Abstract
Programmed death 1 ligand 1 (PD-L1) Immunohistochemistry (IHC) is the key FDA-approved predictive marker to identify responders to anti-PD1 axis drugs. Multiple PD-L1 IHC assays with various antibodies and cut points have been used in clinical trials across tumor types. Comparative performance characteristics of these assays have been extensively studied qualitatively but not quantitatively. Here we evaluate the use of a standardized PD-L1 Index tissue microarray (TMA) to objectively determine agreement between antibody assays for PD-L1 applying quantitative digital image analysis. Using a specially constructed Index TMA containing a panel of ten isogenic cell lines in triplicate, we tested identical but independently grown batches of isogenic cells to prove Index TMAs can be produced in large quantities and hence serve as a standardization tool. Then the Index TMAs were evaluated using quantitative immunofluorescence (QIF) to validate the TMA itself and also to compare antibodies including E1L3N, SP142 and SP263. Next, an inter-laboratory and inter-assay comparison of 5 PD-L1 chromogenic IHC assays (US Food and Drug Administration (FDA) approved and lab developed test (LDT)) were performed at 12 sites around the USA. As previously reported, the SP142 FDA assay failed to detect low levels of PD-L1 in cell lines distinguished by the other four assays. The assays for 22C3 FDA, 28-8-FDA, SP263 FDA, and E1L3N LDT were highly similar across sites and all laboratories showed a high consistency over time for all assays using this Index TMA. In conclusion, we were able to objectively quantify PD-L1 expression on a standardized Index TMA using digital image analysis and we confirmed previous subjective assessments of these assays, but now in a multi-institutional setting. We envision commercial use of this Index TMA or similar smaller version as a useful standardization mechanism to compare results between institutions and to identify abnormalities while running routine clinical samples.
Similar content being viewed by others
Introduction
Immune checkpoint blockade inhibitors have changed the landscape of cancer treatment in the last decade, demonstrating unprecedented clinical success in several tumor types [1,2,3]. Simultaneously, PD-L1 (programmed cell death 1 ligand 1) expression has been identified as a predictive diagnostic marker to select patients that may benefit from anti-PD-1 (programmed cell death 1) axis agents such as nivolumab, pembrolizumab, atezolizumab, and durvalumab [4,5,6,7]. With each drug, a unique proprietary diagnostic test has been developed. Currently, there are multiple qualitative PD-L1 assays, involving various antibodies, to assess the expression of PD-L1 by immunohistochemistry (IHC) using chromogenic methods. The US Food and Drug Administration (FDA) approved some of them as either companion and/or complementary diagnostic tests for specific drugs and cancers. The PD-L1 IHC 22C3 PharmDx kit (Agilent Technologies Inc.) is the only companion diagnostic test approved by FDA for pembrolizumab in non-small cell lung cancer (NSCLC), whereas the Ventana PD-L1 SP142 Assay® (Roche Ventana Medical Systems, Inc.) is a diagnostic test for atezolizumab that the FDA approved as a complementary assay in NSCLC and as a companion assay in patients with urothelial cancer. In addition, the range of cut points for defining positive cases and cell type expression (tumor or immune cells) is widely variable across clinical trials.
In the past years, there has been a significant effort to compare the performance characteristics of these PD-L1 IHC assays in patient samples [8,9,10,11,12,13,14,15]. Recently, Tsao et al. [16] published the results of the second phase of the Blue Print study, confirming the interchangeability of 22C3, 28-8, and SP263 assays and the lower sensitivity of SP142 assay in lung cancer, after PD-L1 scoring by 25 experienced pathologists. Moreover, concordance between PD-L1 IHC assays has been assessed by correlating levels of protein detected by the corresponding assay and the level of PD-L1 mRNA assessed by RNAscope assay [9]. Although there is a high concordance between trained pathologists for PD-L1 scoring in tumor cells [8, 10,11,12, 16], there is a poor reliability when PD-L1 is evaluated in immune cells or at low PD-L1 scores [16, 17].
