Abstract
In recent years, the relevance of RNA metabolism has been increasingly recognized in a variety of diseases. Modifications in the levels of RNA-binding proteins elicit changes in the expression of cancer-related genes. Here we evaluate whether SRSF1 regulates the expression of DNA repair genes, and whether this regulation has a relevant role in lung carcinogenesis. An in silico analysis was performed to evaluate the association between the expression of SRSF1 and DNA repair genes. In vitro functional analyses were conducted in SRSF1 or DNA ligase 1 (LIG1)-downregulated non-small cell lung cancer (NSCLC) cell lines. In addition, the prognostic value of LIG1 was evaluated in NSCLC patients by immunohistochemistry. We found a significant correlation between the DNA repair gene LIG1 and SRSF1 in NSCLC cell lines. Moreover, SRSF1 binds to LIG1 mRNA and regulates its expression by increasing its mRNA stability and enhancing its translation in an mTOR-dependent manner. Furthermore, siRNA-mediated LIG1 inhibition reduced proliferation and increased apoptosis of NSCLC cells. Finally, the expression of LIG1 was an independent prognostic factor for NSCLC, as confirmed in a series of 210 patients. These results show that LIG1 is regulated by the oncoprotein SRSF1 and plays a relevant role in lung cancer cell proliferation and progression. LIG1 is associated with poor prognosis in non-small lung cancer patients.
Similar content being viewed by others
Introduction
RNA-binding proteins (RBPs) play critical roles in human pathogenesis. After transcription, nascent mRNAs undergo different processing steps including pre-mRNA splicing, transport, surveillance, localization, stability, and translation [1]. Thus, aberrant expression of RBPs profoundly affects RNA metabolism, modifying the function of target genes and promoting the development of different diseases, including cancer [2,3,4].
One of the best studied RBPs is serine/arginine-rich splicing factor 1 (SRSF1), formerly known as SF2, ASF, or SF2/ASF. SRSF1 is involved in splicing and other mRNA-dependent and independent processes (reviewed in ref. [5]). Moreover, SRSF1 was identified as a proto-oncogene with roles in both the establishment and maintenance of transformation [6]. We and others have shown that alterations in the levels of this protein accelerate lung cancer progression [7,8,9]. Several hypotheses have been suggested to explain the mechanisms by which SRSF1 promotes tumor development. Firstly, changes in the alternative splicing of some cancer-related mRNAs were proposed to be partially responsible for cell transformation upon SRSF1 overexpression. Specifically, SRSF1 overexpression promoted the appearance of oncogenic isoforms of several genes (BIN1, BCL2L11, MCL1, RON, MKNK2, S6K1, and PRRC2C), affecting proliferation, cell cycle progression, apoptosis, cell motility, and invasion [5, 8, 10,11,12]. Apart from splicing-related mechanisms, alterations in translation through mTOR activation have also been observed after SRSF1 aberrant expression [13]. In line with these findings, we showed that the activation of mTOR by SRSF1 regulates the translation of the antiapoptotic factor survivin [7]. Fu and colleagues also reported that SRSF1 regulates the expression of β-catenin, a key effector of the canonical Wnt signaling cascade, in an mTOR-dependent manner [14]. SRSF1 has also been associated with the maintenance of genomic instability through the suppression of transcription-linked ssDNA formation [15]. Moreover, some RBPs, including SRSF1, are directly involved in DNA repair through interactions with nascent transcripts, non-coding RNA or DNA repair genes [16, 17].
In the present study, we aimed to investigate whether the regulation of DNA repair genes through SRSF1 has a relevant role in lung carcinogenesis. First, we performed an in silico analysis of microarray data to select those DNA repair genes whose expression correlated with SFSR1 mRNA levels in lung cancer patients and cell lines. Among the five DNA repair genes with positive correlation, we found that SFSR1 binds to the mRNA of only one of them: DNA ligase I (LIG1). We found that SRSF1 regulates the expression of LIG1 protein, promoting the stability and translation of LIG1 mRNA. Moreover, in vitro downregulation of LIG1 decreased proliferation and increased cell death in non-small cell lung cancer (NSCLC) cell lines. In this paper, we also describe that LIG1 is up-regulated in human lung cancer tumors. Furthermore, the prognostic value of LIG1 is demonstrated and validated in NSCLC patients. In summary, we provide data suggesting the relevance of LIG1 in lung carcinogenesis and showing that its expression is regulated by the oncogenic mRNA-binding protein SRSF1.
Material and methods
Cell culture
Human NSCLC cell lines were purchased from American Type Culture Collection (ATCC), German Collection of Microorganisms and Cell Culture (DMSZ) or European Collection of Cell Cultures (ECACC) in 1998–1999. Cell lines (H520, H157, SK-MES1, A549, H322, H1395, H1299, 103H, H441, H1703, LXF289, and HCC15) were grown at 37 °C in an atmosphere of 5% CO2 and 95% relative humidity. Culture medium consisted of RPMI 1640 Glutamax (Invitrogen, Barcelona, Spain) supplemented with 10% fetalclone serum synthetic III (Hyclone/SH30109.03), 100 units/ml penicillin and 100 µg/ml streptomycin (Invitrogen/15140-122). Cells were authenticated by PCR analysis on the basis of their known mutations (COSMIC database) in 2012. Cell lines tested negative for mycoplasma using MycoAlert (Lonza: LT07-28). All experiments were performed within 10 passages after thawing cells.
