Abstract
A previous study revealed that therapeutic miR-26a delivery suppresses tumorigenesis in a murine liver cancer model, whereas we found that forced miR-26a expression increased hepatocellular carcinoma (HCC) cell migration and invasion, which prompted us to characterize the causes and mechanisms underlying enhanced invasion due to ectopic miR-26a expression. Gain-of-function and loss-of-function experiments demonstrated that miR-26a promoted migration and invasion of BEL-7402 and HepG2 cells in vitro and positively modulated matrix metalloproteinase (MMP)-1, MMP-2, MMP-9, and MMP-10 expression. In addition, exogenous miR-26a expression significantly enhanced the metastatic ability of HepG2 cells in vivo. miR-26a negatively regulated in vitro proliferation of HCC cells, and miR-26a overexpression suppressed HepG2 cell tumor growth in nude mice. Further studies revealed that miR-26a inhibited cell growth by repressing the methyltransferase EZH2 and promoted cell migration and invasion by inhibiting the phosphatase PTEN. Furthermore, PTEN expression negatively correlated with miR-26a expression in HCC specimens from patients with and without metastasis. Thus, our findings suggest for the first time that miR-26a promotes invasion/metastasis by inhibiting PTEN and inhibits cell proliferation by repressing EZH2 in HCC. More importantly, our data also suggest caution if miR-26a is used as a target for cancer therapy in the future.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is highly malignant and has a very poor prognosis [1]. HCC is usually treated with surgery-based comprehensive therapies. The 5-year postoperative recurrence rate for this disease is greater than 60%, and the median 5-year survival rates after repeat hepatectomy, ablation and transarterial chemoembolization (TACE) were 35.2, 48.3, and 15.5 percent, respectively [2]. HCC diagnosis and treatment are limited by many factors, including difficulties in early diagnosis, high rates of recurrence and metastasis, lack of therapeutic specificity, and insufficient sources of innovative drugs and treatment methods [3, 4]. Therefore, the development of new therapies for HCC is particularly urgent.
Studies have shown that microRNAs (miRNAs) regulate the expression of more than 50% of all human coding genes [5]. miRNAs play important roles in a variety of physiological and pathological processes, including organ development [6], cell proliferation [7], cell differentiation [8, 9], cell apoptosis, and tumor development. Downregulation of oncogenic miRNA expression results in upregulation of genes with tumor suppressive functions. Conversely, upregulation of tumor suppressive miRNA expression can lead to downregulation of oncogene expression levels [10]. miRNAs regulate the expression of a large number of downstream genes as well as activation or inactivation of many signalling pathways. The ultimate effect of a miRNA is determined by the sum of its effects on all regulated genes [11]. Therefore, the biological functions of miRNAs are complex and diverse. The roles and mechanisms of action of miRNAs in various aspects of cancer development and progression need to be further elucidated.
miR-26a, which is 22 nucleotides in length, is located on human chromosome 3 and is widely expressed in various human tissues [12]. Previous studies have shown that miR-26a inhibits the proliferation of nasopharyngeal carcinoma cells, HCC cells and breast cancer cells [13,14,15]. However, miR-26a promotes the proliferation of glioma, ovarian cancer, and cholangiocarcinoma cells [16,17,18]. Additionally, miR-26a inhibits metastasis of nasopharyngeal carcinoma cells and gastric cancer cells [19, 20] but promotes metastasis of lung cancer cells [21]. Interestingly, miR-26a inhibits tumor proliferation in acute myeloid leukemia [22] but promotes tumor proliferation in T-cell acute lymphoblastic leukemia [23]. These findings indicate that miR-26a plays distinct roles in different tumors and in tumors of different histological types. Thus, the functional complexity of miR-26a is undeniable.
In a previous study, miR-26a was overexpressed in tumor-bearing mouse livers. The study found that miR-26a overexpression resulted in regression of liver tumors [14]. These results demonstrate the great potential of miR-26a in the treatment of HCC. However, it remains unclear whether the process of tumor regression is accompanied by reversal of HCC cell proliferation and migration. Therefore, we attempted to investigate the effects of miR-26a in HCC cells and its underlying mechanisms of action, thereby laying a theoretical foundation for application of miR-26a in clinical treatment of HCC. Previously, miR-26a has been reported to inhibit HCC cell proliferation. Our preliminary study found that miR-26a promotes HCC cell migration and invasion in vitro, prompting us to further investigate the role of miR-26a in HCC cell invasion and metastasis.
Materials and methods
Cell lines and cell culture
The human HCC cell lines BEL-7402 and HepG2 were purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). HEK293T cells were obtained from the American Type Culture Collection (ATCC). The cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (high glucose) supplemented with 10% fetal bovine serum (FBS), 1% non-essential amino acids (NEAA), 1 mM l-glutamine and 1 mM sodium pyruvate in a humidified incubator with 5% CO2 at 37 °C.
Lentivirus production and transduction
A 418-bp fragment containing the precursor sequence of human miR-26a (amplified by PCR from pMSCVpuro-miR-26a [15]) and a fragment containing a non-specific microRNA sequence was cloned into the Xba I and BamH I sites of pHAGE-fullEF1a-MCS-IZsGreen [generously provided by Prof. Jeng-Shin Lee (Harvard Gene Therapy Initiative, Harvard Medical School)], respectively. The resulting constructs were designated pLV-miR-26a and pLV-con and were confirmed by PCR, enzyme digestion analysis and sequencing (data not shown). The lentiviral packaging plasmids psPAX2 and pMD2.G were kindly provided by Dr. Didier Trono (University of Geneva, Geneva, Switzerland).
The production, purification, and titration of lentiviruses (LV-con and LV-miR-26a) were performed as described by Tiscornia and colleagues [24]. Briefly, to produce virus particles expressing the empty vector or target genes (miR-26a), HEK293T cells (maintained in 10% FBS) were co-transfected with lentiviral vector and packaging plasmids (psPAX2 and pMD2.G) using Lipofectamine 2000 reagent (Invitrogen) according to the manufacturer’s instructions. The packaged lentiviruses carrying pLV-con or pLV-miR-26a were named LV-con (used as a control) and LV-miR-26a, respectively. At 72 h post-transfection, the supernatant was harvested and subsequently used to infect BEL-7402 and HepG2 cells.
miRNA or siRNA transient transfection
Human miR-26a inhibitor (anti-miR-26a) and inhibitor control (a non-specific anti-miR control, anti-miR con) were purchased from RiboBio Co., Ltd. (Guangzhou, China). miR-26a-expressing cells were transiently transfected with anti-miR-26a (100 nM) and anti-miR con (100 nM) using Lipofectamine 2000 reagent (Invitrogen) according to the manufacturer’s instructions, respectively. miR-26a-expressing HepG2 cells were transiently transfected with pcDNA3.0 (used as blank vector), pcDNA3.0-PTEN or pcDNA3.0-EZH2 via using Lipofectamine 2000 reagent.
siRNA was transfected into cells at a final concentration of 140 nM by using Lipofectamine 2000 reagent as the manufacturer's instructions. The following siRNA was synthesized by RiboBio Co., Ltd. (Guangzhou, China) and used in the study: siEZH2 (Cat. No. siG09121182954): 5′-GCAAATTCTCGGTGTCAAA-3′; siPTEN(Cat. No. siB06817152516): 5′-GGCGCTATGTGTATTATTA-3′. Scrambled siRNA (sicontrol) was described previously [25].
RNA isolation, reverse transcription, and quantitative real-time PCR (qRT-PCR)
RNA isolation, reverse transcription, and qRT-PCR have been well described previously [26,27,28,29,30]. The primers used in the qRT-PCR assay are described below. GAPDH or U6 snRNA was used as an endogenous control. All samples were normalized to internal controls, and fold changes were calculated through relative quantification (2-△△Ct) [31].