The assessment of PD-L1 in all previous studies has been on the entire assay, combining the subjective interpretation with the level of expression and the localization. It is possible that these variables are confounding, and variable levels of expression may not be discernable when combined with both localization and subjective interpretation. Despite the increasing number of publications on this topic, there is very little work done on the analytic assessment of expression separated from the interpretation. Here, we propose the use of a standardized Index TMA to objectively compare PD-L1 IHC assays using quantitative image analysis. To achieve this goal, an Index TMA with isogenic cell lines expressing PD-L1 spanning a predetermined dynamic range was built and quantitatively evaluated for PD-L1 expression using both chromogenic and fluorescent IHC methods. In addition, the Index TMA was tested independently across multiple institutions that used diverse PD-L1 chromogenic IHC assays, both FDA-approved and laboratory developed test (LDT), to compare the assays and to illustrate the utility of analytic standardization between institutions.
Materials and methods
Index TMA construction
A panel of 15 isogenic cell lines expressing various amounts of PD-L1 was sourced from Horizon Dx including production and growth of cell lines. After review, ten lines were selected, and an Index TMA was constructed according to the map in Fig. 1. Three independent batches of cell lines were cultured, each ~2 months apart, to produce three independent batches of formalin-fixed, paraffin-embedded cell pellets blocks for batch-to-batch concordance and correlation assessment.
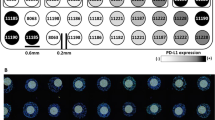
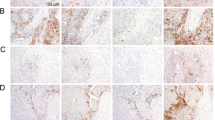
Map of the Index TMA for standardization of PD-L1 measurement. a Schematic map including the cell line code, where the color gradient corresponds to the level of PD-L1 expression (black = high expression; white = no expression). b Representative image of standardized TMA stained with 22C3 IHC assay at low magnification, including three spots at high magnification of negative spot (“1”, cell line 8063), low expresser (“2”, cell line 11221), and high expresser (“3”, cell line 11185). Bar = 200 µm
PD-L1 IHC assays
PD-L1 expression was evaluated by quantitative immunofluorescence (QIF) and chromogenic IHC using five monoclonal antibodies (Supplementary Table 1), including both LDT- and FDA-approved assays. For QIF, clones E1L3N (#13684, Cell Signaling Technology, Inc.), SP142 (#M4420, SpringBio), and SP263 (#790–4905, Ventana Medical Systems, Inc.) were assessed. For chromogenic IHC, automated systems were used for different clones using our own protocol for the LDT E1L3N (#13684, Cell Signaling Technology, Inc.) on multiple platforms, and protocols specified by corresponding manufacturer per the FDA labeling for 22C3 (#SK006, Dako) and 28-8 (#SK005, Dako) with the Dako Autostainer Link 48 Instrument (Dako). Similarly, on-label protocols were used for FDA-approved assays; SP263 (#740–4907) and SP142 (#740–4859) both from Ventana Medical Systems, Inc. on the Benchmark Ultra (Ventana Medical Systems, Inc.). For the multi-institutional comparison, twelve 5-µm sections per PD-L1 assay were cut from a block of Index TMA at Yale University and sent to 12 institutions for staining weekly during 6 consecutive weeks, running two slides per week with their clinical workload using the assay of choice for each institution.
Quantitative immunofluorescence
TMA slides were stained using a protocol previously described by our laboratory [10]. Primary antibodies for PD-L1 E1L3N and SP142 were incubated overnight at 4 °C and clone SP263 was incubated for 20 minutes at 37 °C. An optimized final concentration for each antibody was used [10], except for SP263 which was only available from the company as a prediluted reagent and hence not optimizable (Supplementary Table 1). Then, slides were incubated in rabbit EnVision reagent (K4009, Dako) for 1 h at RT and Cy5-Tyramide (SAT705A001EA, PerkinElmer) was used to amplify target signal. Finally, TMAs were stained with 1:250 4,6-diamidino-2-phenylindole for 10 min at RT and mounted with Prolong Gold antifade mounting reagent (P36394, Life Technologies). Image analysis was performed using AQUA method of QIF (NavigateBP), which generates a score by dividing the sum of target pixel intensities by the area of the molecularly designated compartment. Scores are automatically normalized to lamp hours, bit depth and CC intensity of the microscope to account for day to day variation.