Patient samples and tissue microarray construction
Archived formalin-fixed, paraffin-embedded specimens of patients diagnosed with NSCLC were obtained from Clínica Universidad de Navarra (Pamplona, Spain) (CUN) and University of Texas MD Anderson Cancer Center (Houston, TX) (MDA). Tissue specimens were examined and histologically classified using the 2004 WHO classification system for lung cancer [18]. For staging, the 7th TNM system was used [19]. The inclusion criteria were: complete resection of the primary lung tumor, absence of cancer within the 5 years prior to lung cancer surgery, absence of neoadjuvant therapy and available clinico-pathological information. Tissue microarrays containing representative areas from each tumor were constructed using a manual tissue arrayer. After histological examination of the NSCLC specimens, tissue microarrays were constructed using triplicated 1 mm tissue cores from each tumor. For survival analysis, the follow-up period was restricted to 60 months. Disease-free survival was calculated from the date of surgery to the date of recurrence or last follow-up. Overall survival was calculated from the date of surgery to the date of death or last follow-up. Detailed clinical and pathological information of the patients is summarized in Supplementary Table 1. The study protocol was approved by the institutional ethical committee of each center. Written informed consent was obtained from each patient. Reported recommendations for tumor marker prognostic studies (REMARK) criteria were followed throughout the study [20].
In silico analysis
SRSF1 and DNA repair genes mRNA expression levels were downloaded from TCGA and CCLE databases for correlation studies (https://cancergenome.nih.gov/; https://portals.broadinstitute.org/ccle). Data from three lung cancer microarray experiments deposited in GEO DataSets (http://www.ncbi.nlm.nih.gov/geo) were used for the correlation analysis [21,22,23]. GEO accession numbers and probes used in this part of the work were: GSE8894 (208863_s_at for SRSF1, 202726_at for LIG1 and 202123_s_at for Ki67), GSE26939 and GSE17710 (A_23_P49521 for SRSF1, A_23_P39116 for LIG1 and A_23_P202232 for Ki67). Survival analysis was conducted by Kaplan–Meier curves and log-rank test using the web-tool Kaplan–Meier Plotter (http://kmplot.com/analysis/index.php?p=service&cancer=lung) [24]. Patients were divided according to LIG1 mRNA expression levels (probe 202726_at for LIG1) using the upper tertile as cut-off.
RNA interference
SRSF1 and LIG1 expression was downregulated in NSCLC cell lines by RNA interference. A549, 103H, and HCC15 cells were seeded in six-well plates. The following day, cells were transfected with siRNAs (30 nM; Sigma-Aldrich, St. Louis, MO, USA) against SRSF1 (siSRSF1 #1 and siSRSF1 #2) or LIG1 (LIG1 #1 and LIG1 #2) in Opti-MEM Reduced Serum Medium with GlutaMAX (Gibco, Barcelona, Spain) using Lipofectamine 2000 (Invitrogen). The siRNA sequences were: siSRSF1 #1: 5′-GAAAGAAGAUAUGACCUAU-3′; siSRSF1 #2: 5′-TGAAGCAGGTGATGTATGT-3′; LIG1 #1: 5′-GGAAGAUGCCUGCUGGAAA-3′ and LIG1 #2: 5′-CCAAGAAGAUAGACAUCAU-3′. Cells were incubated for 2 or 3 days after transfection. As a negative control, a scrambled siRNA (Dharmacon, Lafayette, CO, USA) was used. To minimize the possibility of off-target effects, preliminary experiments were performed to determine the minimal concentration of siRNAs needed.
Reverse transcription and real-time quantitative PCR
RNA isolation was performed using RNAeasy kit (RNAeasy Mini Kit; Qiagen, Hilden, Germany). Reverse transcription was carried out using 2 µg of RNA, random hexamers, and SuperScript III RT (Invitrogen). Real-time PCR reactions were performed using the Applied Biosystems 7300 Sequence Detection System (Life Technologies, Carlsbad, CA, USA). Expression values were normalized to those of GAPDH. Primer sequences are listed in Supplementary Table 2.
Western blotting
Total proteins were extracted using RIPA buffer containing proteases inhibitors. Twenty micrograms from each lysate were boiled at 95 °C for 5 min, separated by SDS–PAGE gels and transferred onto nitrocellulose membranes. The membranes were blocked in 5% non-fat milk for one hour and incubated overnight with the primary antibodies for LIG1 (1:2000; Areta International, Gerenzano, VA, Italy); SRSF1 (1:12,000; Zymed, San Francisco, CA, USA) or β-actin (1:10,000; Sigma). Subsequently, horseradish peroxidase-conjugated antibodies were applied (1:5000; Amersham, Piscataway, NJ, USA). Chemoluminiscence detection was performed using Lumilight Plus Kit (Roche diagnostics, Manhein, Germany). Protein expression was quantified utilizing Fiji software [25].
Immunoprecipitation, RNA isolation, and quantification
After trypsinization, A549 cells were incubated with nuclear isolation buffer (1.28 M sucrose; 40 mM Tris pH 7.4; 20 mM MgCl2; 4% Triton X-100). Nuclear pellets were resuspended in RIP buffer (150 mM KCl; 25 mM Tris pH 7.4; 5 mM EDTA; 0.5% NP40, 0.5 mM DTT; and protease and RNase inhibitors). After centrifugation, supernatants were split into two fractions and 10% was saved as input. Twenty micrograms of anti-SRSF1 antibody (Zymed) or 20 µg of IgG1 isotype control antibody (Sigma) were added to each fraction and incubated overnight at 4 °C in continuous rotation. Magnetic beads were added and incubated for 1 h at 4 °C in continuous rotation. Five washes with RIP buffer and two washes with PBS were performed to wash off unbound material. Pellets were resuspended in trizol reagent (Invitrogen). RNA was extracted according to the manufacturer's protocol. RT-PCR was performed by using the total amount of RNA extracted. RNA quantification was represented as percentage of input.