For qRT-PCR quantification of protein-coding genes, the following primer pairs were used: EZH2-FP(FP: forward primer)(5′-GCCAGACTGGGAAGAAATCTG-3′) and EZH2-RP(RP: reverse primer) (5′-TGTGT-TGGAAAATCCAAGTCA-3′); PTEN-FP(5′-TTTGAAGACCATAACCCACCAC-3′) and PTEN-RP(5′-ATTACACCAGTTCGTCCCTTTC-3′); MMP1-FP(5′-AGCTAGCTCAGGATGACA-TTGATG-3′) and MMP1-RP(5′-GCCGATGGGCTGGACAG-3′); MMP2-FP(5′-TACAGGATCATTG GCTACACACC-3′) and MMP2-RP (5′-GGTCACATCGCTCCAGACT-3′); MMP9-FP(5′-GGGACGCAGACATCGTCATC-3′) and MMP9-RP(5′-TCGTCATCGTCGAAATGGGC-3′); MMP10-FP(5′-CAC-TCTACAACTCATTCACA GAGCT-3′) and MMP10-RP (5′-CTTGGATAACCTGCTTGTACCTCAT-3′); GAPDH-FP(5′-ACCCAG AAGACTGTGGATGG-3′) and GAPDH-RP(5′-TCTAGACGGCAGGTCAGG-TC-3′).
For qRT-PCR quantification of human miR-26a, the following primer pairs were used: U6 snRNA-RT primer (AACGCTTCACGAATTTGCGT), U6 snRNA-FP (CTCGCTTCGGCAGCACA) and U6 snRNA-RP (AACGCTTCACGAATTTGCGT); miR-26a-RT primer (GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAGCCT), miR-26a-FP (GACTGTTCAAGTAATCCAGGATA) and miR-26a-RP (GTGCAGGGTCCGAGGTATTC).
Western blot analysis
Protein lysates were separated on 10% SDS-PAGE gels and electrophoretically transferred to PVDF (polyvinylidene difluoride) membranes. Then, the blots were probed with primary antibodies against EZH2 (Cat. No. 612666, BD Biosciences), PTEN (Cat. No. 9188S, Cell Signaling Technology) or KLF4 (Cat. No. ab215036, Abcam), followed by incubation with horseradish peroxidase (HRP)-labeled secondary antibodies. Signals were detected using enhanced chemiluminescence (ECL). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as the protein loading control.
Wound-healing assay
Cells were seeded in a 6-well plate for 24 h. The cell monolayers were scratched with a 200-μl pipette tip and washed with medium without serum to remove the detached cells. The wounded areas were observed and imaged under a microscope.
Transwell migration assay and Boyden chamber invasion assay
Transwell migration and Boyden chamber invasion assays were performed as described previously [29, 30].
Metastasis analysis via tail vein injection assays
Four- to five-week-old female BALB/c nude mice were purchased from the Medical Laboratory Animal Center of Guangdong Province and maintained in microisolator cages under aseptic conditions. HepG2 cells were infected by lentivirus harboring luciferase (Luc) gene to generate the stable cell line (i.e., HepG2 cells) carrying Luc gene, designated Luc-HepG2 cell line. Vector- or miR-26a-expressing luc-HepG2 cells (6 × 106) were suspended in 200 μl of PBS and then injected into the tail vein of the mice (n = 14). Subsequently, luciferase (Luc) signals in nude mice were used to monitor metastatic tumors. When Luc signals of metastatic tumors were observed, the mice were sacrificed, and metastatic tumors in organs were analyzed through hematoxylin and eosin (H&E) staining. This investigation was carried out according to the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health (NIH Publication No. 85-23, revised 1996). All animal protocols were approved by the Institutional Animal Care and Use Committee (IACUC) at the Institute of Laboratory Animal Center, Southern Medical University.
Whole-animal bioluminescence imaging
The procedure for in vivo bioluminescence imaging using a Xenogen IVIS LuminaII Imaging System (Xenogen Corp., Alameda, CA, USA) to noninvasively measure Luc activity has been fully described previously [26, 29, 32,33,34].
CCK8 assay and colony formation assay
CCK8 and colony formation assays were performed as previously described [29]. For the CCK8 assay, the indicated cells were plated in 96-well plates at 1 × 103 per well in a final volume of 200 μl and then cultured for 7 days. For the colony formation assay, cells were counted and plated at 200 per well in 6-well plates for 14 days.
Cell cycle analysis
For cell cycle analysis, the indicated cells were plated in 6-well plates at 2 × 105 per well and transfected with miR-26a inhibitor or inhibitor control. At 48 h post-transfection, cell cycle distribution was analyzed via propidium iodide (PI) staining and flow cytometry as described by Singh and colleagues [35].
Tumor xenografts in animals
Female BALB/c nude mice aged 4–5 weeks were purchased from the Medical Laboratory Animal Center of Guangdong Province. Vector- or miR-26a-expressing HepG2 cells (1.5 × 106 cells) were subcutaneously injected into the left or right dorsal thigh of the mice (n = 10), respectively. Tumor volumes were measured every 2–3 days using a calliper slide rule. After 5 weeks, the mice were sacrificed, and tumor xenografts were dissected. Tumor volumes were calculated as follows: volume = (D × d2)/2, where D is the longest diameter and d is the shortest diameter. All animals were sacrificed on the 35th day after transplantation.
Histological analysis and immunohistochemistry (IHC)
Histological analysis and IHC were performed as described previously [27,28,29, 36]. For histological analysis, human tumor xenografts in nude mice were collected, fixed in 4% phosphate-buffered paraformaldehyde (PFA) at 4 °C overnight, embedded in paraffin, and then cut into 5-mm-thick sections. Subsequently, the tissue sections were mounted on slides, dewaxed, rehydrated, and then stained with H&E staining according to standard procedures.
The immunohistochemical staining procedure followed the standard streptavidin-peroxidase (SP) protocol. After deparaffinization and rehydration, the paraffin-embedded sections were subjected to high-pressure treatment in citrate buffer (pH 6.0) and boiling for 2 min for antigen retrieval. Endogenous peroxidase and non-specific staining were blocked with H2O2 and 1% BSA for 15 min at room temperature, respectively. The sections were then incubated overnight at 4 °C with primary antibodies against BrdU (GE Healthcare, dilution 1:50) or p21 (Abcam, dilution 1:100). PBS was used as the negative control. Immunocomplexes were visualized with DAB, and the tissue sections were counterstained with hematoxylin.
Clinical specimens
Primary HCC specimens and tumor-adjacent noncancerous tissues were collected from Sun Yat-sen Memorial Hospital, Sun Yat-sen University (Guangzhou, China) with informed consent and under institutional review board-approved protocols. The inclusion criteria for HCC cases were (1) a clear pathological diagnosis of HCC and (2) no anticancer treatments before surgery. Ethical approval was given by the Medical Ethics Committee of Southern Medical University, with the following reference number: 2017-002-01. Total RNA was extracted from the specimens and used for qRT-PCR analysis.
Statistical analyses
Statistical analyses were performed using the SPSS 13.0 software package. Two-tailed Student's t-test was used for comparisons of two independent groups. One-way ANOVA was used for comparisons of intergroup differences and differences within groups. The data are presented as the mean ± SEM. The “*” sign denotes P < 0.05 compared with the control, the “**” sign denotes P < 0.01 compared with the control, and the “***” sign denotes P < 0.001 compared with the control.
