transcriptional control of Nrf2 expression and redox homeostasis in neuronal, SH-SY5Y cells. PLoS ONE. 2012;7:e51111.
Maiuri MC, Zalckvar E, Kimchi A, Kroemer G. Self-eating and self-killing: crosstalk between autophagy and apoptosis. Nat Rev Mol Cell Biol. 2007;8:741–52.
Zhang P, Liu X, Li H, Chen Z, Yao X, Jin J, et al. TRPC5-induced autophagy promotes drug resistance in breast carcinoma via CaMKKbeta/AMPKalpha/mTOR pathway. Sci Rep. 2017;7:3158.
Mowers EE, Sharifi MN, Macleod KF. Autophagy in cancer metastasis. Oncogene. 2017;36:1619–30.
Lu Z, Luo RZ, Lu Y, Zhang X, Yu Q, Khare S, et al. The tumor suppressor gene ARHI regulates autophagy and tumor dormancy in human ovarian cancer cells. J Clin Investig. 2008;118:3917–29.
Harrison H, Simoes BM, Rogerson L, Howell SJ, Landberg G, Clarke RB. Oestrogen increases the activity of oestrogen receptor negative breast cancer stem cells through paracrine EGFR and Notch signalling. Breast Cancer Res. 2013;15:R21.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–33.
Papagiannakopoulos T, Kosik KS. MicroRNAs: regulators of oncogenesis and stemness. BMC Med. 2008;6:15.
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C, et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell. 2007;131:1109–23.
Tanaka M, Oikawa K, Takanashi M, Kudo M, Ohyashiki J, Ohyashiki K, et al. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS ONE. 2009;4:e5532.
Ishimoto T, Nagano O, Yae T, Tamada M, Motohara T, Oshima H, et al. CD44 variant regulates redox status in cancer cells by stabilizing the xCT subunit of system xc(-) and thereby promotes tumor growth. Cancer Cell. 2011;19:387–400.
Kreso A, Dick JE. Evolution of the cancer stem cell model. Cell Stem Cell. 2014;14:275–91.
Ponti D, Costa A, Zaffaroni N, Pratesi G, Petrangolini G, Coradini D, et al. Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res. 2005;65:5506–11.
Dontu G, Abdallah WM, Foley JM, Jackson KW, Clarke MF, Kawamura MJ, et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 2003;17:1253–70.
Balatti V, Tomasello L, Rassenti LZ, Veneziano D, Nigita G, Wang HY, et al. miR-125a and miR-34a expression predicts Richter syndrome in chronic lymphocytic leukemia patients. Blood. 2018;132:2179–82.
Yan L, Yu MC, Gao GL, Liang HW, Zhou XY, Zhu ZT, et al. MiR-125a-5p functions as a tumour suppressor in breast cancer by downregulating BAP1. J Cell Biochem. 2018;119:8773–83.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA. 2003;100:3983–8.
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007;1:555–67.
Zhang H, Brown RL, Wei Y, Zhao P, Liu S, Liu X, et al. CD44 splice isoform switching determines breast cancer stem cell state. Genes Dev. 2019;33:166–79.
Iliopoulos D, Hirsch HA, Struhl K. Metformin decreases the dose of chemotherapy for prolonging tumor remission in mouse xenografts involving multiple cancer cell types. Cancer Res. 2011;71:3196–201.
No JH, Kim YB, Song YS. Targeting Nrf2 signaling to combat chemoresistance. J Cancer Prev. 2014;19:111–7.
Willems PH, Rossignol R, Dieteren CE, Murphy MP, Koopman WJ. Redox homeostasis and mitochondrial dynamics. Cell Metab. 2015;22:207–18.
Ursini F, Maiorino M, Forman HJ. Redox homeostasis: the golden mean of healthy living. Redox Biol. 2016;8:205–15.
Bannai S, Ishii T. Transport of cystine and cysteine and cell growth in cultured human diploid fibroblasts: effect of glutamate and homocysteate. J Cell Physiol. 1982;112:265–72.