Immunohistochemistry
Indirect immunohistochemistry was carried out on paraffin-embedded tissue or cells sections using the signal amplification Envision-system-HRP (Dako, Glostrup, Denmark). Endogenous peroxidase activity was blocked with 3% hydrogen peroxidase for 10 min and antigen retrieval was carried out by heating sections in a Lab Vision PT Module with Tris–EDTA buffer (pH 9) (Thermo Scientific, Waltham, MA, USA) for 20 min at 95 °C. Afterwards, tissues were incubated with primary antibodies: LIG1 (Areta International; 1:500), Ki67 (Neomarkers, Fremont, CA, USA; 1:200) or SRSF1 (Zymed; 1:200) at 4 °C overnight. Envision complex was added for 30 min and staining was visualized with 3-3′-diaminobenzidine and H2O2 addition. The specificity of LIG1 and SRSF1 antibodies was demonstrated using a variety of controls, including Western blot analysis, inhibition with siRNAs, isotype control, and omission of the primary antibody.
Immunostaining evaluation
Staining scores were established by semiquantitative analysis as previously described [26]. Briefly, staining was evaluated by two observers independently (MJP and EMT) blinded to the clinical features and outcomes of patients. The extension was scored as percentage of positive cells (0–100%), and the intensity of staining (1, weak; 2, moderate; 3, strong staining). A final score, called the H-score, was calculated by adding the products of the percentage of stained cells at a given staining intensity (0–100) by the staining intensity (0–3). The agreement between the two observers was verified by Gwet's AC1. Discordant independent readings were resolved by simultaneous review by both observers.
Cell proliferation assays
Anchorage-dependent cell proliferation was evaluated by thiazolyl blue tetrazolium bromide (MTT) assay. Twelve hours after transfection, cells were trypsinized and seeded in a 96-well at an appropriate cell density A549 (1500 cells/well), 103H (2000 cells/well) and HCC15 (2000 cells/well). For development, 10 µL of MTT at 5 mg/mL (Sigma/M2128) was added per well and incubated for 4 h at 37 °C. After that time, the formazan crystals were dissolved using a solubilizing agent (50% dimethylformamide and 20% SDS, pH 4.7). Absorbance was measured at 540/690 nm on a Sunrise plate reader (Tecan, Mannedorf, Switzerland). Anchorage-independent cell proliferation was analyzed by colony formation in soft agar. A549 (300 cells/well), 103H (600 cells/well), and HCC15 (400 cells/well) cells were seeded in six-well plates with a bottom layer of 0.6% agar and a top layer of 0.3% agar. Fourteen days later, cells were fixed in 4% buffered formalin for 30 min and stained with crystal violet to quantify colony formation. All experiments were performed in triplicate and repeated at least three times.
Cell cycle analysis
Floating and attached cells were collected and fixed with 70% ethanol for one hour at 4 °C and then washed with PBS. Fixed cells were treated with 0.2 mg/ml RNAase A (Sigma) for one hour at 37 °C and stained with 5 µg/ml of propidium iodide (PI, Sigma). Cells were analyzed in a flow cytometer (FACSCalibur, BD, San Jose, NJ, USA).
Apoptosis assay
The percentage of apoptotic cells was analyzed using the FITC Annexin V Apoptosis detection Kit I (BD). After transfection, floating and attached cells were collected and washed twice with PBS and kit buffer. Afterwards, cells were stained with PI and annexin V (AV) for 15 min and analyzed on a flow cytometer (FACSCalibur, BD). AV and PI positive cells were considered apoptotic cells.
Statistical analysis
For the correlation analysis between SRSF1 and DNA repair (in silico analysis), the Pearson coefficient was used after assuring bivariate normality. When variables did not follow this distribution, the Spearman coefficient was calculated (correlation between LIG1 and SRSF1 or LIG1 and Ki67 expression in cell lines). Chi-square test was used to analyze differences in LIG1 protein levels between two or more than two groups, following Cochran recommendations. Wilcoxon signed rank sum test was applied to analyze differences between tumors and matched normal tissues. Kaplan– Meier survival curves were generated to evaluate the role of LIG1 expression in the prognosis of NSCLC patients. Patients were stratified in two groups according to the upper tertile and long rank test was performed to assess the statistical differences. Univariate and multivariate Cox proportional hazards analyses were also used to assess the prognostic role of LIG1. Only those variables with P < 0.20 in the univariate analysis were included in the multivariate analysis. Student's t-test was used to analyze differences in proliferation, number of colonies, cell cycle, and apoptosis. A P-value < 0.05 was considered as statistically significant (*P < 0.05, **P < 0.01, ***P < 0.001). Statistical analyses were performed using SPSS 15.0 (Chicago, IL, USA) or STATA/IC 12.1 (StataCorp.).