Results
miR-26a is downregulated in human HCC cell lines and clinical specimens
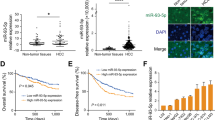
qRT-PCR was used to analyze miR-26a expression levels in human HCC cell lines and clinical specimens. The results showed that miR-26a expression was decreased in all 7 HCC cell lines examined compared with normal liver tissues (Fig. S1). We further examined miR-26a expression levels in 55 HCC specimens and 45 non-tumor liver tissues. Consistent with the data obtained from HCC cell lines, the average miR-26a expression level was significantly lower in the HCC specimens than in the non-tumor liver tissues (Fig. 7a).
miR-26a promotes HCC cell migration and invasion in vitro
To explore the effects of miR-26a on cell migration and invasion, BEL-7402 and HepG2 cells were infected with LV-con or LV-miR-26a (Fig. S2). Wound-healing assays demonstrated that miR-26a overexpression promoted both BEL-7402 and HepG2 cell migration (Fig. 1a). Transwell migration assays and Boyden chamber invasion assays showed that miR-26a overexpression enhanced BEL-7402 cell migration and invasion (Fig. 1b, c), while anti-miR-26a inhibited BEL-7402 cell migration and invasion (Fig. S3 and Fig. 1b, c). Similar results were obtained in HepG2 cells (Fig. S3 and Fig. 1d, e).
miR-26a positively modulated the migration and invasion ability of HCC cells in vitro. a The migration ability of miR-26a-expressing BEL-7402 and HepG2 cells was assessed with wound-healing assays. b–e In vitro migration assay using a transwell chamber and in vitro invasion assay using a Matrigel-coated Boyden chamber were employed to detect the motility and invasion of miR-26a-expressing BEL-7402 (b, c) and HepG2 (d, e) cells, and of BEL-7402 (b, c) and HepG2 (d, e) cells transfected with antimiR-26a inhibitor (100 nM) for 48 h
We then analyzed the expression levels of matrix metalloproteinase (MMP)-1, MMP-2, MMP-9, and MMP-10 using qRT-PCR and found that miR-26a overexpression resulted in the significantly elevated expression of all four MMPs in BEL-7402 and HepG2 cells (Fig. 2a, b). Conversely, inhibition of miR-26a expression led to the notably reduced expression of MMP-1, MMP-2, MMP-9, and MMP-10 expression in BEL-7402 and HepG2 cells (Fig. 2c, d).
qRT-PCR detection of metastasis-related gene expression. a, b MMP-1, MMP-2, MMP-9 and MMP-10 expression was upregulated in miR-26a-expressing BEL-7402 (a) and HepG2 (b) cells. c, d Reduced MMP-1, MMP-2, MMP-9 and MMP-10 expression was observed in BEL-7402 (c) and HepG2 (d) cells transfected with anti-miR-26a inhibitor (100 nM) for 48 h
miR-26a promotes HCC cell metastasis in vivo
To further confirm that miR-26a promotes metastasis in vivo, miR-26a-expressing HepG2-Luc cells were injected into mice via the tail vein. Bioluminescence imaging revealed (Luc) activity in the lungs of nude mice 3 h after tail vein injection of vector- or miR-26a-expressing HepG2-Luc cells (Fig. S4), indicating that the cancer cells were successfully injected. Within 3–4 months post-injection, tumor metastasis nodules in the lung were more numerous in the miR-26a-expressing group than in the control group. Most of the mice in the miR-26a overexpression group developed brain metastases (Fig. 3a, b), lung metastases (Fig. 3c, d), and lymph node metastases (Fig. S5). Histological sections of metastatic tumor foci were stained with H&E and analyzed to confirm the presence of brain, lung and lymph node metastases (Fig. 3e). The results can be summarized as follows: among the 7 mice in the miR-26a overexpression group, 1 developed brain metastases, 1 had brain and lymph node metastases, 4 had lung metastases, and 1 had liver and lymph node metastases. In the control group, only 2 mice developed lung metastases. No metastases were observed in any of the other vital organs examined (Fig. 3f and Fig. S6). These findings demonstrate that miR-26a promotes HCC cell metastasis in vivo.
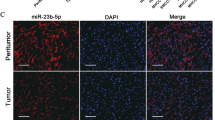
Enforced expression of miR-26a in Luc-HepG2 cells promoted metastasis. a Bioluminescence imaging of a representative nude mouse with brain metastases. b Representative brain metastatic tumour shown by the GFP and Luc signals. c Bioluminescence imaging of a representative nude mouse with lung metastases. d Representative lung metastatic tumours shown by the GFP and Luc signals. e H&E staining of brain, lung, liver and lymph node sections from nude mice 3-4 months after tail vein injection of miR-26a-expressing Luc-HepG2 cells. f The number of brain, lung, liver and lymph node metastatic nodules in nude mice
miR-26a negatively regulates HCC cell proliferation
Previous studies have reported that overexpression of miR-26a inhibits HCC cell proliferation. However, the present study found that overexpression of miR-26a promoted HCC cell invasion and metastasis. To verify the effect of miR-26a on proliferation of the cell lines examined above, Cell Counting Kit-8 (CCK8) assays, cell cycle analyses, and colony formation assays were performed. The CCK8 assay results showed that miR-26a overexpression resulted in a decrease in HCC cell proliferation (Fig. 4a), while inhibition of miR-26a overexpression restored HCC cell proliferative capacity (Fig. 4b). Cell cycle analyses demonstrated that overexpression of miR-26a led to G1-phase cell cycle arrest. In contrast, cells that underwent miR-26a overexpression-induced G1 arrest resumed proliferation following subsequent inhibition of miR-26a expression (Fig. 4c–f). Colony formation assays also demonstrated that miR-26 overexpression inhibited HCC cell proliferation (Fig. 4g. h). These results confirm that miR-26a negatively regulates HCC cell proliferation in vitro.
miR-26a negatively regulated the in vitro proliferation of HCC cells. a Effects of miR-26a on cell proliferation measured with CCK-8 assays in miR-26a-expressing BEL-7402 and HepG2 cells. b CCK-8 assay performed in BEL-7402 and HepG2 cells transfected with anti-miR-26a inhibitor. c-f Effects of miR-26a on cell cycle distribution of BEL-7402 cells (c, d) and HepG2 cells (e, f). g, h Colony formation assay revealing significantly decreased proliferation of miR-26a-expressing HCC cells
miR-26a overexpression suppresses tumor growth of HCC cells in vivo
To confirm the regulatory effect of miR-26a on HCC cell proliferation in vivo, subcutaneous tumor xenograft experiments were performed in nude mice. Significantly smaller tumors were observed in mice injected with miR-26-overexpressing HCC cells than in the mice injected with empty vector-transduced HCC cells (Fig. 5a, b). In addition, immunohistochemical analyses showed that tumors formed from the miR-26-overexpressing HCC cells contained a markedly reduced number of 5-bromo-2′-deoxyuridine (BrdU)-positive cells and a significantly increased number of p21-positive cells than tumors derived from empty vector-transduced HCC cells (Fig. 5c, d).
miR-26a suppressed HCC cell tumour growth in nude mice. Vector- and miR-26a-expressing HepG2 cells were injected subcutaneously into the left and right side of nude mice, respectively. At 2 weeks after implantation, LV-miR-26a-infected cells produced smaller tumours than control cells. a Representative picture of tumours formed. b Growth curve of tumour volumes. c, d BrdU- and p21-stained sections of transplanted tumours formed by HepG2 cells. The percentages of BrdU- or p21- positive cancer cells were calculated by determining the total number of BrdU- or p21-positive cells divided by the total number of cancer cells
miR-26a inhibits cell growth by repressing EZH2 and promotes migration and invasion by inhibiting PTEN in HCC cells
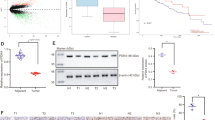
miR-26a promotes HCC cell invasion and metastasis but inhibits HCC cell proliferation. We suspected that miR-26a was able to exert such interesting effects by targeting different genes. Therefore, we explored the mechanisms underlying miR-26a activity. Bioinformatics software predicted that nucleotides 2–8 at the 5′ end of miR-26a were perfectly complementary to the 3′ untranslated regions (3’-UTRs) of enhancer of zeste homolog 2 (EZH2), phosphatase and tensin homolog (PTEN) and Kruppel-like factor 4 (KLF4) (data not shown). Moreover, this complementary region was highly conserved across multiple species (human, mouse, rat, and monkey) (Fig. 6a, b and Fig. S7a). Western blot analysis revealed that miR-26a overexpression resulted in downregulation of EZH2 and PTEN expression in BEL-7402 and HepG2 cells, while subsequent inhibition of miR-26a resulted in upregulation of EZH2 and PTEN expression (Fig. 6c). These results indicate that miR-26a negatively regulates EZH2 and PTEN expression. However, miR-26a did not negatively regulate KLF4 expression in BEL-7402 and HepG2 cells (Fig. S7b). Previous studies have reported that EZH2 is a direct target gene of miR-26a [13, 37]. Additionally, PTEN has also been reported to be a direct target gene of miR-26a [21, 38].
