Abstract
Background:
The majority of prostate cancers (CaP) are detected in early stages with uncertain prognosis. Therefore, an intensive effort is underway to define early predictive markers of CaP with aggressive progression characteristics.
Methods:
In order to define such prognostic markers, we performed comparative analyses of transcriptomes of well- and poorly differentiated (PD) tumor cells from primary tumors of patients (N=40) with 78 months of mean follow-up after radical prostatectomy. Validation experiments were carried out at transcript level by quantitative real-time reverse transcriptase-PCR (RT-PCR) (N=110) and at protein level by immunohistochemistry (N=53) in primary tumors from an independent patient cohort.
Results:
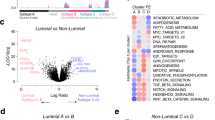
Association of a biochemical network of 12 genes with SPARC gene as a central node was highlighted with PD phenotype. Of note, there was remarkable enrichment of NKXH_NKXH_HOX composite regulatory elements in the promoter of the genes in this network suggesting a biological significance of this gene-expression regulatory mechanism in CaP progression. Further, quantitative expression analyses of SPARC mRNA in primary prostate tumor cells of 110 patients validated the association of SPARC expression with poor differentiation and higher Gleason score. Most significantly, higher SPARC protein expression at the time of prostatectomy was associated with the subsequent development of metastasis (P=0.0006, AUC=0.803).
Conclusions:
In summary, we propose that evaluation of SPARC in primary CaP has potential as a prognostic marker of metastatic progression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 4 print issues and online access
$259.00 per year
only $64.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Siegel R, Ward E, Brawley O, Jemal A . Cancer statistics, 2011: the impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J Clin 2011; 61: 212–236.
Denmeade SR, Isaacs JT . A history of prostate cancer treatment. Nat Rev Cancer 2002; 2: 389–396.
Nelson WG, De Marzo AM, Yegnasubramanian S . Epigenetic alterations in human prostate cancers. Endocrinology 2009; 150: 3991–4002.
Hessels D, Schalken JA . The use of PCA3 in the diagnosis of prostate cancer. Nat Rev Urol 2009; 6: 255–261.
Rubin MA, Zhou M, Dhanasekaran SM, Varambally S, Barrette TR, Sanda MG et al. alpha-Methylacyl coenzyme A racemase as a tissue biomarker for prostate cancer. JAMA 2002; 287: 1662–1670.
Petrovics G, Liu A, Shaheduzzaman S, Furusato B, Sun C, Chen Y et al. Frequent overexpression of ETS-related gene-1 (ERG1) in prostate cancer transcriptome. Oncogene 2005; 24: 3847–3852.
Witte JS . Prostate cancer genomics: towards a new understanding. Nat Rev Genet 2009; 10: 77–82.
Reynolds MA . Molecular alterations in prostate cancer. Cancer Lett 2008; 271: 13–24.
Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 2005; 310: 644–648.
Palanisamy N, Ateeq B, Kalyana-Sundaram S, Pflueger D, Ramnarayanan K, Shankar S et al. Rearrangements of the RAF kinase pathway in prostate cancer, gastric cancer and melanoma. Nat Med 2010; 16: 793–798.
Negrini S, Gorgoulis VG, Halazonetis TD . Genomic instability - an evolving hallmark of cancer. Nat Rev Mol Cell Biol 2010; 11: 220–228.
Hainaut P, Wiman KG . 30 years and a long way into p53 research. Lancet Oncol 2009; 10: 913–919.
Srikantan V, Srivastava S . Molecular dissection of the prostate cancer genome. In: Hofmann R, Heidenreich A, Moul JW (eds). Prostate Cancer. Springer-Verlag: Berlin, 2003, pp 25–40.
Dehm SM, Tindall DJ . Androgen receptor structural and functional elements: role and regulation in prostate cancer. Mol Endocrinol 2007; 21: 2855–2863.
Dobi A, Furusato B, Shaheduzzaman S, Chen Y, Vahey M, Nydam T et al. ERG expression levels in prostate tumors reflect functional status of the androgen receptor (AR) as a consequence of fusion of ERG with AR regulated gene promoters. The Open Cancer J 2010; 3: 101–108.
Sarker D, Reid AH, Yap TA, de Bono JS . Targeting the PI3K/AKT pathway for the treatment of prostate cancer. Clin Cancer Res 2009; 15: 4799–4805.
Varambally S, Dhanasekaran SM, Zhou M, Barrette TR, Kumar-Sinha C, Sanda MG et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 2002; 419: 624–629.
Tomlins SA, Rhodes DR, Yu J, Varambally S, Mehra R, Perner S et al. The role of SPINK1 in ETS rearrangement-negative prostate cancers. Cancer Cell 2008; 13: 519–528.
Clark JP, Cooper CS . ETS gene fusions in prostate cancer. Nat Rev Urol 2009; 6: 429–439.
Thomas R, True LD, Bassuk JA, Lange PH, Vessella RL . Differential expression of osteonectin/SPARC during human prostate cancer progression. Clin Cancer Res 2000; 6: 1140–1149.
