Abstract
Epithelial-to-mesenchymal transition (EMT) is associated with poor prognosis and metastasis in hepatocellular carcinoma. We have previously demonstrated an in vivo model of liver cancer in which mesenchymal cells post-EMT demonstrate a high rate of invasive growth and metastasis. Here, we investigate the role of microRNA 200 (miR-200) family members and epigenetic modifications on the maintenance of mesenchymal/metastatic phenotype after EMT. Mesenchymal cells post-EMT demonstrates high levels of E-box repressors Zeb1 and Zeb2 and downregulation of four miR-200 family members (miR-200a, miR-200b, miR-200c and miR-429). In addition, DNA sequencing after bisulfite modification demonstrates that several CpG sites within the E-cadherin promoter are methylated in mesenchymal cells. In mesenchymal cells, forced expression of miR-200b results in a significant increase in E-cadherin and a reduction in cell migration/invasion. Despite these mesenchymal-to-epithelial transition (MET) changes in vitro, there is no significant change in metastatic potential after miR-200b upregulation in vivo. After the mesenchymal cells were treated with combination of DNA methyltransferase (DNMT) inhibitor and upregulation of miR-200b, invasive phenotype was significantly reduced and metastatic potential was eliminated. Direct targeting of E-cadherin with short hairpin RNA does not restore metastatic potential after DNMT inhibition and miR-200b re-expression. In addition, restoration of E-cadherin alone was unable to block metastatic potential in primary mesenchymal cells. In conclusion, targeting mesenchymal liver cancer cells with miR-200b and DNMT inhibitor reduces metastatic potential irrespective of E-cadherin expression. Thus, the broader differentiation and MET effects of DNMT inhibition and miR-200b must be considered in terms of rescuing metastatic potential.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) remains the third leading cause of cancer-related death world-wide,1 and the incidence of HCC related to Hepatitis C virus infection in the United States has increased 10 fold in the last decade.2 Although the mortality of HCC patients is primarily related to metastatic disease,3 the mechanisms underlying HCC invasion and metastasis are not well defined.4
Metastasis is a multi-step process that requires fixed epithelial carcinomas to invade into surrounding tissue, survive the circulatory system and establish a distant colony. One proposed theory regarding metastasis is that epithelial carcinomas undergo epithelial-to-mesenchymal transition (EMT).5, 6 After EMT, mesenchymal cells demonstrate decreased adhesion and increased motility, typically associated with the downregulation of E-cadherin, a component of the epithelial adherens junction.7, 8 During EMT, a number of transcription factors, such as Snail, Slug, Zeb1, Zeb2, Twist1/2 and E12/E47, bind to the E-cadherin promoter and inhibit transcription.7, 9, 10
EMT and downregulation of E-cadherin correlate with poor outcomes in HCC.11, 12, 13 In a recent work, we demonstrated that epithelial HCC tumor cells acquired a mesenchymal phenotype via EMT in vivo.14 In this model, mesenchymal cells post-EMT demonstrated invasive growth and established metastatic colonies compared with epithelial counterparts, which have slower local growth without significant invasion or metastasis.14
Recent work links the alterations of microRNA species to EMT. Specifically, downregulation of the microRNA 200 (miR-200) family has a significant role in the initiation and maintenance of EMT in a canine kidney epithelial cell.15 In this system, forced expression of miR-200 in mesenchymal cells induced mesenchymal-to-epithelial transition (MET). In addition, the expression of the miR-200 family was reduced in mesenchymal-phenotype cancer cells from the 60 human lines of the drug screening panel at the National Cancer Institute (NCI 60).16 However, the direct role of miR-200 family in tumor metastasis remains controversial. On the one hand, increased expression of miR-200 in metastatic breast cancer cells induces MET and reduces tumor cell motility.16 Conversely, miR-200 may facilitate breast cancer metastasis by enhancing colonization.17, 18
The miR-200 family is highly conserved across different species, and consists of five members organized as two clusters, miRs-200b/a/429 and miRs-200c/141, on chromosomes 1 and 12 in humans, and chromosomes 4 and 6 in mice.19 In epithelial cells, miR-200 targets the 3′-UTR of Zeb1 and Zeb2 mRNA.15, 20, 21, 22 In HCC, miR-200a is downregulated in tumor tissue compared with adjacent liver tissues.23 In terms of survival, decreased miR-200b expression correlates with poor patient outcomes.24 However, the precise role of the miR-200 family in the HCC progression and metastasis is not well defined.
In addition to transcriptional repressors, epigenetic modification is associated with EMT and metastatic disease. Specifically, the CpG island of E-cadherin promoter can be highly methylated in a variety of human cancers, such as colorectal,25, 26 prostate,27 cervical,28 and breast cancer.29 In HCC, E-cadherin promoter hyper-methylation was associated with reduced E-cadherin expression and disease progression.30
In this current study, we demonstrate that the miR-200 family was significantly downregulated in mesenchymal HCC cells compared with counterpart epithelial cells. Forced expression of miR-200b in mesenchymal cells results in Zeb1 and Zeb2 suppression, E-cadherin upregulation and the inhibition of tumor migration and invasion in vitro, but does not reduce metastatic potential in vivo. Further analysis demonstrates that the E-cadherin promoter is highly methylated in mesenchymal cells. For the first time, we demonstrate that the combination of a DNA methyltransferase (DNMT) inhibitor and restoration of miR-200b results in significant E-cadherin expression, MET, and complete loss of metastatic potential. However, this loss of metastatic potential was not overcome with direct targeting of E-cadherin.