miR-26a promoted migration and invasion by inhibiting PTEN and inhibited cell growth by repressing EZH2 in HCC cells. a Sequence alignment of the 3' UTR of the mouse (Mmu), rat (Rno), human (Hsa) and rhesus (Mml) PTEN protein, highlighting the miR-26a binding site. b Sequence alignment of the 3' UTR of the mouse (Mmu), rat (Rno), human (Hsa) and rhesus (Mml) EZH2 protein, highlighting the miR-26a binding site. c miR-26a negatively regulated the expression of its target genes (i.e., EZH2 and PTEN) in HCC cells. d The protein levels of EZH2 and PTEN in HepG2 cells transfected with different plasmids determined by Western blot analysis. e miR-26a inhibited cell growth by repressing EZH2 but not PTEN in HepG2 cells, based on CCK-8 assays. f, g miR-26a promoted migration and invasion by inhibiting PTEN but not EZH2 in HepG2 cells. h The protein levels of EZH2 and PTEN in BEL-7402 and HepG2 cells transfected with the indicated siRNA (140 nM) determined by Western blot analysis. i, j CCK-8 assay performed in BEL-7402 (i) and HepG2 (j) cells transfected with the indicated siRNA (140 nM). k, l, m In vitro migration assay using a transwell chamber was employed to detect the motility of BEL-7402 (k, l) and HepG2 (k, m) cells transiently transfected with the indicated siRNA (140 nM)
To further elucidate whether miR-26a regulates HCC cell proliferation, migration, and invasion through EZH2 and PTEN, we carried out HepG2 cell-based functional analyses. EZH2 and PTEN were separately transiently overexpressed in miR-26a-overexpressing HepG2 cells (Fig. 6d). CCK8 assay results showed that stable overexpression of miR-26a significantly reduced the proliferative capacity of HepG2 cells, whereas transient overexpression of EZH2 in these cells resulted in an increase in cell proliferation (Fig. 6e). In contrast, transient overexpression of PTEN in miR-26a-overexpressing HepG2 cells had no effect on cell proliferation (Fig. 6e). The results of the Transwell and Boyden chamber assays showed that migration and invasion were significantly enhanced in miR-26a-overexpressing HepG2 cells compared with empty vector-transduced HepG2 cells (Fig. 6f, g). Transient overexpression of PTEN in miR-26a-overexpressing HepG2 cells reduced both cell migration and invasion (Fig. 6f, g). However, transient overexpression of EZH2 in miR-26a-overexpressing HepG2 cells did not affect the increased cell migration and invasion ability induced by miR-26a overexpression (Fig. 6f, g). In summary, miR-26a inhibits cell growth by repressing EZH2 and promotes migration and invasion by inhibiting PTEN in HCC cells.
To further examine the effects of PTEN or/and EZH2 downregulation in these HCC cells, BEL-7402 and HepG2 cells were separately or combinedly transfected with siEZH2 and siPTEN (Fig. 6h), respectively. CCK8 assay revealed that transient expression of siEZH2 or siEZH2 and siPTEN significantly decreased the proliferative ability of BEL-7402 (Fig. 6i) and HepG2 (Fig. 6j) cells, whereas transient expression of siPTEN had slight proliferation-promoting effect on BEL-7402 cells (Fig. 6i) and had no effect on HepG2 cell proliferation (Fig. 6j). Furthermore, there is no significant difference in cell proliferation-inhibition effect between HCC cells transfected with siEZH2 and HCC cells transfected with siEZH2 & siPTEN (Fig. 6i, j). The transwell assays showed that the migration was very significantly enhanced in siPTEN-transfected BEL-7402 cells (Fig. 6k, l) and HepG2 cells (Fig. 6k, m), whereas the transient expression of siEZH2 in BEL-7402 cells (Fig. 6k, l) and HepG2 cells (Fig. 6k, m) only led to a slight decrease in cell migration capacity. Therefore, our findings demonstrate that EZH2 and PTEN might play key roles in modulating the proliferation and motility ability of these examined HCC cells, respectively.
Confirmation of the relationship between miR-26a expression and PTEN and EZH2 expression levels using clinical HCC specimens
To confirm the relationship between miR-26a expression and PTEN or EZH2 expression levels in clinical HCC specimens, we obtained 55 HCC tissue specimens and 45 paracancerous tissue specimens. Total RNA was extracted from the specimens and reverse transcribed into complementary DNA (cDNA). Subsequently, qRT-PCR was performed to examine the expression levels of miR-26a, PTEN, and EZH2. The results demonstrated that miR-26a (Fig. 7a) and PTEN (Fig. 7c) expression levels were significantly downregulated in HCC tissues compared with paracancerous tissues, while EZH2 expression was significantly upregulated in HCC tissues (Fig. 7e). These data indicate that miR-26a, PTEN, and EZH2 play important roles in HCC malignancy. miR-26a expression was significantly upregulated in HCC specimens from patients with metastasis compared with HCC specimens from patients without metastasis (Fig. 7b), while PTEN expression was markedly downregulated in HCC specimens from patients with metastasis (Fig. 7d). These results demonstrate that PTEN expression is negatively correlated with miR-26a expression in HCC specimens from patients with and without metastasis. In addition, our results suggest that miR-26a might play an important role in HCC metastasis through regulation of PTEN expression. Moreover, we found that there was no significant difference in EZH2 expression in HCC specimens between patients with metastasis and patients without metastasis (Fig. 7f). Through examination of clinical tissue specimens, we obtained strong clinical evidence supporting our findings in the HCC cell lines. In both sample sets, we demonstrated that miR-26a inhibits HCC cell proliferation by downregulating EZH2 expression and promotes HCC cell migration and invasion by downregulating PTEN expression.
miR-26a, PTEN and EZH2 expression levels in HCC clinical specimens and their prognostic value in HCC patients. a, c, e Expression levels of mature miR-26a (a), PTEN (c) and EZH2 (e) in 55 HCC specimens and 45 adjacent non-tumour tissues. b, d, f Expression levels of mature miR-26a (b), PTEN (d) and EZH2 (f) in HCC specimens compared between patients with metastasis and patients without metastasis
In summary, we have reached the following conclusions: miR-26a inhibits HCC cell proliferation by downregulating EZH2 expression. Additionally, miR-26a promotes HCC cell migration and invasion by downregulating PTEN expression. Further, the upregulation of miR-26a expression leads to increased metastatic capacity and decreased proliferative capacity in HCC cells (Fig. 8). This miR-26a activity is likely to occur during invasion and metastasis of HCC cells.
Discussion
miRNAs can have oncogenic, tumor suppressive, or both roles depending on the type of tumor and target mRNA whose translation they suppress [39]. The expression levels and biological functions of a single miRNA may vary across various cancer types and subtypes and even between different cell lines of the same cancer, suggesting that a miRNA signature may be tissue- and cell-specific [39]. Additionally, miRNA levels of expression also vary during disease formation and propagation, suggesting the presence of a time profile for their expression [39].