Results
SRSF1 expression correlates and regulates LIG1 expression
To evaluate whether SRSF1 modulates the expression of DNA repair genes in NSCLC, we first studied the correlation between the mRNA levels of SRSF1 and different DNA repair genes through an in silico analysis of three different datasets (CCLE, TCGA ADC, and TCGA SCC). Pearson correlation coefficients are shown in Supplementary Table 3. We selected five DNA repair genes based on the correlation coefficients of the first 10 ranked genes in the three databases studied. Genes were: BLM, BRCA1, LIG1, MRE11A, and PARP1. Next, we tested whether SRSF1 binds to the mRNA of those genes. Endogenous SRSF1 was immunoprecipitated from A549 cells and total RNA bound to SRSF1 was extracted and analyzed by quantitative PCR to quantify BLM, BRCA1, LIG1, MRE11A, and PARP1 transcripts. SRSF1 immunoprecipitate was highly enriched with LIG1 mRNA, but not with mRNA from the others genes studied (Fig. 1a). Moreover, LIG1 mRNA presents the SRSF1-binding motif GAAGAA at exons 2, 6, and 24 [27]. Therefore, we selected LIG1 for further studies. LIG1 encodes for DNA ligase 1, an enzyme that catalyzes the joining of the Okazaki fragments during DNA replication.
SRSF1 binds to LIG1 mRNA, and correlates with LIG1 expression and regulates its expression in NSCLC cell lines. a Level of BLM, BRCA1, LIG1, MRE11A, and PARP1 mRNA enrichment in SRSF1 immunoprecipitates by RT-qPCR. Immunoprecipitation with an isotype control antibody (IgG) was used as negative control. Means ± SEM are shown for LIG1 gene (n = 2). b mRNA levels of SRSF1 and LIG1 in a panel of 11 NSCLC cell lines (left) and scatter plot showing the correlation of the expression between both genes (right). c Western blotting showing the expression level of LIG1 and SRSF1 proteins in NSCLC cell lines (left) and its correlation analysis after densitometry analysis (right). d Representative immunocytochemical staining for LIG1 and SRSF1 NSCLC cell lines (left) and immunocytochemical semi-quantification of both proteins in the panel of 11 NSCLC cell lines (right). Scale bar: 50 µm
To further confirm the correlation found between SRSF1 and LIG1, we conducted a new in silico analysis using three additional microarray databases containing a large number of patients with NSCLC (GSE8894, GSE26939, and GSE17710). The three databases showed a significant correlation between LIG1 and SRSF1 mRNA levels (P < 0.01; Supplementary Figure 1). Moreover, we evaluated by RT-PCR the correlation of SRSF1 gene expression with that of LIG1 in a panel of 11 NSCLC cell lines. We found a significant correlation between SRSF1 and LIG1 mRNA levels (r = 0.673, P = 0.023; Fig. 1b). The correlation was confirmed at the protein level by Western blotting (r = 0.673, P = 0.023; Fig. 1c and Supplementary Figure 3A) and immunocytochemistry in the same panel of 11 NSCLC cell lines (r = 0.697, P = 0.017; Fig. 1d). Taken together, these and the previously described data show that SRSF1 expression is associated with LIG1 expression in lung cancer.
To examine whether SRSF1 regulates LIG1 expression, we downregulated SRSF1 in lung cancer cell lines (A549, 103H, and HCC15) using two different siRNAs. SRSF1 down-regulation resulted in a marked reduction of LIG1 expression, both at the mRNA (Fig. 2a and Supplementary Figure 2) and protein level (Fig. 2b and Supplementary Figure 3B).
SRSF1 regulates LIG1 expression in NSCLC cell lines. RT-qPCR (a) and Western blot (b) analysis after inhibition of SRSF1 expression in A549, 103H, and HCC15 cell lines. β-actin was used as a loading control and densitometry analysis was performed to quantify the protein levels. RT-qPCR data in each cell line were normalized with its own Scramble
SRSF1 increases the stability of LIG1 mRNA and promotes its translation
We next studied the mechanism by which SRSF1 regulates LIG1 expression. Michlewski and colleagues have proposed that SRSF1 interacts with mTOR kinase facilitating the phosphorylation of the translational repressor 4E-BP1, which allows the release and activation of the translation initiation factor eIF4E [13]. We also demonstrated in a previous work that SRSF1 regulates the expression of survivin through the phosphorylation and inactivation of 4E-BP1, but also through the promotion of the stability of survivin mRNA [7]. Thus, we assessed whether SRSF1 increases LIG1 mRNA translation initiation and/or stability. First, we evaluated whether SRSF1 regulates LIG1 expression through the activation of mTOR. Treatment of A549 cells with the mTOR inhibitor rapamycin, produced a concentration-dependent decrease in the levels of LIG1 (Fig. 3a and Supplementary Figure 3C). Next, we tested the expression of LIG1 in A549 cells after SRSF1 downregulation in the presence of the transcription inhibitor actinomycin-D at different time points. Total protein extracts were obtained and LIG1 expression was analyzed by Western blotting. We found a time-dependent decrease of LIG1 protein in SRSF1 downregulated cells compared to control cells after actinomycin D treatment (Fig. 3b and Supplementary Figure 3D). These results suggest that SRSF1 regulates LIG1 expression by promoting the translation of LIG1 mRNA in an mTOR-dependent manner, and increasing its mRNA stability (Fig. 3c).