Results
The miR-200 family is downregulated in mesenchymal cells
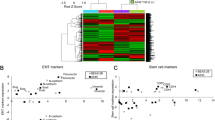
The miR-200 family members are downregulated in many tumor cells with mesenchymal phenotype, and miR-200 levels inversely correlate with target genes, Zeb1 and Zeb2.16 In a previous report, Pten−/− HCC epithelial cells acquired EMT after serial in vivo transplantation. The mesenchymal cells post-EMT demonstrate fibroblastoid morphology compared with cobble-stone morphology of the epithelial cells, and these mesenchymal cells are characterized by aggressive growth, invasion and metastasis in vivo.14 In the mesenchymal cells, E-cadherin is significantly downregulated, and transcriptional repressors that drive EMT, Zeb1 and Zeb2, are significantly upregulated.14 Further analysis demonstrates that four members of the miR-200 family (miR-200a, miR-200b, miR-200c and miR-429) are significantly downregulated in the mesenchymal cells (Figure 1a). miR-205, which is not in the miR-200 family, but can target Zeb1 and Zeb2,15 demonstrates no significant change after EMT (data not shown).
Transient delivery of pre-miR-200a or pre-miR-200b in mesenchymal cells partially reversed EMT. (a) Mesenchymal (M) cells demonstrate loss of four miR-200 family members including miR-200a, miR-200b, miR-200c and miR-429 compared with parent epithelial (E) cells (n=3, *P<0.05). (b) Zeb1 and Zeb2 were significantly downregulated and E-cadherin was significantly upregulated in precursor miR-200-transfected mesenchymal cells compared with cells transfected with NC pre-miRNA (n=3, *P<0.05). (c) Protein expression of E-cadherin was significantly upregulated after miR-200 transfection in mesenchymal cells. (d) Compared with NC miRNA, the transient expression of miR-200a or miR-200b significantly inhibited tumor cell migration on wound-healing assay (n=3, NC 149±21, miR-200a 54±10 and miR-200b 26±8, *P<0.05 compared with NC).
Forced expression of miR-200b precursor results in partial MET in vitro
To investigate whether restoration of miR-200 can induce MET, we performed transient transfection with miR-200a or miR-200b precursors in mesenchymal cells. In mesenchymal cells, treatment with miR-200a or miR-200b precursors downregulates Zeb1 and Zeb2 expression, increases E-cadherin expression and inhibits cell migration (Figures 1b–d). There is no significant change in cell proliferation between control, miR-200a and miR-200b groups (data not shown). As miR-200b is the strongest inducer of E-cadherin and inhibitor of cell migration, we developed a stable miR-200b expression vector (Materials and methods). The negative control (NC) empty vector only expresses GFP (green fluorescence protein) (miR-NC). Using this model, the expression of miR-200b is 120 fold greater than that of parent epithelial cells (Supplementary Figure 1a). This upregulation of miR-200b has no effect on the expression of other miRNAs, such as miR-200c and miR-16 (Supplementary Figures 1b and c). This stable expression of miR-200b is associated with change in cell morphology to larger cells with increased cell–cell contact in vitro, significant suppression of Zeb1 and Zeb2 and upregulation of E-cadherin (Figures 2a–c, and Supplementary Figures 2a and 3). In time-lapse video, the stable expression of miR-200b in mesenchymal cells results in reduced cell mobility (Supplementary Video 1 (miR-NC) and Video 2 (miR-200b)). As shown in Figure 2d, cell invasion is significantly inhibited in mesenchymal cells with stable expression of miR-200b compared with miR-NC cells. In this model of forced miR-200b expression in mesenchymal cells, Zeb1 overexpression results in increased migration in vitro, confirming that miR-200b targets Zeb1 specifically as one mechanism of miR-200b-driven MET (Supplementary Figure 4).
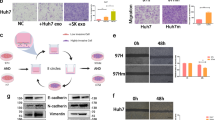
Stable expression of miR-200b reverses EMT. (a) In mesenchymal cells, miR-200b expression results in downregulation of Zeb1 and Zeb2, and upregulation of E-cadherin compared with NC miRNA (miR-NC, n=3/group, *P<0.01). (b) The protein level of E-cadherin was upregulated in miR-200b expressing mesenchymal cells compared with control miR-NC cells, but remained below the level of parent epithelial cells (n=3, representative image shown). (c) Green fluorescence protein (GFP) fluorescence demonstrates that mesenchymal cells undergo morphology change to larger cells with more cell–cell contact after miR-200b upregulation (n=10, representative image shown). (d) Compared with miR-NC control mesenchymal cells, invasion is significantly inhibited after miR-200b expression (expressed as mean number of cells invading into lower chamber, n=3, *P<0.05). (e) Mesenchymal cells expressing miR-NC or miR-200b spread to multiple abdominal organs following orthotopic transplant in nude mice, as demonstrated by in vivo imaging (n=8–9 mice/group, representative images shown). (f) Histology demonstrates mesenchymal cells with miR-200b expression invade the pancreas (P) and colon (C) (L, liver; T, tumor). Metastatic lesions (arrow) are detectable in the lungs of both miR-NC and miR-200b groups (miR-NC, n=9, lung metastasis 22.2% (2/9), miR-200b, n=8, lung metastasis 37.5% (3/8), H&E stain).
Forced expression of miR-200b is unable to inhibit metastasis
To investigate whether restoration of miR-200b expression is capable of inhibiting metastasis, we utilized mesenchymal cells with stable expression of miR-200b in an orthotopic tumor model. Histology confirms suppressed Zeb1 and Zeb2 and increased E-cadherin in tumors from miR-200b treated mesenchymal cells compared with miR-NC mesenchymal controls (Supplementary Figures 2c and 5). Despite these changes, tumors from mesenchymal cells with forced miR-200b expression demonstrate significant invasion within multiple abdominal organs similar to miR-NC mesenchymal control cells (Figures 2e and 2f, Table 1). Histology of lung tissue confirms no significant difference between the number of mice with lung tumor metastatic nodules (Figure 2f and Table 1, miR-NC 22% vs miR-200b 38%). These findings demonstrate that the restoration of miR-200b expression in mesenchymal cells is insufficient to inhibit lung metastasis.