Based on accumulated evidence, miR-26a is primarily known for its tumor suppressive functions in cancer cell proliferation, tumorigenesis, invasion, metastasis and resistance to radiotherapy and chemotherapy in numerous tumors, including HCC [14, 40,41,42,43,44,45], nasopharyngeal carcinoma [13, 19], bladder cancer [46], breast cancer [47, 48], pancreatic cancer [49], prostate cancer [50, 51], osteosarcoma [52], papillary thyroid carcinoma [53], squamous cell cancer [54], acute myeloid leukemia [22], tongue squamous cell carcinoma [55], and endometrial carcinoma [56]. Increasing evidence supports the notion that miR-26a suppresses HCC cell proliferation and tumorigenesis [14, 40, 45] and inhibits HCC cell growth and/or tumorigenesis by suppressing the expression of one or more of the following target genes: IL-6 [43], ST3GAL6 [42], hepatocyte growth factor (HGF) [44] or Lin28B and Zcchc11 [57], while miR-26a suppresses HCC cell migration, invasion and/or metastasis by targeting the interleukin-6-stat3 pathway [43], EZH2 and CDK8 in Wnt pathway [58], DNMT3B-MEG3 axis [59], HGF-cMet pathway [44], EZH2 [60], [42] ST3GAL6 or Lin28B and Zcchc11 [57]. Moreover, miR-26a/b can promote apoptosis and sensitize HCC to chemotherapy by suppressing the expression of autophagy initiator ULK1 [61]. miR-26a suppresses angiogenesis by targeting the HGF-cMet pathway in HCC [44]. These aforementioned findings support the tumor suppressive function of miR-26a in HCC. In this study, we found that forced miR-26a expression suppressed HCC cell proliferation by repressing EZH2 and suppressed HCC cell tumor growth in vivo, whereas miR-26a overexpression significantly promoted migration and invasion by inhibiting PTEN and enhanced the metastatic ability of HepG2 cells in vivo, which was further supported by analysis of HCC clinical specimens. The in vitro and in vivo HCC cell growth-suppressing role of miR-26a revealed in this study is consistent with the above-described findings in HCC [14, 40, 42,43,44,45, 57], whereas in vitro and in vivo cell migration-, invasion- and metastasis-promoting miR-26a activity has been shown to enhance the migration, invasion and metastasis potential of lung cancer cells by suppressing PTEN to activate the PTEN/AKT pathway [21], by regulating the ITGβ8-JAK2/STAT3 axis [62] or by suppressing GSK3β [63]. Moreover, miR-26a was found to be downregulated in lung cancer specimens compared with normal lung tissue, and interestingly, the miR-26a expression level was higher in lymph node metastasis tumor tissues than in primary tumor tissues [21]. Additionally, the expression level of PTEN, a target gene of miR-26a, was lower in lymph node metastasis tumor tissues than in primary tumor tissues and was inversely correlated with the miR-26a level [21]. Thus, the above findings in lung cancer [21] are consistent with our results in HCC. Moreover, miR-26a promoted ovarian cancer proliferation and tumorigenesis by targeting ER-α [17], whereas miR-26a inhibited ovarian cancer cell proliferation and induced cell apoptosis by regulating CDC6 expression [64]. Sustained expression of miR-26a promoted chromosomal instability and tumorigenesis through regulation of CHFR in breast cancer [65], whereas miR-26a suppressed tumor proliferation and metastasis by targeting metadherin in triple negative breast cancer [48], and miR-26a inhibited the proliferation and migration of breast cancer cells through repression of MCL-1 [47]. In summary, these findings clearly demonstrate that the biological functions of miR-26a vary across different cancer types and subtypes and even between different cell lines of the same cancer, suggesting that the biological roles of miR-26a in the progression of various malignancies and the underlying mechanisms are very complex. Furthermore, miR-26a plays diametrically opposing biological roles in cell proliferation, apoptosis, tumorigenesis, invasion, and metastasis by regulating different target genes in various cancers.
An oncomiRNA selectively represses the expression of its tumor suppressor target genes in tumorigenesis but does not affect the expression of oncogenic potential target genes; similarly, a tumor suppressor miRNA selectively targets oncogene(s) but not tumor suppressor(s) in normal development or cell differentiation [66,67,68]. Thus, most researchers follow the aforementioned logic when trying to identify candidate target genes of a given miRNA. A single miRNA is well known to have tens or hundreds of putative target genes, including oncogenes and tumor suppressors [69]. Therefore, it might not be practical for a given oncogenic or tumor suppressor miRNA to selectively repress only the expression of targets with functions opposite to those proposed to be exerted by the miRNA, and this idea is strongly supported by the following evidence. miR-196b is involved in MLL-fusion-mediated leukemogenesis by simultaneously repressing the expression of both oncogenic (HOXA9/MEIS1) and tumor suppressor (FAS) target genes [70]. By providing compelling evidence, here, we demonstrate that miR-26a promoted cell migration and invasion by inhibiting the tumor suppressor PTEN and inhibited cell proliferation by repressing the oncogene EZH2 in HCC, and this was supported by analysis of HCC clinical specimens. The information obtained from HCC clinical specimens revealed that miR-26a was downregulated in HCC specimens compared with tumor-adjacent noncancerous tissues, and interestingly, miR-26a expression was higher in HCC specimens from patients with metastasis than in HCC specimens from patients without metastasis, while the expression level of PTEN, a target gene of miR-26a, was lower in HCC specimens from patients with metastasis than in HCC specimens from patients without metastasis. Collectively, the ability of a single miRNA to target both oncogenes and tumor suppressors simultaneously, or sequentially, is quite important for cancer malignant progression. More importantly, we should update our present concepts about miRNA regulation in normal development, cell differentiation, and tumorigenesis and refine our definition of oncogenic miRNA and tumor suppressor miRNA.
Interestingly, we found that miR-26a-overexpressing HCC cells exhibited reduced cell proliferation, and increased cell migration, invasion and metastasis ability. Thus, it seems that the migration, invasion or metastasis promotion roles of miR-26a in BEL-7402 and HepG2 cells (Figs. 1–3) is inconsistent with its tumor suppressive role [14, 40, 42,43,44,45, 60]. Similar phenomena are observed with other miRNAs and protein-coding genes. miR-200c upregulation inhibited pancreatic cancer invasion but increased cell proliferation by targeting E-cadherin [71]. miR-19 triggered epithelial–mesenchymal transition (EMT) and then promoted migration and invasion of lung cancer cells, accompanied by reduced proliferation of cells [72]. Forced expression of Y-box binding protein-1 (YB-1) in noninvasive breast epithelial cells induced EMT, accompanied by enhanced metastatic potential and reduced cell proliferation rates [73]. Bone morphogenetic protein 4 (BMP4) inhibited breast cancer cell proliferation and induced an MMP-dependent migratory phenotype by targeting and regulating different target genes [74,75,76]. EMT has been reported to contribute to tumor cell invasion and migration and to metastatic dissemination [73, 77, 78]. Tumor cells undergoing EMT are typically found at the edge of tumors, account for a small proportion of the primary tumor and have low proliferation rates. Strong evidence suggests that reduced proliferation cooperates with EMT to drive cancer cells to reach their new destination before efficient proliferation [73, 79,80,81,82,83]. In addition, the products of EMT-induced genes, such as Snail1 [81], and ZEB2/SIP1 [83], exhibited strong antiproliferative activity by inhibiting cell cycle progression in tumor cells undergoing EMT, which is consistent with the abovementioned results. Previous studies [73, 80,81,82,83] and our results (Fig. 8) suggest that enhanced cell proliferation is important for initiation and maintenance of primary tumors, while growth inhibition is crucial for invasion and migration of tumor cells. Collectively, our results suggest that downregulation of tumor suppressive miR-26a can promote tumor cell proliferation during initiation and maintenance of primary tumors and upregulation of tumor suppressive miR-26a in HCC specimens from patients with metastasis inhibits cell proliferation by repressing EZH2 and enhances tumor invasion and metastasis by inhibiting PTEN expression (Fig. 8), suggesting functional transition of miR-26a at different stages of tumor development.