Digital image analysis on chromogenic IHC
After chromogenic staining, as described above, slides were scanned on the Aperio ScanScope XT platform. Then, PD-L1 expression on the cell lines was quantified using the open-source software QuPath [18] (Queen’s University of Belfast, Northern Ireland). Briefly, once images were loaded, cores were automatically selected through the TMA annotation module, and later classified by the module “positive cell detection”. An optimized algorithm was used for cell segmentation based on the size of the nucleus and cell expansion, and for DAB intensity quantification of PD-L1 expression for all antibodies looking at the “cell: DAB optical density (OD) mean” score compartment, among the possible options (nucleus, cytoplasm, cell). The settings were adjusted to avoid false positive detection. Results were shown as percentage of PD-L1+ cells or as OD of the chromogenic staining divided by mm2.
Statistical analysis
All datasets were analyzed and plotted using GraphPad Prism v7.0 software for Windows (GraphPad Software, Inc.). Bar graphs were used to plot average of QIF scores and percentage of PD-L1+ cells and OD/mm2 for DAB staining quantified by QuPath, per cell line from for each assay. Due to the complexity of this work, we define the following terms and concepts related to standardization of measurement: reproducibility, a component of the precision, determines the agreement among results obtained from testing the same substance by using the same test protocol under repeatable conditions (different operators, locations and time); correlation, a statistical relationship to quantify the strength of association between two variables; and concordance, the proportion of the outcomes of a specific test which are identical to an agreed upon reference [19,20,21]. Comparison between blocks, antibodies/assays and laboratories was assessed for correlation using a linear regression coefficient (R2) and Bland–Altman plots [22] were used to assess correlation and concordance, respectively, between PD-L1 assays. Levey–Jennings plots were used to evaluate the quality and the consistency of measurement over time for every laboratory and assay, using both the average of all cell lines or each cell line.
Results
Validation of the Index TMA as a standardized PD-L1 assay
To validate the Index TMA, we quantitatively showed the dynamic range of PD-L1 expression on the TMA and then assess the correlation of Index TMA production. Using the Index TMA described above (Fig. 1 and Supplementary Figure 1), we first measured PD-L1 by QIF using different antibodies with previously optimized conditions [10] (Supplementary Table 1). Lack of staining in cell lines 8063 and 11198 with all antibodies confirmed them as negative controls, as none of them expressed detectable levels of protein by western blot and only 11198 showed very low levels of PD-L1 mRNA (Horizon Dx, unpublished data). The highest levels of PD-L1 were detected on cell lines 11228, 11190, and 11185 by all antibodies. The rest of the clones expressed intermediate amounts of PD-L1 protein in different proportions. As previously described [10, 11, 16], the cellular location of PD-L1 was membranous with all the antibodies in all the cell lines. To quantitatively determine the amount of PD-L1 in each line on the TMA, we performed QIF with each antibody (Fig. 2) showing the full dynamic range of the assay and the relative expression levels determined by each antibody.
Correlation between three independent blocks was assessed by both QIF and chromogenic assays using linear regression (Table 1). For QIF analysis, the correlation between blocks was high for antibodies SP142 (R2 = 0.983–0.995) and E1L3N (R2 = 0.834–0.857), but somewhat lower for SP263 (R2 = 0.655–0.962) most likely due to our inability to optimize the titration for this antibody. Chromogenic assessment utilized QuPath analysis as described above and showed results comparable to the QIF results (Table 1), showing the ability to construct highly reproducible Index TMAs over time using isogenic cell lines.
Comparison of PD-L1 antibodies by QIF and DAB staining using the Index TMA
Correlation and concordance amongst the antibodies or assays were analyzed by linear regression and Bland–Altman plots, respectively. For QIF quantification (Fig. 3a), an average of 27 spots per cell line were evaluated per PD-L1 antibody (three spots per cell line/three independent experiments/three different blocks or batches/per antibody). Correlation of QIF scores obtained with antibodies E1L3N, SP142, and SP263 under LDT conditions was extremely high by linear regression, with coefficients >0.96 similar to that previously described [10]. In addition, the concordance was high as confirmed by Bland–Altman analysis.