E-cadherin promoter is highly methylated in mesenchymal cells
In addition to transcriptional repressors, E-cadherin promoter methylation is a mechanism for suppression of E-cadherin transcription in HCC.30, 31 Previous reports demonstrate that methylation of the CpG island in the E-cadherin promoter correlates to poor HCC clinical outcomes.30, 32 We utilized DNA sequencing after bisulfite modification to demonstrate that 14 cytosine residues in the E-cadherin promoter CpG island are methylated in mesenchymal cells (Figure 3a and Supplementary Figure 6). This CpG methylation is reduced using DNMT inhibitor decitabine (Figure 3a).
E-cadherin promoter CpG island is methylated in mesenchymal cells. (a) Compared with epithelial cells, DNA of mesenchymal cells analyzed by sequencing after bisulfite treatment demonstrates CpG sites that are methylated in the E-cadherin promoter (red arrows). Graphic representation of E-cadherin promoter with the open and filled circles denoting non-methylated and methylated CpG sites, respectively. The methylation sites of E-cadherin were reduced in mesenchymal cells treated with decitabine. (b) Inhibition of DNA methylation with decitabine in combination with miR-200b upregulation results in synergistic increase in E-cadherin expression (n=3, *P<0.01). (c) Western blot confirms E-cadherin was upregulated following decitabine treatment of mesenchymal cells expressing miR-200b (n=3, representative image shown).
Combination of miR-200b upregulation with DNMT inhibition results in the strongest induction of E-cadherin.
Mesenchymal cells treated with decitabine demonstrate no significant change in E-cadherin mRNA and protein expression (Figures 3b and c), no significant change in morphology (data not shown) and no reduction in migration using time-lapse video (Supplementary Video 3 (miR-NC+decitabine)). However, combination treatment with decitabine and upregulation of miR-200b results in a synergistic increase in E-cadherin (Figures 3b and c, 1408-fold increase in E-cadherin in decitabine+miR-200b group, P<0.01). In addition, we note an increase in cell-to-cell contact and reduction in cell motility in time-lapse video in this combination miR-200b and decitabine treatment group (Figure 4a and Supplementary Video 4 (miR-200b+decitabine)).
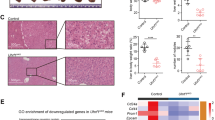
Metastatic potential inhibited after miR-200b upregulation and DNMT inhibition. (a) Mesenchymal cells treated with miR-200b and decitabine have significant reduction in migration compared with mesenchymal cells treated with decitabine (miR-NC+vehicle 350±13, miR-NC+decitabine 331±21, miR-200b+decitabine 104±34, *P<0.01 for miR-200b+decitabine compared with other groups). (b) In orthotopic liver tumor models, compared with miR-NC vehicle control cells and miR-NC+decitabine, combination treatment with miR-200b and decitabine results in reduction of invasive growth, as demonstrated by in vivo luciferase bioluminescent imaging (n=4–5/group, representative images shown). (c) Histological examination demonstrates that both miR-NC vehicle control and miR-NC+decitabine-treated groups have invasive growth into the pancreas (P) and colon (C) (T, tumor, L, liver). No significant invasion of tumors is observed in miR-200b+decitabine group. Metastatic tumors in the lungs (arrows) after inoculation with miR-NC vehicle control (1/4), miR-NC+decitabine (2/5), but no metastasis nodules are observed in miR-200b+decitabine group (0/5, H&E stain).
One potential mechanism for this synergistic change after DNMT inhibitor treatment is epigenetic modification of the miR-200 promoter. After decitabine treatment, miR-200b expression is increased threefold compared with control mesenchymal cells, but remains extremely low level compared with both epithelial cells and mesenchymal cells with forced miR-200b expression (data not shown). Other miR-200 family members demonstrate no increase after decitabine treatment. These results indicate that decitabine is not synergizing with miR-200b by specifically increasing miR-200b levels.
Combination of DNMT inhibition and miR-200b overexpression inhibits invasion and metastasis
In order to analyze whether the combination of miR-200b and DNMT inhibition could inhibit tumor migration in vitro, we utilized wound-healing assays. Mesenchymal cells treated with miR-200b and decitabine combination demonstrate significant reduction in cell migration compared with miR-NC vehicle control group and miR-NC+decitabine-treated mesenchymal cells (n=3, miR-NC vehicle 350±13, miR-NC+decitabine 331±21, miR-200b+decitabine 104±34, *P<0.01 for miR-200b+decitabine compared with other groups).
Before in vivo tumor metastasis assays, we confirmed that decitabine treatment would result in a sustained effect in E-cadherin expression in mesenchymal cells expressing miR-200b. Cells were treated with decitabine for 48 h and maintained without decitabine for 1 week. In combination with miR-200b upregulation, this treatment results in stable E-cadherin upregulation in mesenchymal cells (Supplementary Figure 7). We then orthotopically inoculated mesenchymal cells with miR-200b and decitabine treatment for analysis of in vivo metastasis. As shown in Figure 4b, in vivo bioluminescence images demonstrate that miR-NC mesenchymal cells treated with decitabine produce tumors with invasive growth similar to vehicle controls. However, the animals that received mesenchymal cells with combination miR-200b and decitabine treatment demonstrate tumors with restricted growth around the orthotopic inoculation area. Histology confirms increased E-cadherin in the tumors of this miR-200b and decitabine group (Supplementary Figure 5). Using an invasion score of pancreatic and colon invasion, we confirm the loss of invasive disease in the miR-200b and decitabine combination group (Figure 4c, P<0.01 when invasion score from mesenchymal miR-200b+decitabine group in Table 1 is compared with mesenchymal miR-NC, mesenchymal miR-NC+vehicle, mesenchymal miR-NC+decitabine and mesenchymal miR-200b groups). In addition, no mice (0%) demonstrate lung tumor nodules in the mesenchymal miR-200b and decitabine combination group, compared with 22–40% in the other treatment groups (Table 1).