A single miRNA can regulate a large number of target protein-coding genes involved in different signal transduction pathways that participate in many physiological and pathological processes, including tumor formation and progression [84,85,86,87,88]. Therefore, certain miRNAs are being considered as potential therapeutic targets for various diseases, including hepatitis and cancers. Several miRNA mimics and molecules that target miRNAs (anti-miRs) have shown promise for clinical application in preclinical or clinical trials [89, 90], and miRNA-targeted therapeutics have already been tested in clinical trials, including a mimic of the tumor suppressor miR-34, which reached phase I clinical trials for cancer treatment [89] and anti-miRs targeting miR-122, which reached phase II trials for hepatitis treatment [91]. Systemic administration of miR-26a in a mouse model of HCC using adeno-associated virus (AAV) resulted in nearly complete regression of liver tumors, inhibition of cancer cell proliferation, induction of tumor-specific apoptosis and dramatic protection from disease progression without toxicity [14]. However, these data along with the above discussion suggest that the possibility of distant organ metastasis triggered by AAV-mediated miR-26a delivery cannot be ruled out in an HCC mouse model after long-term follow-up observation. Therefore, our data call for caution if investigators plan to target miR-26a for cancer therapy in the future, and further studies on the multiple functions of the miR-26a should be performed before clinical application is pursued.
In conclusion, here, we report a previously unappreciated miR-26a-mediated regulatory model in which miR-26a can promote invasion/metastasis by inhibiting the tumor suppressor target gene PTEN and inhibit cell proliferation by repressing the oncogenic target gene EZH2 in HCC, indicating that miRNA regulation is much more complex than previously thought. Although therapeutic miR-26a delivery significantly suppressed tumorigenesis in a murine liver cancer model [14], further study of the multiple functions and dichotomous roles of miR-26a are warranted before it can be claimed relevant as a potential agent or target for HCC therapy.
References
Akinyemiju T, Abera S, Ahmed M, Alam N, Alemayohu MA, Allen C, et al. The burden of primary liver cancer and underlying etiologies from 1990 to 2015 at the global, regional, and national level: Results From the Global Burden of Disease Study 2015. JAMA Oncol. 2017;3:1683–91.
Erridge S, Pucher PH. Meta-analysis of determinants of survival following treatment of recurrent hepatocellular carcinoma. Br J Surg. 2017;104:1433–42.
Ni W, Zhang Y, Zhan Z, Ye F, Liang Y, Huang J, et al. A novel lncRNA uc.134 represses hepatocellular carcinoma progression by inhibiting CUL4A-mediated ubiquitination of LATS1. J Hematol Oncol. 2017;10:91.
Li M, Zhang M, Zhang ZL, Liu N, Han XY, Liu QC, et al. Induction of apoptosis by berberine in hepatocellular carcinoma HepG2 cells via downregulation of NF-kappaB. Oncol Res. 2017;25:233–9.
Leclercq M, Diallo AB, Blanchette M. Prediction of human miRNA target genes using computationally reconstructed ancestral mammalian sequences. Nucleic Acids Res. 2017;45:556–66.
Schratt GM, Tuebing F, Nigh EA, Kane CG, Sabatini ME, Kiebler M, et al. A brain-specific microRNA regulates dendritic spine development. Nature. 2006;439:283–9.
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell. 2003;113:25–36.
Chen CZ, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentiation. Science. 2004;303:83–86.
Card DA, Hebbar PB, Li L, Trotter KW, Komatsu Y, Mishina Y, et al. Oct4/Sox2-regulated miR-302 targets cyclin D1 in human embryonic stem cells. Mol Cell Biol. 2008;28:6426–38.
Chuang JC, Jones PA. Epigenetics and microRNAs. Pediatr Res. 2007;61:24r–29r.
Mishra PJ, Merlino G. MicroRNA reexpression as differentiation therapy in cancer. J Clin Invest. 2009;119:2119–23.
Chen J, Zhang K, Xu Y, Gao Y, Li C, Wang R, et al. The role of microRNA-26a in human cancer progression and clinical application. Tumour Biol. 2016;37:7095–108.
Lu J, He ML, Wang L, Chen Y, Liu X, Dong Q, et al. MiR-26a inhibits cell growth and tumorigenesis of nasopharyngeal carcinoma through repression of EZH2. Cancer Res. 2011;71:225–33.
Kota J, Chivukula RR, O’Donnell KA, Wentzel EA, Montgomery CL, Hwang HW, et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137:1005–17.
Zhang B, Liu XX, He JR, Zhou CX, Guo M, He M, et al. Pathologically decreased miR-26a antagonizes apoptosis and facilitates carcinogenesis by targeting MTDH and EZH2 in breast cancer. Carcinogenesis. 2011;32:2–9.
Huse JT, Brennan C, Hambardzumyan D, Wee B, Pena J, Rouhanifard SH, et al. The PTEN-regulating microRNA miR-26a is amplified in high-grade glioma and facilitates gliomagenesis in vivo. Genes Dev. 2009;23:1327–37.
Shen W, Song M, Liu J, Qiu G, Li T, Hu Y, et al. MiR-26a promotes ovarian cancer proliferation and tumorigenesis. PLoS ONE. 2014;9:e86871.
Zhang J, Han C, Wu T. MicroRNA-26a promotes cholangiocarcinoma growth by activating beta-catenin. Gastroenterology. 2012;143:246–56. e248
Yu L, Lu J, Zhang B, Liu X, Wang L, Li SY, et al. miR-26a inhibits invasion and metastasis of nasopharyngeal cancer by targeting EZH2. Oncol Lett. 2013;5:1223–8.
Deng M, Tang HL, Lu XH, Liu MY, Lu XM, Gu YX, et al. miR-26a suppresses tumor growth and metastasis by targeting FGF9 in gastric cancer. PLoS ONE. 2013;8:e72662.
Liu B, Wu X, Liu B, Wang C, Liu Y, Zhou Q, et al. MiR-26a enhances metastasis potential of lung cancer cells via AKT pathway by targeting PTEN. Biochim Biophys Acta. 2012;1822:1692–704.
Salvatori B, Iosue I, Djodji Damas N, Mangiavacchi A, Chiaretti S, Messina M, et al. Critical role of c-Myc in acute myeloid leukemia involving direct regulation of miR-26a and histone methyltransferase EZH2. Genes Cancer. 2011;2:585–92.
Mavrakis KJ, Van Der Meulen J, Wolfe AL, Liu X, Mets E, Taghon T, et al. A cooperative microRNA-tumor suppressor gene network in acute T-cell lymphoblastic leukemia (T-ALL). Nat Genet. 2011;43:673–8.
Tiscornia G, Singer O, Verma IM. Production and purification of lentiviral vectors. Nat Protoc. 2006;1:241–5.
Guo Y, Li D, Zhang S, Yang Y, Liu JJ, Wang X, et al. Visualizing intracellular organelle and cytoskeletal interactions at nanoscale resolution on millisecond timescales. Cell. 2018;175:1430–42. e1417
Lin X, Jia J, Du T, Li W, Wang X, Wei J, et al. Overexpression of miR-155 in the liver of transgenic mice alters the expression profiling of hepatic genes associated with lipid metabolism. PLoS ONE. 2015;10:e0118417.
Lin X, Qin Y, Jia J, Lin T, Lin X, Chen L, et al. MiR-155 enhances insulin sensitivity by coordinated regulation of multiple genes in Mice. PLoS Genet. 2016;12:e1006308.
Lin X, Chen L, Li H, Liu Y, Guan Y, Li X, et al. miR-155 accelerates proliferation of mouse hepatocytes during liver regeneration by directly targeting SOCS1. Am J Physiol Gastrointest Liver Physiol. 2018;315:G443–g453.
Rong XX, Wei F, Lin XL, Qin YJ, Chen L, Wang HY, et al. Recognition and killing of cancer stem-like cell population in hepatocellular carcinoma cells by cytokine-induced killer cells via NKG2d-ligands recognition. Oncoimmunology. 2016;5:e1086060.