Validation of Index TMA with cell lines using three clones of anti-PD-L1 antibody by QIF. a Linear regression and Bland–Altman plots for PD-L1 antibodies (SP142, SP263, and E1L3N) tested by an immunofluorescence protocol developed at the lab (LDT) and previously published [10]. Data represent the average of three independent experiments with three different blocks per antibody used. b Linear regression and Bland–Altman plots for PD-L1 DAB IHC assays (22C3 FDA, 28-8 FDA, E1L3N LDT, SP263 FDA, and SP142 FDA) performed by our CLIA-certified laboratory. Data represent the mean of % PD-L1+ cells per cell line, measured by QuPath platform, of two TMAs per run for each week during ten consecutive weeks (n = 20 slides per assay). For a and b: R2 = coefficient of determination; for Bland–Altman diagrams, continued line represents bias and dotted lines represent upper and lower 95% limits of agreement
Regarding the chromogenic IHC assays, we tested 5 PD-L1 IHC assays (22C3 FDA, 28-8 FDA, SP263 FDA, SP142 FDA, and E1L3N LDT) by running two slides weekly over 10 consecutive weeks in our CLIA-certified research laboratory (Fig. 3b). The chromogenic IHC assays also showed comparably high degree of agreement between 22C3 FDA, 28-8 FDA, and E1L3N LDT assays by Bland–Altman analysis, whereas the concordance between SP142 and SP263 FDA assays was equally poor when compared with the three other assays and themselves (Fig. 3b and Table 2), identifying some of the intermediate expressers as outliers. On the other hand, linear regressions showed the highest correlation between 22C3 FDA, 28-8 FDA and E1L3N LDT assays (R2 = 0.943–0.993) and a slightly lower correlation between SP142 FDA and the above three assays than SP263 FDA and the above three assays (R2 = 0.823–0.871) (Fig. 3b). After comparing SP142 and SP263 FDA assays, linear regression showed a high correlation (R2 = 0.992).
Comparison of PD-L1 chromogenic IHC assays among multiple institutions using the Index TMA
The vision for use of the Index TMA is to be used as a tool for calibration for development of standardization of assays, as shown above, and also to enable methodology transfer and facilitate standardization of assays between institutions, and even more importantly between clinical trials and diagnostic IHC laboratories that will be performing the predictive IHC assays for patient care. To assess the efficiency of this Index TMA for methodology transfer and comparison of analytical performance between different laboratories, the Index cell line TMA was tested independently in 12 institutions across the country, including hospitals and clinical laboratories, using diverse PD-L1 IHC assays and platforms (Table 3). Although they were not run in parallel with clinical samples as the slides for the multi-institutional study, we included the results for SP142 FDA assay from our CLIA-certified research laboratory to increase statistical power for this assay. The chromogenic assays were evaluated both by intensity (OD/mm2) and percentage of positive cells (counting cells above the detection threshold for the assay). As shown in Fig. 4, summarizing 240 slides from 12 institutions and measuring both by % PD-L1+ cells or OD/mm2, all PD-L1 assays performed essentially identically for the very high expressers (cell lines 11228, 11190, and 11185) and for the negative cases (cell lines 8063, 11198). Although, in the negative clones 28-8 and E1L3N show some measurable background signal. Confirming previous qualitative results [11, 16], the SP142 FDA assay showed lower sensitivity failing to detect low levels of PD-L1 in the intermediate cell lines (cell lines 11186–11187) measurable by the other 4 assays (Supplementary Fig. 2). Similar conclusions may be drawn when assessing intensity, although the background signal is more easily illustrated in the nonexpressing cases, showing that even optimized assays show low levels of signal inherent in the IHC technique.
PD-L1 distribution on Index TMA using different PD-L1 DAB IHC assays by multiple institutions. Bar graphs show mean ± SD, as percentage of PD-L1+ cells (a) or OD/mm2 (b) measured for each cell line including all the slides (two per run) run during six consecutive weeks by all the laboratories that ran the specific PD-L1 assay: 22C3 FDA: six labs; 28-8 FDA: six labs; SP263 FDA: three labs; SP142 FDA: two labs (including the CLIA-certified laboratory that participated in the validation of the Index TMA); E1L3N LDT: six labs. The open-source platform QuPath was used to quantify DAB staining. OD optical density for DAB
Having an assay perform with the same sensitivity and specificity is critical for a companion diagnostic test. Figure 5 shows cross institutional comparisons for each assay. To compare the assays from each site to each other we have used regressions (Supplementary Fig. 3 and Table 4).