Arnold SA, Brekken RA . SPARC: a matricellular regulator of tumorigenesis. J Cell Commun Signal 2009; 3: 255–273.
De S, Chen J, Narizhneva NV, Heston W, Brainard J, Sage EH et al. Molecular pathway for cancer metastasis to bone. J Biol Chem 2003; 278: 39044–39050.
Podhajcer OL, Benedetti LG, Girotti MR, Prada F, Salvatierra E, Llera AS . The role of the matricellular protein SPARC in the dynamic interaction between the tumor and the host. Cancer Metastasis Rev 2008; 27: 691–705.
Framson PE, Sage EH . SPARC and tumor growth: where the seed meets the soil? J Cell Biochem 2004; 92: 679–690.
Motamed K . SPARC (osteonectin/BM-40). Int J Biochem Cell Biol 1999; 31: 1363–1366.
Petrovics G, Zhang W, Makarem M, Street JP, Connelly R, Sun L et al. Elevated expression of PCGEM1, a prostate-specific gene with cell growth-promoting function, is associated with high-risk prostate cancer patients. Oncogene 2004; 23: 605–611.
Werner T . Regulatory networks: linking microarray data to systems biology. Mech Ageing Dev 2007; 128: 168–172.
Werner T . Bioinformatics applications for pathway analysis of microarray data. Curr Opin Biotechnol 2008; 19: 50–54.
Seifert M, Scherf M, Epple A, Werner T . Multievidence microarray mining. Trends Genet 2005; 21: 553–558.
Cohen CD, Lindenmeyer MT, Eichinger F, Hahn A, Seifert M, Moll AG et al. Improved elucidation of biological processes linked to diabetic nephropathy by single probe-based microarray data analysis. PLoS One 2008; 3: e2937.
Sharad S, Srivastava A, Ravulapalli S, Parker P, Chen Y, Li H et al. Prostate cancer gene expression signature of patients with high body mass index. Prostate Cancer Prostatic Dis 2011; 14: 22–29.
Cornford PA, Dodson AR, Parsons KF, Desmond AD, Woolfenden A, Fordham M et al. Heat shock protein expression independently predicts clinical outcome in prostate cancer. Cancer Res 2000; 60: 7099–7105.
Lapointe J, Li C, Higgins JP, van de Rijn M, Bair E, Montgomery K et al. Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc Natl Acad Sci USA 2004; 101: 811–816.
Santagata S, Demichelis F, Riva A, Varambally S, Hofer MD, Kutok JL et al. JAGGED1 expression is associated with prostate cancer metastasis and recurrence. Cancer Res 2004; 64: 6854–6857.
Szász AM, Nyirády P, Majoros A, Szendrõi A, Szûcs M, Székely E et al. beta-catenin expression and claudin expression pattern as prognostic factors of prostatic cancer progression. BJU Int 2010; 105: 716–722.
Li R, Dai H, Wheeler TM, Sayeeduddin M, Scardino PT, Frolov A et al. Prognostic value of Akt-1 in human prostate cancer: a computerized quantitative assessment with quantum dot technology. Clin Cancer Res 2009; 15: 3568–3573.
Gurel B, Iwata T, Koh CM, Yegnasubramanian S, Nelson WG, De Marzo AM . Molecular alterations in prostate cancer as diagnostic, prognostic, and therapeutic targets. Adv Anat Pathol 2008; 15: 319–331.
Quinn DI, Henshall SM, Sutherland RL . Molecular markers of prostate cancer outcome. Eur J Cancer 2005; 41: 858–887.
Acknowledgements
This work was supported by grant RO1-DK065977 to SS and GP from the National Institutes of Health, and by the Center for Prostate Disease Research, a program of the Henry M. Jackson Foundation for the Advancement of Military Medicine (Rockville, MD), funded by the US Army Medical Research and Materiel Command.
Disclaimer
The views expressed in this manuscript are those of the authors and do not reflect the official policy of the Department of the Army, Department of Defense or the US Government.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Prostate Cancer and Prostatic Diseases website
Supplementary information
Rights and permissions
About this article
Cite this article
DeRosa, C., Furusato, B., Shaheduzzaman, S. et al. Elevated osteonectin/SPARC expression in primary prostate cancer predicts metastatic progression. Prostate Cancer Prostatic Dis 15, 150–156 (2012). https://doi.org/10.1038/pcan.2011.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pcan.2011.61
Keywords
This article is cited by
-
Quantitative proteomics identifies and validates urinary biomarkers of rhabdomyosarcoma in children
Clinical Proteomics (2023)
-
Study on the role of SLC14A1 gene in biochemical recurrence of prostate cancer
Scientific Reports (2022)
-
Re-sensitization of 5-FU resistance by SPARC through negative regulation of glucose metabolism in hepatocellular carcinoma
Tumor Biology (2015)
-
SPARC/osteonectin is involved in metastatic process to the lung during melanoma progression
Virchows Archiv (2014)
-
SPARC mediates metastatic cooperation between CSC and non-CSC prostate cancer cell subpopulations
Molecular Cancer (2014)