Wang SC, Lin XL, Li J, Zhang TT, Wang HY, Shi JW, et al. MicroRNA-122 triggers mesenchymal-epithelial transition and suppresses hepatocellular carcinoma cell motility and invasion by targeting RhoA. PLoS ONE. 2014;9:e101330.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Du T, Jia J, Lin X, Xie R, Li J, Xiao D, et al. Generation of Rm21LG transgenic mice: a powerful tool to generate conditional overexpression of miR-21 that is involved in oncogenesis. Biotechnol Lett. 2014;36:9–20.
Lin X, Jia J, Qin Y, Lin X, Li W, Xiao G, et al. Simple and rapid determination of homozygous transgenic mice via in vivo fluorescence imaging. Oncotarget. 2015;6:39073–87.
Wei F, Rong XX, Xie RY, Jia LT, Wang HY, Qin YJ, et al. Cytokine-induced killer cells efficiently kill stem-like cancer cells of nasopharyngeal carcinoma via the NKG2D-ligands recognition. Oncotarget. 2015;6:35023–39.
Singh SV, Herman-Antosiewicz A, Singh AV, Lew KL, Srivastava SK, Kamath R, et al. Sulforaphane-induced G2/M phase cell cycle arrest involves checkpoint kinase 2-mediated phosphorylation of cell division cycle 25C. J Biol Chem. 2004;279:25813–22.
Lin X, Zhao W, Jia J, Lin T, Xiao G, Wang S, et al. Ectopic expression of Cripto-1 in transgenic mouse embryos causes hemorrhages, fatal cardiac defects and embryonic lethality. Sci Rep. 2016;6:34501.
Sander S, Bullinger L, Klapproth K, Fiedler K, Kestler HA, Barth TF, et al. MYC stimulates EZH2 expression by repression of its negative regulator miR-26a. Blood. 2008;112:4202–12.
Kim H, Huang W, Jiang X, Pennicooke B, Park PJ, Johnson MD. Integrative genome analysis reveals an oncomir/oncogene cluster regulating glioblastoma survivorship. Proc Natl Acad Sci USA 2010;107:2183–8.
Petrovic N, Ergun S, Isenovic ER. Levels of MicroRNA Heterogeneity in Cancer Biology. Mol Diagn Ther. 2017;21:511–23.
Chen L, Zheng J, Zhang Y, Yang L, Wang J, Ni J, et al. Tumor-specific expression of microRNA-26a suppresses human hepatocellular carcinoma growth via cyclin-dependent and -independent pathways. Mol Ther. 2011;19:1521–8.
Ma Y, Deng F, Li P, Chen G, Tao Y, Wang H. The tumor suppressive miR-26a regulation of FBXO11 inhibits proliferation, migration and invasion of hepatocellular carcinoma cells. Biomed Pharmacother. 2018;101:648–55.
Sun M, Zhao X, Liang L, Pan X, Lv H, Zhao Y. Sialyltransferase ST3GAL6 mediates the effect of microRNA-26a on cell growth, migration, and invasion in hepatocellular carcinoma through the protein kinase B/mammalian target of rapamycin pathway. Cancer Sci. 2017;108:267–76.
Yang X, Liang L, Zhang XF, Jia HL, Qin Y, Zhu XC, et al. MicroRNA-26a suppresses tumor growth and metastasis of human hepatocellular carcinoma by targeting interleukin-6-Stat3 pathway. Hepatology. 2013;58:158–70.
Yang X, Zhang XF, Lu X, Jia HL, Liang L, Dong QZ, et al. MicroRNA-26a suppresses angiogenesis in human hepatocellular carcinoma by targeting hepatocyte growth factor-cMet pathway. Hepatology. 2014;59:1874–85.
Zhu Y, Lu Y, Zhang Q, Liu JJ, Li TJ, Yang JR, et al. MicroRNA-26a/b and their host genes cooperate to inhibit the G1/S transition by activating the pRb protein. Nucleic Acids Res. 2012;40:4615–25.
Lin Y, Chen H, Hu Z, Mao Y, Xu X, Zhu Y, et al. miR-26a inhibits proliferation and motility in bladder cancer by targeting HMGA1. FEBS Lett. 2013;587:2467–73.
Gao J, Li L, Wu M, Liu M, Xie X, Guo J, et al. MiR-26a inhibits proliferation and migration of breast cancer through repression of MCL-1. PLoS ONE. 2013;8:e65138.
Liu P, Tang H, Chen B, He Z, Deng M, Wu M, et al. miR-26a suppresses tumour proliferation and metastasis by targeting metadherin in triple negative breast cancer. Cancer Lett. 2015;357:384–92.
Deng J, He M, Chen L, Chen C, Zheng J, Cai Z. The loss of miR-26a-mediated post-transcriptional regulation of cyclin E2 in pancreatic cancer cell proliferation and decreased patient survival. PLoS ONE. 2013;8:e76450.
Guo K, Zheng S, Xu Y, Xu A, Chen B, Wen Y. Loss of miR-26a-5p promotes proliferation, migration, and invasion in prostate cancer through negatively regulating SERBP1. Tumour Biol. 2016;37:12843–54.
Zhao S, Ye X, Xiao L, Lian X, Feng Y, Li F, et al. MiR-26a inhibits prostate cancer progression by repression of Wnt5a. Tumour Biol. 2014;35:9725–33.
Lu J, Song G, Tang Q, Yin J, Zou C, Zhao Z, et al. MiR-26a inhibits stem cell-like phenotype and tumor growth of osteosarcoma by targeting Jagged1. Oncogene. 2017;36:231–41.
Lv M, Zhang X, Li M, Chen Q, Ye M, Liang W, et al. miR-26a and its target CKS2 modulate cell growth and tumorigenesis of papillary thyroid carcinoma. PLoS ONE. 2013;8:e67591.
Shao Y, Li P, Zhu ST, Yue JP, Ji XJ, Ma D, et al. MiR-26a and miR-144 inhibit proliferation and metastasis of esophageal squamous cell cancer by inhibiting cyclooxygenase-2. Oncotarget. 2016;7:15173–86.
Jia LF, Wei SB, Gan YH, Guo Y, Gong K, Mitchelson K, et al. Expression, regulation and roles of miR-26a and MEG3 in tongue squamous cell carcinoma. Int J Cancer. 2014;135:2282–93.
Jiang FZ, He YY, Wang HH, Zhang HL, Zhang J, Yan XF, et al. Mutant p53 induces EZH2 expression and promotes epithelial-mesenchymal transition by disrupting p68-Drosha complex assembly and attenuating miR-26a processing. Oncotarget. 2015;6:44660–74.
Fu X, Meng Z, Liang W, Tian Y, Wang X, Han W, et al. miR-26a enhances miRNA biogenesis by targeting Lin28B and Zcchc11 to suppress tumor growth and metastasis. Oncogene. 2014;33:4296–306.
Zhang X, Zhang X, Wang T, Wang L, Tan Z, Wei W, et al. MicroRNA-26a is a key regulon that inhibits progression and metastasis of c-Myc/EZH2 double high advanced hepatocellular carcinoma. Cancer Lett. 2018;426:98–108.
Li Y, Ren M, Zhao Y, Lu X, Wang M, Hu J, et al. MicroRNA-26a inhibits proliferation and metastasis of human hepatocellular carcinoma by regulating DNMT3B-MEG3 axis. Oncol Rep. 2017;37:3527–35.
Ma DN, Chai ZT, Zhu XD, Zhang N, Zhan DH, Ye BG, et al. MicroRNA-26a suppresses epithelial-mesenchymal transition in human hepatocellular carcinoma by repressing enhancer of zeste homolog 2. J Hematol Oncol. 2016;9:1.
Jin F, Wang Y, Li M, Zhu Y, Liang H, Wang C, et al. MiR-26 enhances chemosensitivity and promotes apoptosis of hepatocellular carcinoma cells through inhibiting autophagy. Cell Death Dis. 2017;8:e2540.