Forced E-cadherin expression in mesenchymal cells is unable to reduce the metastatic potential
As combination treatment of mesenchymal cells with miR-200b and decitabine results in synergistic upregulation of E-cadherin, we utilized an E-cadherin expression vector in mesenchymal cells to drive E-cadherin to levels above those in the original epithelial cells (Figure 5a). We sought to test the hypothesis that focused E-cadherin upregulation reduces the metastatic potential of mesenchymal cells. Immunohistochemistry demonstrates that E-cadherin is abundantly localized to cell–cell connections in parent epithelial cells. In mesenchymal cells with forced E-cadherin upregulation, increased E-cadherin staining does not localize to cell–cell contacts and the cells maintain fibroblastiod morphology (Figure 5b). In addition, mesenchymal cells with forced E-cadherin expression demonstrate no significant change in migration or invasion (Figures 5c and d). Using the orthotopic liver tumor model, forced expression of E-cadherin in mesenchymal cells does not reduce invasion and lung metastasis, despite increased E-cadherin observed on tumor histology (Figure 5e, Table 1, and Supplementary Figure 8).
Direct expression of E-cadherin in mesenchymal cells is not sufficient to inhibit lung metastasis. (a) E-cadherin expression was restored in mesenchymal cells post-EMT using stable expression system. High levels of E-cadherin expression are confirmed in all four stable expression pools, mesenchymal Ecad p1, p2, p3 and p4. (b) Immunofluorescence staining demonstrates that E-cadherin is localized to the cell membrane in parental epithelial cells, and E-cadherin is localized to the cytoplasm and reduced in the mesenchymal cells. In mesenchymal cells with stable E-cadherin expression, the E-cadherin staining is increased but remains in a pattern suggestive of mesenchymal cells. No difference in the capability of cell migration (c) and invasion (d) between primary mesenchymal cells and mesenchymal cells with stable E-cadherin overexpression (n=3/group, P>0.05 for both migration and invasion assays). (e) Restoration of E-cadherin expression fails to inhibit tumor invasion into the pancreas (P) and colon (C) and lung metastasis (T, tumor; L, liver; arrows indicate lung metastasis, H&E stain).
Knockdown of E-cadherin in mesenchymal cells treated with miR-200b and inhibitor does not restore metastatic potential
In mesenchymal cells treated with miR-200b and decitabine, we utilized a stable E-cadherin short hairpin RNA (shRNA) vector to test the hypothesis that the synergistic upregulation of E-cadherin is a primary mechanism for the inhibition of metastasis. In this model, E-cadherin shRNA results in downregulation of E-cadherin expression (Figures 6a and b). This downregulation of E-cadherin is associated with increased migration and invasion in vitro (Figures 6c and d). In vivo, targeting E-cadherin in mesenchymal cells treated with miR-200b and decitabine results in increased invasion without evidence of increasing metastatic disease (Figure 6e and Table 1, P<0.05 comparing invasion score of mesenchymal NC-shRNA+miR-200b+decitabine group to mesenchymal E-cad-shRNA+miR-200b+decitabine group).
Knock-down of E-cadherin in mesenchymal cells treated with miR-200b and DNMT inhibitor promotes invasion. Stable E-cadherin knock-down using shRNA demonstrated in mesenchymal cells expressing miR-200b+decitabine treatment detected on (a) western blot and (b) immune-fluorescence staining. (c) Compare with NC-shRNA, knockdown of E-cadherin in miR-200b+decitabine-treated cells increased tumor cell migration (NC-shRNA 341±22 vs Ecad-shRNA 542±35, P<0.05) and invasion (d) (NC-shRNA 40±5 vs Ecad-shRNA 66±8, *P<0.05). (e) H&E staining demonstrates that the knock-down of E-cadherin in miR-200b and decitabine-treated mesenchymal cells promotes tumor invasion of pancreas (P) and colon (C) compared with stable NC-shRNA cells (T, tumor; L, liver). E-cadherin shRNA knock-down did not increase lung metastasis (n=5/group, NC-shRNA 0/5, E-cad-shRNA 0/5).
In Table 1, we summarize all in vivo tumor invasion and metastasis results. We developed a scoring system (Materials and methods) to evaluate the capacity of peritoneal invasion, as the pancreas and colon are two major organs invaded by the mesenchymal cell tumors.14 As shown in Table 1, the invasion score equals 1.0 in all groups treated with combination of miR-200b and decitabine, and the invasion score for other groups is ⩾2.0, indicating that tumor invasion is significantly inhibited with the combination of miR-200b restoration and DNMT inhibition (P<0.01). Although targeting E-cadherin specifically in this combination miR-200b and decitabine group induces increased invasion, loss of E-cadherin does not increase metastasis.
Additional markers of the EMT/MET spectrum in liver cancer
As focused E-cadherin expression does inhibit a metastatic phenotype, and as targeting E-cadherin in miR-200b and decitabine-treated cells does not re-induce a metastatic phenotype, we sought to understand alternative mechanisms driving changes in the metastatic phenotype, independent of E-cadherin. After miR-200b upregulation and decitabine treatment, Cytokeratin-19 (Krt-19) and HNF4α are both synergistically upregulated (Supplementary Figure 9). Krt-19 is a marker of epithelial cholangiocytes and HNF4α is a marker of hepatocyte differentiation. As alterations in E-cadherin expression alone are not sufficient to directly affect the metastatic process in our model, we propose that the broader MET and differentiation effects of miR-200b and DNMT inhibition may be a mechanism inhibiting metastatic potential.