SRSF1 inhibition decreases LIG1 mRNA translation and stability. a Protein levels of A549 cells were analyzed by Western blotting after treatment with different concentrations of the mTOR inhibitor rapamycin. β-actin was used as a loading control, and survivin as a positive control. b Relative levels of LIG1 protein were analyzed by Western blotting at different times after actinomycin-D treatment in A549 cells, 2 days after siSRSF1 transfection. c Densitometry LIG1 values were normalized for each condition with β-actin (left) and natural logarithms of the values were used to statistically compare growth slopes (right). d Proposed model for the role of SRSF1 in the post-transcriptional regulation of LIG1 mRNA. SRSF1 promotes translation initiation by suppressing the activity of 4E-BP, a competitive inhibitor of cap-dependent translation. This activity is mediated by interactions of SRSF1 with mTOR, a key regulator of 4E-BP phosphorylation. Additionally, SRSF1 binds to LIG1 mRNA, protecting it from degradation
LIG1 inhibition decreases proliferation and increases cell death
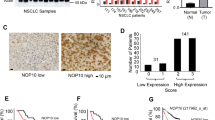
To further examine the role of LIG1 in lung cancer, we inhibited LIG1 in NSCLC cell lines and evaluated the effect on cell proliferation. LIG1 inhibition was performed in A549, 103H, and HCC15 cell lines using two different siRNAs. These cell lines represent the three main NSCLC histological subtypes. Both LIG1 siRNAs significantly reduced the proliferation rate of NSCLC cell lines (Fig. 4a–c, left), as well as their colony formation ability (Fig. 4a–c, right). We next analyzed the effect of LIG1 downregulation on the cell cycle. An increase in the percentage of cells in SubG0 phase was observed in LIG1-downregulated A549 cells as compared with the scrambled ones (Fig. 4d), suggesting an effect on cell death. Using the annexin V/PI assay, we confirmed that LIG1 downregulation results in an increase of apoptosis (Fig. 4e).
LIG1 inhibition reduces proliferation and induces apoptosis in NSCLC cell lines. a–c Left panels. MTT assay in A549 (a), 103H (b), and HCC15 (c) cell lines. Absorbance values were normalized to time 0. Means ± SEM of triplicates are shown. Natural logarithms of the values were used to evaluate the statistical differences between growth slopes. a–c Right panels. Clonogenic assays were performed in the same cell lines. Means ± SEM of triplicates and representative images are shown. d Representative experiment of the cell cycle analysis at 4 days after LIG1 inhibition. Percentages of cells in each phase (sub-G0–G1, G0–G1, S, and G2–M) are indicated. e Percentages of Annexin V-positive and PI-positive cells after 4 days of LIG1 downregulation. Attached and floating cells were collected and analyzed by flow cytometry. Means ± SEM from three independent experiments are shown
LIG1 expression in NSCLC patients
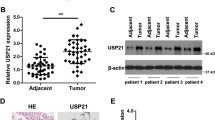
To assess the relevance of LIG1 in lung cancer, we evaluated by immunohistochemistry its expression in normal and tumor lung tissues from 97 patients from the CUN cohort. In non-tumor tissues, the vast majority of the alveolar parenchyma (including pneumocytes, macrophages, and inflammatory cells) showed no LIG1 staining; only some nuclei of the mucosa of the bronchi and bronchioli expressed LIG1 (Fig. 5a left). On the contrary, an intense LIG1 nuclear staining was found in most tumor samples (Fig. 5a right). In fact, a significant increase in LIG1 expression was found in tumors as compared with their non-tumor counterparts (P < 0.001).
LIG1 is overexpressed and correlates with cell proliferation and in NSCLC. a Representative immunohistochemical staining for LIG1 in a normal human lung and a tumor. Scale bar: 100 µm. b Immunohistochemical analysis of LIG1 and Ki67 expression in NSCLC patients. Representative images from two specimens with different expression levels for both proteins are shown. Patient 1 and 2 were diagnosed with SCC and ADC lung tumor, respectively. Scale bar: 100 µm. c Scatter plot showing the differences in the levels of LIG1 between ADC and SCC histologies in the CUN cohort (P < 0.001). d Scatter plot showing the differences in the levels of LIG1 between ADC and SCC histologies in the GSE8894 database (P < 0.001)
To further extend our in vitro results, we examined whether there was a correlation between LIG1 expression and proliferation, measured as the expression of the proliferation antigen Ki67 by immunochemistry. Ki67 is a well-established marker to detect proliferating cells, which is present during all phases of the cell cycle but absent in quiescent cells (reviewed in ref. [28]). Statistical analysis revealed a highly significant correlation between both proteins (r = 0.572; P < 0.001). Representative images are shown in Fig. 5b. Furthermore, this significant difference was confirmed in the three in silico mRNA databases (r = 0.551, P < 0.001 in GSE8894; r = 0.350, P < 0.001 in GSE26939; and r = 0.281, P = 0.036 in GSE17710). As observed in the in vitro experiments, LIG1 expression is associated with proliferation markers in NSCLC tumors.
We next evaluated the relationship between LIG1 levels in tumors and clinicopathological features of the patients from CUN cohort. Clinical data from these patients are listed in Supplementary Table 1. No correlation was found between LIG1 expression and gender, age, smoking status, or stage (Supplementary Table 4). Interestingly, a significant correlation was found between LIG1 expression and histology. SCC showed higher levels of LIG1 protein than ADC (P < 0.001; Supplementary Table 4 and Fig. 5c). Moreover, we validated the association between LIG1 mRNA levels and histology on the GSE8894 microarray database, which contains both ADC and SCC cases (62 and 66 cases, respectively). Congruently, lung SCC exhibited significantly higher mRNA LIG1 levels than ADC (P < 0.001, Fig. 5d). To corroborate these results, we analyzed LIG1 expression in tumor tissue from 210 NSCLC patients from MD Anderson Cancer Center. No correlation was found between LIG1 expression and gender, age, or stage in lung tumors; however, significant differences were found in SCC patients compared to ADC histology (P < 0.001, Supplementary Table 4). In this cohort, a significant association between LIG1 and smoking status was also observed (P = 0.001).