Song Q, Liu B, Li X, Zhang Q, Cao L, Xu M, et al. MiR-26a-5p potentiates metastasis of human lung cancer cells by regulating ITGbeta8- JAK2/STAT3 axis. Biochem Biophys Res Commun. 2018;501:494–500.
Lin G, Liu B, Meng Z, Liu Y, Li X, Wu X, et al. MiR-26a enhances invasive capacity by suppressing GSK3beta in human lung cancer cells. Exp Cell Res. 2017;352:364–74.
Sun TY, Xie HJ, He H, Li Z, Kong LF. miR-26a inhibits the proliferation of ovarian cancer cells via regulating CDC6 expression. Am J Transl Res. 2016;8:1037–46.
Castellano L, Dabrowska A, Pellegrino L, Ottaviani S, Cathcart P, Frampton AE, et al. Sustained expression of miR-26a promotes chromosomal instability and tumorigenesis through regulation of CHFR. Nucleic Acids Res. 2017;45:4401–12.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Guo H, Ingolia NT, Weissman JS, Bartel DP. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature. 2010;466:835–40.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105.
Li Z, Huang H, Chen P, He M, Li Y, Arnovitz S, et al. miR-196b directly targets both HOXA9/MEIS1 oncogenes and FAS tumour suppressor in MLL-rearranged leukaemia. Nat Commun. 2012;3:688.
Yu J, Ohuchida K, Mizumoto K, Sato N, Kayashima T, Fujita H, et al. MicroRNA, hsa-miR-200c, is an independent prognostic factor in pancreatic cancer and its upregulation inhibits pancreatic cancer invasion but increases cell proliferation. Mol Cancer. 2010;9:169
Li J, Yang S, Yan W, Yang J, Qin YJ, Lin XL, et al. MicroRNA-19 triggers epithelial-mesenchymal transition of lung cancer cells accompanied by growth inhibition. Lab Invest. 2015;95:1056–70.
Evdokimova V, Tognon C, Ng T, Sorensen PH. Reduced proliferation and enhanced migration: two sides of the same coin? Molecular mechanisms of metastatic progression by YB-1. Cell Cycle. 2009;8:2901–6.
Ampuja M, Rantapero T, Rodriguez-Martinez A, Palmroth M, Alarmo EL, Nykter M, et al. Integrated RNA-seq and DNase-seq analyses identify phenotype-specific BMP4 signaling in breast cancer. BMC Genomics. 2017;18:68.
Ampuja M, Jokimaki R, Juuti-Uusitalo K, Rodriguez-Martinez A, Alarmo EL, Kallioniemi A. BMP4 inhibits the proliferation of breast cancer cells and induces an MMP-dependent migratory phenotype in MDA-MB-231 cells in 3D environment. BMC Cancer. 2013;13:429.
Ketolainen JM, Alarmo EL, Tuominen VJ, Kallioniemi A. Parallel inhibition of cell growth and induction of cell migration and invasion in breast cancer cells by bone morphogenetic protein 4. Breast Cancer Res Treat. 2010;124:377–86.
Thiery JP, Sleeman JP. Complex networks orchestrate epithelial-mesenchymal transitions. Nat Rev Mol Cell Biol. 2006;7:131–42.
Polyak K, Weinberg RA. Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer. 2009;9:265–73.
Barrallo-Gimeno A, Nieto MA. The Snail genes as inducers of cell movement and survival: implications in development and cancer. Development. 2005;132:3151–61.
Browne G, Sayan AE, Tulchinsky E. ZEB proteins link cell motility with cell cycle control and cell survival in cancer. Cell Cycle. 2010;9:886–91.
Vega S, Morales AV, Ocana OH, Valdes F, Fabregat I, Nieto MA. Snail blocks the cell cycle and confers resistance to cell death. Genes Dev. 2004;18:1131–43.
Evdokimova V, Tognon C, Ng T, Ruzanov P, Melnyk N, Fink D, et al. Translational activation of snail1 and other developmentally regulated transcription factors by YB-1 promotes an epithelial-mesenchymal transition. Cancer Cell. 2009;15:402–15.
Mejlvang J, Kriajevska M, Vandewalle C, Chernova T, Sayan AE, Berx G, et al. Direct repression of cyclin D1 by SIP1 attenuates cell cycle progression in cells undergoing an epithelial mesenchymal transition. Mol Biol Cell. 2007;18:4615–24.
Biswas S. MicroRNAs as therapeutic agents: The future of the battle against cancer. Curr Top Med Chem. 2018;18:2544–54.
Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10:704–14.
Garzon R, Calin GA, Croce CM. MicroRNAs in Cancer. Annu Rev Med. 2009;60:167–79.
Nadeem A, Ashraf MR, Javed M, Hussain T, Tariq MS, Babar ME. Review - MicroRNAs: A new paradigm towards mechanistic insight of diseases. Pak J Pharm Sci. 2018;31:2017–26.
Saliminejad K, Khorram Khorshid HR, Soleymani Fard S, Ghaffari SH. An overview of microRNAs: Biology, functions, therapeutics, and analysis methods. J Cell Physiol. 2019;234:5451–65.
Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov. 2013;12:847–65.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16:203–22.
Janssen HL, Reesink HW, Lawitz EJ, Zeuzem S, Rodriguez-Torres M, Patel K, et al. Treatment of HCV infection by targeting microRNA. N Engl J Med. 2013;368:1685–94.
Acknowledgements
We thank Prof. Qian Zhao (School of Medicine, Jiao-Tong University) and Prof. Jeng-Shin Lee (Harvard Gene Therapy Initiative, Harvard Medical School) for generously providing plasmids. This work was supported by the National Natural Science Foundation of China (Grant No. 81872209, 81672689, 81372896, and 81172587, to D. Xiao; Grant No. 81600086 and 81770100, to Y. Sun; Grant No. 81600488 and 81870602, to X.-L. Lin; Grant No. 81702778, to J.S. Jia), the Natural Science Foundation of Guangdong Province of China (Grant No. 2014A030313294 to D. Xiao), the Science and Technology Planning Project of Guangdong Province of China (Grant No. 2009B060300008, 2013B060300013 and 2017A010105017, to D. Xiao; Grant No. 2017A030303018, to J.S. Jia; Grant No. 2015A030302024, to X.-L. Lin), the Applied Basic Research Foundation of Yunnan Province Science and Technology Department & Kunming Medical University of China (Grant No. 2018FE001(-249), to W.T. Zhao), the China Postdoctoral Science Foundation(Grant No. 2015M572338, 2016T90792, 2017M622740 and 2018T110884, to X.-L. Lin) and the Medical Scientific Research Foundation of Guangdong Province of China (Grant No. A2017420, to J.S. Jia).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhao, WT., Lin, XL., Liu, Y. et al. miR-26a promotes hepatocellular carcinoma invasion and metastasis by inhibiting PTEN and inhibits cell growth by repressing EZH2. Lab Invest 99, 1484–1500 (2019). https://doi.org/10.1038/s41374-019-0270-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-019-0270-5
This article is cited by
-
ThermomiR-377-3p-induced suppression of Cirbp expression is required for effective elimination of cancer cells and cancer stem-like cells by hyperthermia
Journal of Experimental & Clinical Cancer Research (2024)
-
Synthetic miR-26a mimics delivered by tumor exosomes repress hepatocellular carcinoma through downregulating lymphoid enhancer factor 1
Hepatology International (2023)
-
The long and short non-coding RNAs modulating EZH2 signaling in cancer
Journal of Hematology & Oncology (2022)
-
Cordycepin inhibits colon cancer proliferation by suppressing MYC expression
BMC Pharmacology and Toxicology (2022)
-
Lineage tracing: technology tool for exploring the development, regeneration, and disease of the digestive system
Stem Cell Research & Therapy (2020)