Discussion
In our previous investigation, murine mesenchymal cells derived from epithelial liver tumor after EMT in vivo demonstrate invasive growth with significant metastatic potential.14 In human HCC, growing evidence demonstrates that the acquisition of mesenchymal phenotype has an important role in tumor progression and metastatic behavior.33, 34, 35 Thus, we hypothesized that targeting the initiation and maintenance of EMT will result in a reduction in liver tumor metastasis. In murine lung adenocarcinoma, forced expression of miR-200 blocks the capability of tumor cells to invade and metastasize.36 In the present study, we discovered a significant loss of miR-200 in mesenchymal tumor cells, which is correlated with tumor invasion and metastasis. Although re-expression of miR-200b reduces mesenchymal cell migration and invasion in vitro, miR-200b expression in mesenchymal cells does not inhibit a metastatic phenotype in vivo. After forced expression of miR-200b was combined with DNMT inhibition in mesenchymal cells, we demonstrated that tumor invasion and metastasis capability was significantly inhibited. This finding led us to the initial conclusion that E-cadherin may have a critical role in liver tumor metastasis. However, focused re-expression of E-cadherin in mesenchymal cells failed to reduce the invasive and metastatic phenotype, implying that the effects of miR-200b and DNMT inhibition are not mediated by the direct restoration of E-cadherin.
In this study, forced expression of miR-200b does inhibit cell migration and invasion in vitro, but does not reduce metastasis in vivo. These findings are consistent with a recent work in which miR-200 expression reduces breast cancer migration and invasion in vitro and enhances metastasis by increasing colonization.18 Thus, the therapeutic implication of miR-200 as an inhibitor of metastasis provides a classic double-edge sword. miR-200b may reverse EMT and block the tumor migration/invasion features that are characterized by a mesenchymal phenotype, but miR-200 may enhance metastatic colonization by promoting an epithelial phenotype. In current study, the rate of lung metastasis is not significantly different in mesenchymal cells treated with either miR-200b or decitabine compared with control mesenchymal cells (Table 1), but the slight increase in metastasis indicates that MET and modest increases in E-cadherin expression may promote metastatic colonies. However, a histological analysis of lung metastases indicates that the underlying mechanism of metastatic colonization may not be mediated by restoration of E-cadherin expression as the lung metastases demonstrate sustained loss of E-cadherin in the mesenchymal miR-200b group (Supplementary Figure 10). Although these findings challenge the paradigm of MET and increased E-cadherin expression as requirements for metastatic colonization,7, 37 our results are consistent with a recent investigation in breast cancer, in which E-cadherin is not essential for determining metastatic potential.18
In epithelial cells, E-cadherin is an important cell-to-cell adhesion molecule. E-cadherin promoter methylation is reported in EMT, and in HCC, E-cadherin promoter methylation is associated with poor prognosis and metastatic disease.32, 34 In this current work, the E-cadherin promoter is hyper-methylated in mesenchymal cells. Further investigation is needed to assess whether E-cadherin expression can be used as a marker to evaluate invasion and metastatic potential in HCC. In this study, E-cadherin expression correlates with final outcome in the combination miR-200b and decitabine treatment group, which strongly supports E-cadherin as a marker to monitor EMT/MET and tumor metastasis. However, follow-up investigations demonstrate that upregulation of E-cadherin alone in primary mesenchymal tumors is not able to reverse invasive and metastatic phenotype. In addition, although suppression of E-cadherin does increase tumor local invasion, suppression of E-cadherin is not able to induce a metastatic phenotype after treatment with miR-200b and decitabine combination. Thus, we conclude that, in our model of liver cancer EMT, E-cadherin is a marker for cell migration and tumor invasion, but E-cadherin expression is not sufficient to evaluate metastatic potential.
Correlation of additional markers of liver epithelial cells, Cytokeratin-19 (Krt-19), which is primarily expressed in cholangiocytes, and HNF4α, primarily expressed in hepatocytes, may be used as alternative markers for HCC metastasis specifically. The downregulation of HNF4α is associated with HCC progression, loss of epithelial phenotype and metastasis.38 Forced expression of HNF4α promotes a less invasive, well-differentiated phenotype with slower growth.39 These reports are consistent with our findings that HNF4α is downregulated in mesenchymal cells and is significantly induced after combination of miR-200b expression and DNMT inhibition. Thus, broader MET and differentiation effects of miR-200b and DNMT inhibition may inhibit metastasis in an E-cadherin-independent mechanism. In future studies, we will investigate the specific mechanisms underlying the HNF4α upregulation after combination of miR-200b restoration and DNMT inhibition, as miR-200b and DNMT inhibitors have multiple targets and can have broad effects in driving differentiation.
In conclusion, we demonstrate that targeting metastasis in liver cancer may require a combination therapy with miR-200b expression and DNMT inhibition, which is not necessarily mediated by restoration of E-cadherin expression.