LIG1 is a marker for poor prognosis in NSCLC
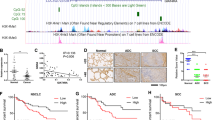
We next evaluated the relationship between LIG1 protein expression and prognosis in three different sets of NSCLC patients. Patients were stratified in two groups according to the upper tertile of the LIG1 expression and differences were evaluated using log-rank test. First, we analyzed the prognostic value of LIG1 mRNA levels using data retrieved from Kaplan–Meier Plotter web-tool. LIG1 mRNA levels were significantly associated with poor prognosis (P < 0.001 for DFS and OS, Fig. 6a). A further analysis at protein level was performed in two independent cohorts (CUN and MDA cohorts). In the CUN cohort, LIG1 protein expression was not statistically associated with outcome (P = 0.235 for DFS and P = 0.123 for OS), although a trend was observed in both cases (Fig. 6b). Taking into account the limited number of patients in the CUN cohort and the tendency observed, another cohort composed of 210 patients was evaluated (MDA cohort). We found that high levels of LIG1 protein were significantly associated with shorter DFS (P = 0.015, Fig. 6c). The same trend was observed for OS, although differences did not reach statistical significance (P = 0.128, Fig. 6c). We also evaluated the prognostic impact of LIG1 in this cohort by Cox regression analysis. High LIG1 expression was an independent prognostic factor for DFS when adjusting for gender, stage, and tobacco use (P = 0.044, HR = 1.63; 95% CI, 1.21–3.14, Supplementary Table 5). Altogether, these results suggest that the expression of LIG1 is associated with poor prognosis in NSCLC.
LIG1 expression is associated with prognosis. Kaplan–Meier survival curves for data collected from the Kaplan–Meier Plotter web-tool (a), for patients with NSCLC from CUN cohort (b), and for patients with NSCLC from MD Anderson Cancer Center (c). Patients were stratified in two groups according to the upper tertile of the H-scores. In all cases, differences between groups were evaluated using the log-rank test
Discussion
SRSF1 is an oncoprotein up-regulated in many tumors, including lung cancer [7]. In addition to its main role as a splicing regulator, SRSF1 has multiple activities on mRNA metabolism, which can be dysregulated in cancer development [5]. For instance, we previously reported that SRSF1 is implicated in the posttranscriptional regulation of the anti-apoptotic protein survivin, leading to NSCLC progression [7]. Since genomic instability is an underlying characteristic of cancer [29], numerous studies and clinical trials have been developed based on the inhibition of DNA damage response [30]. In this line, inhibitors of DNA-repairing enzymes, such as the PARP inhibitor olaparib have been recently approved for breast and ovary cancers [31].
In this work, we hypothesized that the oncoprotein SRSF1 could modulate the mRNA metabolism of DNA repair genes, contributing to a more aggressive phenotype of NSCLC. To address our objective, we first studied the correlation between the expression of SRSF1 and DNA repair-related genes. SRSF1 correlated with genes such as BLM, BRCA1, LIG1, MRE11A, and PARP1. More importantly, we observed that extracts of SRSF1 protein were enriched in LIG1 mRNA. In agreement with this observation, LIG1 mRNA presents the SRSF1-binding motif GAAGAA at exons 2, 6, and 24 [27]. LIG1 is an enzyme essential for joining the Okazaki fragments during DNA replication [32, 33]. This gene plays a role in DNA repair processes, such as base excision repair (BER), nucleotide excision repair (NER), single-strand break repair (SSB), and non-homologous end joining double-strand break repair (NHEJ) [34, 35]. Moreover, Sun and colleagues showed a higher expression of LIG1 in cancer cell lines in comparison with normal cells, as well as an increase in LIG1 protein associated with tumor growth in nude mice [36]. We hypothesized that SRSF1 regulates LIG1 supported by the strong positive correlation observed between their expression in lung cancer cell lines at mRNA and protein levels. In fact, SRSF1 downregulation resulted in the decrease of LIG1 mRNA and protein. Studies performed by Michlewsky et al. showed that SRSF1-activated mTOR and its target gene 4E-BP1 facilitating the liberation of the translation initiation factor eIF4E [13, 37]. We have demonstrated that SRSF1 induction of LIG1 is dependent of mTOR activation. We observed a significant decrease of LIG1 mRNA and protein levels upon SRSF1 downregulation. Moreover, the addition of mTOR inhibitor rapamycin, showed a dose-dependent reduction of LIG1 expression. For this reason and given the aforementioned studies, we propose mTOR as a plausible mechanism by which SRSF1 may be regulating LIG1. Additionally, as we previously reported for survivin [7], we have demonstrated herein that SRSF1 regulates LIG1 mRNA stability. Hence, SRSF1 regulates LIG1 mRNA in a post-transcriptional manner by promoting its translation and stability.
Considering that LIG1 has been reported as a protein upregulated in different cancer cell lines [36], we aimed to study its role in NSCLC. As a protein directly implicated in DNA synthesis and repair, we expected LIG1 to be implicated in proliferation and survival of NSCLC cells. Sun et al. have previously downregulated LIG1 in MCF-7 breast cancer cells by antisense oligodeoxynucleotide inhibition, showing a reduction in cell proliferation in a concentration-dependent manner [36]. Consistently with these data, we observed a significant reduction in the proliferation rate and clonogenicity potential in three NSCLC cell lines upon LIG1 downregulation. This reduction was also associated with an increase cell death (SubG0 phase) via apoptosis (Annexin V and PI positive cells), suggesting that LIG1 knock-down is sufficient to cause this effect. Taken all together, these data suggest that LIG1 plays a key role not only in proliferation but also in survival of lung cancer cells. One possible mechanism to explain this effect is that LIG1, as a DNA repair protein acts impairing cell death under oncogenic stress. To reinforce our in vitro results, we evaluated the proliferation rate of a cohort of lung primary tumors by analyzing the expression of the proliferation antigen Ki67 [28]. We correlated Ki67 and LIG1 levels in the same tumors. Remarkably, we found that patients with high LIG1 expression showed higher levels of Ki67, indicating that LIG1 was significantly elevated in highly proliferative lung primary tumors.