Okuno S, Sato H, Kuriyama-Matsumura K, Tamba M, Wang H, Sohda S, et al. Role of cystine transport in intracellular glutathione level and cisplatin resistance in human ovarian cancer cell lines. Br J Cancer. 2003;88:951–6.
Vera-Ramirez L, Vodnala SK, Nini R, Hunter KW, Green JE. Autophagy promotes the survival of dormant breast cancer cells and metastatic tumour recurrence. Nat Commun. 2018;9:1944.
Gong C, Bauvy C, Tonelli G, Yue W, Delomenie C, Nicolas V, et al. Beclin 1 and autophagy are required for the tumorigenicity of breast cancer stem-like/progenitor cells. Oncogene. 2013;32:2272e 2261–2211.
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Braun S, Vogl FD, Naume B, Janni W, Osborne MP, Coombes RC, et al. A pooled analysis of bone marrow micrometastasis in breast cancer. N Engl J Med. 2005;353:793–802.
Pantel K, Schlimok G, Braun S, Kutter D, Lindemann F, Schaller G, et al. Differential expression of proliferation-associated molecules in individual micrometastatic carcinoma cells. J Natl Cancer Inst. 1993;85:1419–24.
Guttilla IK, White BA. Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. J Biol Chem. 2009;284:23204–16.
Mertens-Talcott SU, Chintharlapalli S, Li X, Safe S. The oncogenic microRNA-27a targets genes that regulate specificity protein transcription factors and the G2-M checkpoint in MDA-MB-231 breast cancer cells. Cancer Res. 2007;67:11001–11.
Alexandre J, Batteux F, Nicco C, Chereau C, Laurent A, Guillevin L, et al. Accumulation of hydrogen peroxide is an early and crucial step for paclitaxel-induced cancer cell death both in vitro and in vivo. Int J Cancer. 2006;119:41–8.
Alexandre J, Hu Y, Lu W, Pelicano H, Huang P. Novel action of paclitaxel against cancer cells: bystander effect mediated by reactive oxygen species. Cancer Res. 2007;67:3512–7.
Lu H, Samanta D, Xiang L, Zhang H, Hu H, Chen I, et al. Chemotherapy triggers HIF-1-dependent glutathione synthesis and copper chelation that induces the breast cancer stem cell phenotype. Proc Natl Acad Sci USA. 2015;112:E4600–9.
Funding
This research was supported by Grants-in-Aid for Scientific Research (17K10564 and 17H04067), a Grant-in-Aid for Exploratory Research from Japan Society for the Promotion of Science (JSPS), and the Strategic Research Foundation Grant-aided Project for Private Universities from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT).
Author information
Authors and Affiliations
Contributions
SU designed the study; SU, MT, and KS performed the wet-lab experiments; SU did data analysis and statistical analysis; SU, MT, and KK performed interpretation of data; SU, MT, KK, and MK drafted the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ueda, S., Takanashi, M., Sudo, K. et al. miR-27a ameliorates chemoresistance of breast cancer cells by disruption of reactive oxygen species homeostasis and impairment of autophagy. Lab Invest 100, 863–873 (2020). https://doi.org/10.1038/s41374-020-0409-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-020-0409-4
This article is cited by
-
Targeting autophagy in prostate cancer: preclinical and clinical evidence for therapeutic response
Journal of Experimental & Clinical Cancer Research (2022)
-
MicroRNAs as a clue to overcome breast cancer treatment resistance
Cancer and Metastasis Reviews (2022)
-
Regulation of autophagy by microRNAs in human breast cancer
Journal of Biomedical Science (2021)
-
Quantitative mapping of the cellular small RNA landscape with AQRNA-seq
Nature Biotechnology (2021)
-
Revisiting cancer hallmarks: insights from the interplay between oxidative stress and non-coding RNAs
Molecular Biomedicine (2020)