Materials and methods
Cell culture
Parental HCC tumor cells were isolated from Pten−/− mice. We obtained two cell lines from tumor tissues after serial transplantations and termed with ‘P1’ and ‘P2’, respectively. P2 cells were mixed with epithelial and mesenchymal cell population, and these two populations were separated into pooled cell lines, named P2E epithelial cells and P2M mesenchymal cells, as described.14 Pten−/− P2E epithelial and P2M mesenchymal cells were cultured in Dulbecco's modified Eagle medium:F12 at 1:1 medium (Mediatech, Manassas, VA, USA) supplemented with 10% fetal bovine serumFBS, as described.14
Antibodies and Plasmid Constructs
The polyclonal E-cadherin antibody for western blot, immunohistochemistry and immunofluorescence, and polyclonal Zeb1 antibody for western blot were purchased from Cell Signaling (Cat# 3195s and 3396s, respectively, Danvers, MA, USA), and Zeb1 antibody for immunohistochemistry and Zeb2 antibody for immunohistochemistry and western blot were purchased from Novus Biologicals (Littleton, CO, USA) with catalog numbers NBP1-05987 and NBP1-67782, respectively. Retroviral construct for mouse E-cadherin was obtained from Addgene (Cambridge, MA, USA) with product ID#18804 and was developed by the Weinberg laboratory (Cambridge, MA, USA).40 The mouse E-cadherin shRNA (Ecad-shRNA) was developed using Invivogen (San Diego, CA, USA) psiRNA-h7SKzeo G1 vector (Cat# ksirna3-z21) by inserting a target sequence 5′-GGAGATGCAGAATAATTAT-3′ to specifically knockdown mouse E-cadherin, and the psiRNA-h7SKzeo G1 empty vector was used as a NC-shRNA. pEGFP-C1-Zeb1 construct was kindly provided by Dr K Morishita from the University of Miyazaki, Japan.41
Animals
Nude mice were fed ad libitum (Harlan Teklad-irradiated mouse diet 7912, Madison, WI, USA) and housed in a temperature-controlled animal facility with a 12-h light/dark cycle. All procedures were in compliance with our institution's guidelines for the use of laboratory animals and approved by the Institutional Animal Care and Use Committee.
Real-time PCR for mRNA and miRNA
Total RNA was extracted using Trizol (Invitrogen, Carlsbad, CA, USA). RNA was quantified with ND-1000 spectrophotometer (Nanodrop, Wilmington, DE, USA) and cDNA was generated. Real-time PCR experiments were conducted as described previously.42 The housekeeping gene GAPDH and β-actin were used for ΔΔCt calculations. Relative expression was calculated for mouse E-cadherin, Zeb1, Zeb2, HNF4α and Krt-19, using assay-on-demand primer/probe sets (Applied Biosystems, Foster City, CA, USA). For miRNA real-time PCR, miRNA species were isolated using mirVara miRNA isolation kit from Applied Biosystems, and cDNA amplified using primers for U6, miR-200 family members, miR-205 and miR-16 from Applied Biosystems, and data were normalized using U6 for ΔΔCt calculations.
Bisulfite DNA C-T conversion and bisulfite DNA sequencing
Five hundred nanogram of mouse genomic DNA was treated with bisulfite at 60 °C water bath for 16 h, and DNA after bisulfite treatment was purified using the EZ DNA methylation kit according to manufacturer's instruction (Zymo Research, Irvine, CA, USA). The methylated CpG sites of E-cadherin were confirmed by DNA amplification and bisulfite sequencing using the following primers: 5′-GTTAGAGTATAGTTAGGTTAGGA-3′ (sense) and 5′-CTACAACACCAAACTCCCATA-3′ (antisense), which amplify a 370-bp fragment. The PCR cycling conditions were 95 °C for 4 min followed by 35 cycles with 95 °C for 30 s, 55 °C for 30 s and 72 °C for 1 min, and finally 72 °C for 10 min. The PCR products were loaded on a 2% agarose gel and visualized with ethidium bromide, and purified using QIAGEN (Valencia, CA, USA) QIAquick gel extraction kit for DNA sequencing.
Tet-on stable miRNA system
Mir-X inducible pmRi-ZsGreen1 miRNA system was purchased from Clontech (Mountain View, CA, USA), and the construct expressing miR-200b was developed by PCR strategy using the following PCR primers: upstream 5′-TAGTTCTCGAGGATCCT ATAAGACCCCCATCCTAAT-3′, and downstream 5′-CTCCAAAGCTAAGCTTGCCTGGGTATTGTATGCAGAA-3′. Constructs were verified by DNA sequencing. The stable miRNA system was developed as described in the manufacturer's protocol. Pooled mesenchymal cells with stable miR-200b or miR-NC expression were isolated per manufactures instructions.
Decitabine treatment
Tumor cells were treated with either vehicle (DMSO) or d1 μM of decitabine (Sigma-Aldrich, St Louis, MO, USA) for indicated times.
Orthotopic transplantation
Five days before tumor inoculation, nude mice were fed with doxycycline chow (200 mg/kg) purchased from Bio-Serv (Frenchtown, NJ, USA). Mice were anaesthetized with 100 mg/kg of ketamine and 10 mg/kg of xylazine, and the liver was exposed through a surgical incision. In all, 3 × 105 cells in 30 μl of 1:1 Matrigel:media were orthotopically inoculated. Pressure was applied to the injection site for 2 min. Abdominal wound was closed using wound clips after tumor cell injection.14
Bioluminescent image collection
Nude mice were anesthetized at defined times post cell transplant, and 150 mg/kg of potassium D-luciferin (Gold Biotechnology, St Louis, MO, USA) was injected intraperitoneal injected. Ten minutes after luciferin injection, bioluminescence images were collected using a Xenogen IVIS 50 system per manufacturer's instruction.
Invasion scoring system
The pancreas and colon were each graded based on the highest extent of tumor invasion on H&E stained slides (two independent sections at different levels more than 20 μm apart/organ). Extensive invasion equals a score of 2 points; for minor invasion, the score will be 1 point; and if there is no invasion, the score will be 0. The average invasion score for each experimental group will be the total points divide the number of animals.
Microscopy
Histology images were viewed with an Olympus CKX41 microscope (Olympus, Melville, NY, USA) using × 40 phase objective lens and ACDSee 3.1 (SR-1) software (ACD Systems, Seattle, WA, USA), and wound-healing migration images and immunofluorescent images were collected with Nikon eclipse TE2000-5 microscope (Nikon, Melville, NY, USA) using × 4 objective lens and × 40/0.60 Plan Fluor objective lens, respectively, and processed with MagnaFIRE SP software (Optronics, Goleta, CA, USA).