Due to the role of LIG1 in proliferation and cell survival we studied its potential prognostic value in NSCLC patients. Newman and colleagues previously revealed the prognostic value of LIG1 mRNA overexpression, among others DNA repair-related proteins, in neuroblastoma [38]. In accordance with the obtained results in neuroblastoma, high levels of LIG1 protein tended to be associated with a shorter DFS and OS in NSCLC.
Several anti-cancer drugs act inducing DNA damage in the cell. For this reason, DNA repair inhibition has emerged as a promising target for anticancer therapy [39]. Accordingly, DNA ligases have been suggested as therapeutic targets; in particular, specific LIG1 inhibitors as L82 and L82-G17 have been developed and applied to preclinical cell models in cancer, leading to a decrease in cell viability [40,41,42]. Of note, previous studies have pointed out that cells expressing LIG1 are more sensitive to L82-G17 inhibitor than isogenic LIG1 null cells [42]. These effects are in line with our findings, suggesting that further exploration of these molecules would help to determine their potential synergic use with the DNA damaging agents in the future. Nevertheless, the field of drug development is still struggling to find specific and uncompetitive LIG1 inhibitors. Our results support the necessity of further exploration of LIG1 inhibitors as anticancer drugs in NSCLC.
In summary, we have demonstrated a causal relationship between SRSF1 and LIG1 expression in lung cancer through posttranscriptional regulation. Besides, LIG1 inhibition reduces the proliferation rate, as well as the capacity to form colonies in the three histological subtypes of NSCLC and increases cell death. Moreover, LIG1 is overexpressed in primary non-small cell lung tumors compared with their normal counterparts, and may be associated with poor prognosis. Taking all into account, our results strongly suggest that LIG1 mediates some of the oncogenic effects of SRSF1 in the carcinogenesis of the lung.
References
Glisovic T, Bachorik JL, Yong J, et al. RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett. 2008;582:1977–86.
Kechavarzi B, Janga SC. Dissecting the expression landscape of RNA-binding proteins in human cancers. Genome Biol. 2014;15:R14.
Wang J, Liu Q, Shyr Y. Dysregulated transcription across diverse cancer types reveals the importance of RNA-binding protein in carcinogenesis. BMC Genom. 2015;16:S5. Suppl 7
Dvinge H, Kim E, Abdel-Wahab O, et al. RNA splicing factors as oncoproteins and tumour suppressors. Nat Rev Cancer. 2016;16:413–30.
Das S, Krainer AR. Emerging functions of SRSF1, splicing factor and oncoprotein, in RNA metabolism and cancer. Mol Cancer Res. 2014;12:1195–204.
Karni R, de Stanchina E, Lowe SW, et al. The gene encoding the splicing factor SF2/ASF is a proto-oncogene. Nat Struct Mol Biol. 2007;14:185–93.
Ezponda T, Pajares MJ, Agorreta J, et al. The oncoprotein SF2/ASF promotes non-small cell lung cancer survival by enhancing survivin expression. Clin Cancer Res. 2010;16:4113–25.
de Miguel FJ, Sharma RD, Pajares MJ, et al. Identification of alternative splicing events regulated by the oncogenic factor SRSF1 in lung cancer. Cancer Res. 2014;74:1105–15.
Jiang L, Huang J, Higgs BW, et al. Genomic landscape survey identifies SRSF1 as a key oncodriver in small cell lung cancer. PLoS Genet. 2016;12:e1005895.
Anczukow O, Rosenberg AZ, Akerman M, et al. The splicing factor SRSF1 regulates apoptosis and proliferation to promote mammary epithelial cell transformation. Nat Struct Mol Biol. 2012;19:220–8.
Gautrey HL, Tyson-Capper AJ. Regulation of Mcl-1 by SRSF1 and SRSF5 in cancer cells. PLoS ONE. 2012;7:e51497.
Ghigna C, Giordano S, Shen H, et al. Cell motility is controlled by SF2/ASF through alternative splicing of the Ron protooncogene. Mol Cell. 2005;20:881–90.
Michlewski G, Sanford JR, Caceres JF. The splicing factor SF2/ASF regulates translation initiation by enhancing phosphorylation of 4E-BP1. Mol Cell. 2008;30:179–89.
Fu Y, Huang B, Shi Z, et al. SRSF1 and SRSF9 RNA binding proteins promote Wnt signalling-mediated tumorigenesis by enhancing beta-catenin biosynthesis. EMBO Mol Med. 2013;5:737–50.
Li X, Manley JL. Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell. 2005;122:365–78.
Dutertre M, Lambert S, Carreira A, et al. DNA damage: RNA-binding proteins protect from near and far. Trends Biochem Sci. 2014;39:141–9.
Naro C, Bielli P, Pagliarini V, et al. The interplay between DNA damage response and RNA processing: the unexpected role of splicing factors as gatekeepers of genome stability. Front Genet. 2015;6:142.