Finally, another use for the Index TMA is to assure consistency of measurements over time. We assessed this for each of the collaborating institutions across a period of 6 weeks (Fig. 6, Supplementary Figs. 4–13). As shown by the Levey–Jennings plots for assays from collaborating labs, all PD-L1 assays and laboratories showed values between the ±2 SD limits, leading to ~10% false rejections when two levels of control are being analyzed [23].
Levey–Jennings plots of PD-L1 IHC chromogenic assays among all the laboratories during 6 consecutive weeks. Each dot represents the mean of % PD-L1+ cells on all cell lines of two TMAs per run for each week for 22C3 FDA, 28-8 FDA, E1L3N LDT, SP263 FDA, and SP142 FDA assays. “Mean”: total average of all cell lines in all experiments, SD standard deviation of “Mean”, where ±1/2/3 SD corresponds to “Mean” ±1/2/3 SD
Discussion
PD-L1 expression by IHC is used in clinical trials to predict response to anti-PD1/PD-L1 therapies in a variety of cancers [4,5,6,7]. There are 4 PD-L1 IHC assays approved by the FDA, either as companion or complementary diagnostic tests, with different cut points and score systems associated with each assay. In evaluation of these assays, the FDA required proof of quality, reproducibility, and sensitivity for prediction, but was less stringent on analytic sensitivity. Several studies have compared the performance of many PD-L1 assays available by qualitative analysis on tumor samples [8,9,10,11,12,13,14,15], with variable results. In this study, we used an Index TMA with isogenic cell lines expressing PD-L1 in a dynamic range to quantitatively compare PD-L1 IHC assays using two IHC methods and digital image analysis. This approach allowed us to quantitatively assess analytic sensitivity, concordance and correlation of the assays, although the use of isogenic cell lines, not clinical samples, precludes assessment of predictive accuracy [24]. After confirming regression of PD-L1 levels between three independent blocks using diverse antibodies and two immunohistochemical techniques, we observed a high correlation and concordance among the antibodies by QIF by linear regressions and Bland–Altman plots, respectively, confirming previous results [10]. The Index TMA also allowed us to assess the analytic sensitivity of each FDA-approved assay. We find that three of the four FDA-approved assays, and an LDT assay, have comparable analytic sensitivity, but the SP142 assay is less sensitive, failing to detect low level expression that is detected by the other four assays. This observation supports previous qualitative studies [11, 16] and two comparative studies using a commercial cell line TMA, also showing reduced PD-L1 staining with SP142 assay on moderate expressers compared with 22C3, 28-8, and SP263 assays [10, 25].
The use of this Index array allowed the comparison of the PD-L1 assays across both time and institution. The assessment of data from 12 independent institutions revealed that the similarity of the pair 22C3/SP263 and 28-8/E1L3N was the highest (Fig. 4). However, the staining patterns were extremely similar for the negative and the saturated cases for all the DAB IHC assays. Most of the studies suggest the method of detection as the principal parameter for discrepancy among assays rather than the antibody itself [10, 25, 26]. The 12 institutions study also showed good measurement consistency over time at every institution and good correlation by linear regression between institutions doing the same assay. This suggests that the assays themselves should not lead to discordant readings amongst pathologists or, with the exception of the SP142 assay, between assays from different vendors.
There are several limitations to this study. Perhaps the most significant is that this study only tests the analytic sensitivity of the assays or antibodies in the context of isogenic cell lines. It is possible that levels of PD-L1 in actual tumors may be different or that the protein may have posttranslational modifications that are not seen in cell lines. While this concern is recognized, the testing of PD-L1 in actual cancers has been done in many other studies and is not the goal of this work. A second weakness is the use of TMAs. In clinical usage, all of these assays are done on full histologic sections. As such, these data must be taken in the context of its usage as a control, not as an evaluation of issues related to assay performance in patient tissue. Finally, another weakness of this work is the absence of quantitative information from patients with outcome data. While we explore the dynamic range in a model system, we cannot be sure we have replicated the dynamic range seen in patient tumors that have been treated with immunotherapy.