Statistical analyses
Paired two-tailed Student's t-test was used when comparing two groups. A P value <0.05 was considered statistically significant. Ordinal logistic regression was used to compare the invasion score and determine significance between treatment groups.
Abbreviations
- DMEM:
-
Dulbecco's modified Eagle medium
- DNMT:
-
DNA methyltransferase
- EMT:
-
epithelial-to-mesenchymal transition
- FBS:
-
fetal bovine serum
- GFP:
-
green fluorescence protein
- IF:
-
immunofluorescence
- IHC:
-
immunohistochemistry
- HCC:
-
hepatocellular carcinoma
- HNF4α:
-
Hepatocyte Nuclear Factor 4α
- MET:
-
mesenchymal-to-epithelial transition
- PI3K:
-
phosphoinositide 3-kinase
- PTEN:
-
phosphatase and tensin homolog deleted on chromosome 10.
References
El-Serag HB, Rudolph KL Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology, 2007 132: 2557–2576.
Kanwal F, Hoang T, Kramer JR, Asch SM, Goetz MB, Zeringue A et al. Increasing Prevalence of HCC and Cirrhosis in Patients With Chronic Hepatitis C Virus Infection. Gastroenterology, 2011 140: 1182–8 e1.
Carr BI Hepatocellular carcinoma: current management and future trends. Gastroenterology, 2004 127 (5 Suppl 1): S218–S224.
Whittaker S, Marais R, Zhu AX The role of signaling pathways in the development and treatment of hepatocellular carcinoma. Oncogene, 2010 29: 4989–5005.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell, 2011 144: 646–674.
Yang J, Weinberg RA Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell, 2008 14: 818–829.
Kalluri R, Weinberg RA The basics of epithelial-mesenchymal transition. J Clin Invest, 2009 119: 1420–1428.
Nieto MA Epithelial-Mesenchymal Transitions in development and disease: old views and new perspectives. Int J Dev Biol, 2009 53: 1541–1547.
Acloque H, Adams MS, Fishwick K, Bronner-Fraser M, Nieto MA Epithelial-mesenchymal transitions: the importance of changing cell state in development and disease. J Clin Invest, 2009 119: 1438–1449.
Perez-Moreno MA, Locascio A, Rodrigo I, Dhondt G, Portillo F, Nieto MA et al. A new role for E12/E47 in the repression of E-cadherin expression and epithelial-mesenchymal transitions. J Biol Chem, 2001 276: 27424–27431.
Gao ZH, Tretiakova MS, Liu WH, Gong C, Farris PD, Hart J Association of E-cadherin, matrix metalloproteinases, and tissue inhibitors of metalloproteinases with the progression and metastasis of hepatocellular carcinoma. Mod Pathol, 2006 19: 533–540.
Yang MH, Chen CL, Chau GY, Chiou SH, Su CW, Chou TY et al. Comprehensive analysis of the independent effect of twist and snail in promoting metastasis of hepatocellular carcinoma. Hepatology, 2009 50: 1464–1474.
Hu L, Lau SH, Tzang CH, Wen JM, Wang W, Xie D et al. Association of Vimentin overexpression and hepatocellular carcinoma metastasis. Oncogene, 2004 23: 298–302.
Ding W, You H, Dang H, Leblanc F, Galicia V, Lu SC et al. Epithelial-to-mesenchymal transition of murine liver tumor cells promotes invasion. Hepatology, 2010 52: 945–953.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol, 2008 10: 593–601.
Park SM, Gaur AB, Lengyel E, Peter ME The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev, 2008 22: 894–907.
Dykxhoorn DM, Wu Y, Xie H, Yu F, Lal A, Petrocca F et al. miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS One, 2009 4: e7181.
Korpal M, Ell BJ, Buffa FM, Ibrahim T, Blanco MA, Celia-Terrassa T et al. Direct targeting of Sec23a by miR-200s influences cancer cell secretome and promotes metastatic colonization. Nat Med, 2011 17: 1101–1108.
Korpal M, Kang Y The emerging role of miR-200 family of microRNAs in epithelial-mesenchymal transition and cancer metastasis. RNA Biol, 2008 5: 115–119.
Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res, 2008 68: 7846–7854.
Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep, 2008 9: 582–589.
Korpal M, Lee ES, Hu G, Kang Y The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem, 2008 283: 14910–14914.
Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T et al. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene, 2006 25: 2537–2545.
Budhu A, Jia HL, Forgues M, Liu CG, Goldstein D, Lam A et al. Identification of metastasis-related microRNAs in hepatocellular carcinoma. Hepatology, 2008 47: 897–907.
Garinis GA, Menounos PG, Spanakis NE, Papadopoulos K, Karavitis G, Parassi I et al. Hypermethylation-associated transcriptional silencing of E-cadherin in primary sporadic colorectal carcinomas. J Pathol, 2002 198: 442–449.
Wheeler JM, Kim HC, Efstathiou JA, Ilyas M, Mortensen NJ, Bodmer WF Hypermethylation of the promoter region of the E-cadherin gene (CDH1) in sporadic and ulcerative colitis associated colorectal cancer. Gut, 2001 48: 367–371.
Li LC, Zhao H, Nakajima K, Oh BR, Ribeiro Filho LA, Carroll P et al. Methylation of the E-cadherin gene promoter correlates with progression of prostate cancer. J Urol, 2001 166: 705–709.
Chen CL, Liu SS, Ip SM, Wong LC, Ng TY, Ngan HY E-cadherin expression is silenced by DNA methylation in cervical cancer cell lines and tumours. Eur J Cancer, 2003 39: 517–523.