Travis W, Brambilla E, Müller-Hermelink K, et al.. Pathology and genetics of tumours of the lung, pleura, thymus and heart. Lyon: IARC Press; 2004.
Goldstraw P. IASLC Staging manual in thoracic oncology. Orange Park, FL: Editorial Rx Press; 2009.
Altman DG, McShane LM, Sauerbrei W, et al. Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK): explanation and elaboration. PLoS Med. 2012;9:e1001216.
Lee ES, Son DS, Kim SH, et al. Prediction of recurrence-free survival in postoperative non-small cell lung cancer patients by using an integrated model of clinical information and gene expression. Clin Cancer Res. 2008;14:7397–404.
Wilkerson MD, Yin X, Walter V, et al. Differential pathogenesis of lung adenocarcinoma subtypes involving sequence mutations, copy number, chromosomal instability, and methylation. PLoS ONE. 2012;7:e36530.
Wilkerson MD, Yin X, Hoadley KA, et al. Lung squamous cell carcinoma mRNA expression subtypes are reproducible, clinically important, and correspond to normal cell types. Clin Cancer Res. 2010;16:4864–75.
Gyorffy B, Surowiak P, Budczies J, et al. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE. 2013;8:e82241.
Schindelin J, Arganda-Carreras I, Frise E, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Pajares MJ, Agorreta J, Larrayoz M, et al. Expression of tumor-derived vascular endothelial growth factor and its receptors is associated with outcome in early squamous cell carcinoma of the lung. J Clin Oncol. 2012;30:1129–36.
Sanford JR, Wang X, Mort M, et al. Splicing factor SFRS1 recognizes a functionally diverse landscape of RNA transcripts. Genome Res. 2009;19:381–94.
Jakobsen JN, Sorensen JB. Clinical impact of ki-67 labeling index in non-small cell lung cancer. Lung Cancer. 2013;79:1–7.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
O’Connor MJ. Targeting the DNA damage response in cancer. Mol Cell. 2015;60:547–60.
Robson M, Im S-A, Senkus E, et al. Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. New Engl J Med 2017;377:523–533
Montecucco A, Savini E, Weighardt F, et al. The N-terminal domain of human DNA ligase I contains the nuclear localization signal and directs the enzyme to sites of DNA replication. EMBO J. 1995;14:5379–86.
Howes TR, Tomkinson AE. DNA ligase I, the replicative DNA ligase. Subcell Biochem. 2012;62:327–41.
Montecucco A, Biamonti G, Savini E, et al. DNA ligase I gene expression during differentiation and cell proliferation. Nucleic Acids Res. 1992;20:6209–14.
Tomkinson AE, Howes TR, Wiest NE. DNA ligases as therapeutic targets. Transl Cancer Res. 2013;2:3.
Sun D, Urrabaz R, Nguyen M, et al. Elevated expression of DNA ligase I in human cancers. Clin Cancer Res. 2001;7:4143–8.
Karni R, Hippo Y, Lowe SW, et al. The splicing-factor oncoprotein SF2/ASF activates mTORC1. Proc Natl Acad Sci USA. 2008;105:15323–7.
Newman EA, Lu F, Bashllari D, et al. Alternative NHEJ pathway components are therapeutic targets in high-risk neuroblastoma. Mol Cancer Res. 2015;13:470–82.
Abbotts R, Thompson N, Madhusudan S. DNA repair in cancer: emerging targets for personalized therapy. Cancer Manag Res. 2014;6:77–92.
Chen X, Zhong S, Zhu X, et al. Rational design of human DNA ligase inhibitors that target cellular DNA replication and repair. Cancer Res. 2008;68:3169–77.
Zhong S, Chen X, Zhu X, et al. Identification and validation of human DNA ligase inhibitors using computer-aided drug design. J Med Chem. 2008;51:4553–62.
Howes TRL, Sallmyr A, Brooks R, et al. Structure-activity relationships among DNA ligase inhibitors: characterization of a selective uncompetitive DNA ligase I inhibitor. DNA Repair (Amst). 2017;60:29–39.
Acknowledgements
The authors thank Amaya Lavín for technical assistance. This work was supported by Foundation for Applied Medical Research (FIMA), Spanish Ministry of Economy and Innovation and Fondo de Investigación Sanitaria-Fondo Europeo de Desarrollo Regional (12/02040, PI13/00806, PI16/01821, RTICC RD12/0036/0040), and AECC Scientific Foundation (GCB14-2170). E. Martínez-Terroba was supported by a predoctoral fellowship from FIMA.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Martínez-Terroba, E., Ezponda, T., Bértolo, C. et al. The oncogenic RNA-binding protein SRSF1 regulates LIG1 in non-small cell lung cancer. Lab Invest 98, 1562–1574 (2018). https://doi.org/10.1038/s41374-018-0128-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-018-0128-2
This article is cited by
-
MicroRNAs from Holarrhena pubescens stems: Identification by small RNA Sequencing and their Potential Contribution to Human Gene Targets
Functional & Integrative Genomics (2023)
-
Autophagy regulation by RNA alternative splicing and implications in human diseases
Nature Communications (2022)
-
Expression profile of RNA binding protein in cervical cancer using bioinformatics approach
Cancer Cell International (2021)
-
Splicing factor SRSF1 promotes breast cancer progression via oncogenic splice switching of PTPMT1
Journal of Experimental & Clinical Cancer Research (2021)
-
ZRANB2/SNHG20/FOXK1 Axis regulates Vasculogenic mimicry formation in glioma
Journal of Experimental & Clinical Cancer Research (2019)