In conclusion, the results generated with this Index TMA are concordant and correlated between independently constructed blocks as assessed by two immunohistochemical methods suggesting this sort of control can be manufactured and used over a period of time. We also suggest that Index TMAs can be a useful tool to compare results from multiple PD-L1 IHC assays and institutions and is also useful as an internal control to identify abnormal runs in a routine clinical laboratory.
References
Brahmer JR, Tykodi SS, Chow LQ, Hwu WJ, Topalian SL, Hwu P, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012;366:2455–65.
Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med. 2012;366:2443–54.
Herbst RS, Soria JC, Kowanetz M, Fine GD, Hamid O, Gordon MS, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014;515:563–7.
Herbst RS, Baas P, Kim DW, Felip E, Perez-Gracia JL, Han JY, et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet. 2016;387:1540–50.
Horn L, Spigel DR, Vokes EE, Holgado E, Ready N, Steins M, et al. Nivolumab versus docetaxel in previously treated patients with advanced non-small-cell lung cancer: two-year outcomes from two randomized, open-label, phase III trials (CheckMate 017 and CheckMate 057). J Clin Oncol. 2017;35:3924–33.
McDermott DF, Sosman JA, Sznol M, Massard C, Gordon MS, Hamid O, et al. Atezolizumab, an anti-programmed death-ligand 1 antibody, in metastatic renal cell carcinoma: long-term safety, clinical activity, and immune correlates from a phase Ia study. J Clin Oncol. 2016;34:833–42.
Antonia SJ, Villegas A, Daniel D, Vicente D, Murakami S, Hui R, et al. Durvalumab after chemoradiotherapy in stage III non-small-cell lung cancer. N Engl J Med. 2017;377:1919–29.
Hirsch FR, McElhinny A, Stanforth D, Ranger-Moore J, Jansson M, Kulangara K, et al. PD-L1 immunohistochemistry assays for lung cancer: results from phase 1 of the blueprint PD-L1 IHC assay comparison project. J Thorac Oncol. 2017;12:208–22.
Tretiakova M, Fulton R, Kocherginsky M, Long T, Ussakli C, Antic T, et al. Concordance study of PD-L1 expression in primary and metastatic bladder carcinomas: comparison of four commonly used antibodies and RNA expression. Mod Pathol. 2018;31:623–32.
Gaule P, Smithy JW, Toki M, Rehman J, Patell-Socha F, Cougot D, et al. A quantitative comparison of antibodies to programmed cell death 1 ligand 1. JAMA Oncol. 2017; 3(2): 256–9.
Rimm DL, Han G, Taube JM, Yi ES, Bridge JA, Flieder DB, et al. A prospective, multi-institutional, pathologist-based assessment of 4 immunohistochemistry assays for PD-L1 expression in non-small cell lung cancer. JAMA Oncol. 2017;3:1051–8.
Batenchuk C, Albitar M, Zerba K, Sudarsanam S, Chizhevsky V, Jin C, et al. A real-world, comparative study of FDA-approved diagnostic assays PD-L1 IHC 28-8 and 22C3 in lung cancer and other malignancies. J Clin Pathol. 2018 71(12):1078–83.
Fujimoto D, Sato Y, Uehara K, Ishida K, Fukuoka J, Morimoto T, et al. Predictive performance of four programmed cell death ligand 1 assay systems on nivolumab response in previously treated patients with non-small cell lung cancer. J Thorac Oncol. 2018;13:377–86.
Hendry S, Byrne DJ, Wright GM, Young RJ, Sturrock S, Cooper WA, et al. Comparison of four PD-L1 immunohistochemical assays in lung cancer. J Thorac Oncol. 2018;13:367–76.
Ratcliffe MJ, Sharpe A, Midha A, Barker C, Scott M, Scorer P, et al. Agreement between programmed cell death ligand-1 diagnostic assays across multiple protein expression cutoffs in non-small cell lung cancer. Clin Cancer Res. 2017;23:3585–91.
Tsao MS, Kerr KM, Kockx M, Beasley MB, Borczuk AC, Botling J, et al. PD-L1 immunohistochemistry comparability study in real-life clinical samples: results of blueprint phase 2 project. J Thorac Oncol. 2018;13:1302–11.