Nass SJ, Herman JG, Gabrielson E, Iversen PW, Parl FF, Davidson NE et al. Aberrant methylation of the estrogen receptor and E-cadherin 5' CpG islands increases with malignant progression in human breast cancer. Cancer Res, 2000 60: 4346–4348.
Kanai Y, Ushijima S, Hui AM, Ochiai A, Tsuda H, Sakamoto M et al. The E-cadherin gene is silenced by CpG methylation in human hepatocellular carcinomas. Int J Cancer, 1997 71: 355–359.
Zhang CY, Mao L, Li L, Tian Z, Zhou XJ, Zhang ZY et al. Promoter methylation as a common mechanism for inactivating E-cadherin in human salivary gland adenoid cystic carcinoma. Cancer, 2007 110: 87–95.
Kwon GY, Yoo BC, Koh KC, Cho JW, Park WS, Park CK Promoter methylation of E-cadherin in hepatocellular carcinomas and dysplastic nodules. J Korean Med Sci, 2005 20: 242–247.
Kim J, Hong SJ, Park JY, Park JH, Yu YS, Park SY et al. Epithelial-mesenchymal transition gene signature to predict clinical outcome of hepatocellular carcinoma. Cancer Sci, 2010 101: 1521–1528.
Lee TK, Poon RT, Yuen AP, Ling MT, Kwok WK, Wang XH et al. Twist overexpression correlates with hepatocellular carcinoma metastasis through induction of epithelial-mesenchymal transition. Clin Cancer Res, 2006 12: 5369–5376.
van Zijl F, Zulehner G, Petz M, Schneller D, Kornauth C, Hau M et al. Epithelial-mesenchymal transition in hepatocellular carcinoma. Future Oncol, 2009 5: 1169–1179.
Gibbons DL, Lin W, Creighton CJ, Rizvi ZH, Gregory PA, Goodall GJ et al. Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression. Genes Dev, 2009 23: 2140–2151.
Thiery JP, Acloque H, Huang RY, Nieto MA Epithelial-mesenchymal transitions in development and disease. Cell, 2009 139: 871–890.
Lazarevich NL, Shavochkina DA, Fleishman DI, Kustova IF, Morozova OV, Chuchuev ES et al. Deregulation of hepatocyte nuclear factor 4 (HNF4)as a marker of epithelial tumors progression. Exp Oncol, 2010 32: 167–171.
Lazarevich NL, Cheremnova OA, Varga EV, Ovchinnikov DA, Kudrjavtseva EI, Morozova OV et al. Progression of HCC in mice is associated with a downregulation in the expression of hepatocyte nuclear factors. Hepatology, 2004 39: 1038–1047.
Onder TT, Gupta PB, Mani SA, Yang J, Lander ES, Weinberg RA Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res, 2008 68: 3645–3654.
Nakahata S, Yamazaki S, Nakauchi H, Morishita K Downregulation of ZEB1 and overexpression of Smad7 contribute to resistance to TGF-beta1-mediated growth suppression in adult T-cell leukemia/lymphoma. Oncogene, 2010 29: 4157–4169.
Ding W, Mouzaki M, You H, Laird JC, Mato J, Lu SC et al. CD133+ liver cancer stem cells from methionine adenosyl transferase 1A-deficient mice demonstrate resistance to transforming growth factor (TGF)-beta-induced apoptosis. Hepatology, 2009 49: 1277–1286.
Acknowledgements
We thank Dr Yvette Liu, from the El-Deiry Laboratory, The Penn State Hershey Cancer Institute, for assisting with luciferase bioluminescence image collection. We thank Dr K Morishita, Department of Medical Sciences, University of Miyazaki, for providing us with pEGFP-C1-Zeb1 construct. We acknowledge Drs Kent Vrana and Willard Freeman of the Functional Genomics Core (The Pennsylvania State University College of Medicine). Important Functional Genomics Core Facility instrumentation purchases were made possible through Tobacco Settlement Funds and through a Penn State Cancer Institute contract with the Department of the Navy. This publication was made possible by generous support from the National Institute of Health, R03DK088013 (CBR); the American Cancer Society, Research Scholar Award, RSG-10-073-01-TBG (CBR); The Department of the Navy, Penn State Center for Pharmacogenetics Pilot Projects (CBR) and the Four Diamonds Foundation (CBR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Dr CB Rountree declares a small research grant (less than $10,000), which does not include direct salary support, from Bayer Pharmaceuticals. The remaining authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogenesis website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Ding, W., Dang, H., You, H. et al. miR-200b restoration and DNA methyltransferase inhibitor block lung metastasis of mesenchymal-phenotype hepatocellular carcinoma. Oncogenesis 1, e15 (2012). https://doi.org/10.1038/oncsis.2012.15
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/oncsis.2012.15
Keywords
This article is cited by
-
Non-homogeneous dispersion of graphene in polyacrylonitrile substrates induces a migrastatic response and epithelial-like differentiation in MCF7 breast cancer cells
Cancer Nanotechnology (2022)
-
Dynamics and plasticity of the epithelial to mesenchymal transition induced by miR-200 family inhibition
Scientific Reports (2016)
-
Roles and epigenetic regulation of epithelial–mesenchymal transition and its transcription factors in cancer initiation and progression
Cellular and Molecular Life Sciences (2016)
-
Combinatorial interventions inhibit TGFβ-driven epithelial-to-mesenchymal transition and support hybrid cellular phenotypes
npj Systems Biology and Applications (2015)
-
Dynamic imbalance between cancer cell subpopulations induced by Transforming Growth Factor Beta (TGF-β) is associated with a DNA methylome switch
BMC Genomics (2014)