Brunnstrom H, Johansson A, Westbom-Fremer S, Backman M, Djureinovic D, Patthey A, et al. PD-L1 immunohistochemistry in clinical diagnostics of lung cancer: inter-pathologist variability is higher than assay variability. Mod Pathol. 2017;30:1411–21.
Bankhead P, Loughrey MB, Fernandez JA, Dombrowski Y, McArt DG, Dunne PD, et al. QuPath: open source software for digital pathology image analysis. Sci Rep. 2017;7:16878.
Ferrario D, Brustio R, Hartung T. Glossary of reference terms for alternative test methods and their validation. ALTEX. 2014;31:319–35.
Fitzgibbons PL, Bradley LA, Fatheree LA, Alsabeh R, Fulton RS, Goldsmith JD, et al. Principles of analytic validation of immunohistochemical assays: guideline from the college of American Pathologists Pathology and Laboratory Quality Center. Arch Pathol Lab Med. 2014;138:1432–43.
Francis DP, Coats AJ, Gibson DG. How high can a correlation coefficient be? Effects of limited reproducibility of common cardiological measures. Int J Cardiol. 1999;69:185–9.
Bland JM, Altman DG. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet. 1986;1:307–10.
Westgard JO. Statistical quality control procedures. Clin Lab Med. 2013;33:111–24.
Howat WJ, Lewis A, Jones P, Kampf C, Ponten F, van der Loos CM, et al. Antibody validation of immunohistochemistry for biomarker discovery: recommendations of a consortium of academic and pharmaceutical based histopathology researchers. Methods. 2014;70:34–8.
Scheel AH, Baenfer G, Baretton G, Dietel M, Diezko R, Henkel T, et al. Interlaboratory concordance of PD-L1 immunohistochemistry for non-small-cell lung cancer. Histopathology. 2018;72:449–59.
Schats KA, Van Vre EA, De Schepper S, Boeckx C, Schrijvers DM, Waelput W, et al. Validated programmed cell death ligand 1 immunohistochemistry assays (E1L3N and SP142) reveal similar immune cell staining patterns in melanoma when using the same sensitive detection system. Histopathology. 2017;70:253–63.
Acknowledgements
We would like to thank Lori Charette and the team at the Yale Pathology Tissue Service for production of the high-quality sections used in this study. We also thank Dr Emina Torlakovic for critical review of the manuscript. This work was supported by a research agreement from Astra Zeneca and support for the Yale SPORE in Lung Cancer (P50-CA196530) and the Yale Cancer Center (P30-CA016359).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
DC is employee of Horizon Discovery; LMS received honoraria from Foghorn Therapeutics and Astra Zeneca and travel support from Bristol Myers Squibb, and she is a consultant for LOXO Oncology; JQ is a US Pathology Advisory Board member for Merck & Co. Inc.; MM-K is a consultant for Merrimack Pharmaceuticals and H3 Biomedicine; JW is employee of Astra Zeneca; MBB has been a prior consultant for Merck and Genentech; HY is at the Speakers Bureau for the HER2 program and is a consultant for Genentech; in the past 12 months, DLR has served as a consultant for advisor to Astra Zeneca, Agendia, Agilent, Biocept, BMS, Cell Signaling Technology, Cepheid, Daiichi Sankyo, GSK, Merck, Konica/Minolta, PerkinElmer/Akoya, and Ultivue. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Martinez-Morilla, S., McGuire, J., Gaule, P. et al. Quantitative assessment of PD-L1 as an analyte in immunohistochemistry diagnostic assays using a standardized cell line tissue microarray. Lab Invest 100, 4–15 (2020). https://doi.org/10.1038/s41374-019-0295-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-019-0295-9
This article is cited by
-
A phase Ib trial of pembrolizumab plus paclitaxel or flat-dose capecitabine in 1st/2nd line metastatic triple-negative breast cancer
npj Breast Cancer (2023)
-
Standardization of PD-L1 immunohistochemistry
Modern Pathology (2022)
-
High-throughput analysis of tissue microarrays using automated desorption electrospray ionization mass spectrometry
Scientific Reports (2022)
-
Validation of E1L3N antibody for PD-L1 detection and prediction of pembrolizumab response in non-small-cell lung cancer
Communications Medicine (2022)
-
Incorporation of TILs in daily breast cancer care: how much evidence can we bear?
Virchows Archiv (2022)