Abstract
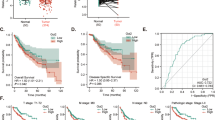
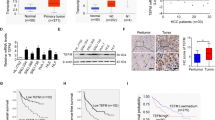
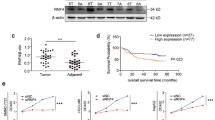
Hepatocellular carcinoma (HCC) is a frequent form of cancer with a poor prognosis and with limited possibilities of medical intervention. It has been shown that over 100 putative driver genes are associated with multiple recurrently altered pathways in HCC, suggesting that multiple pathways will need to be inhibited for any therapeutic method. mRNA processing is regulated by a complex RNA–protein network that is essential for the maintenance of homeostasis. THOC5, a member of mRNA export complex, has a role in less than 1% of mRNA processing, and is required for cell growth and differentiation, but not for cell survival in normal fibroblasts, hepatocytes and macrophages. In this report, we show that 50% depletion of THOC5 in human HCC cell lines Huh7 and HepG2 induced apoptosis. Transcriptome analysis using THOC5-depleted cells revealed that 396 genes, such as transmembrane BAX inhibitor motif containing 4 (TMBIM4), transmembrane emp24-like trafficking protein 10 (Tmed10) and D-tyrosyl-tRNA deacylase 2 (Dtd2) genes were downregulated in both cell lines. The depletion of one of these THOC5 target genes in Huh7 or HepG2 did not significantly induce cell death, suggesting that these may be fine tuners for HCC cell survival. However, the depletion of a combination of these genes synergistically increased the number of TUNEL (terminal deoxynucleotidyl transferase dUTP nick end labeling)-positive HCC. It must be noted that the depletion of these genes did not induce cell death in the hepatocyte cell line, THLE-2 cells. THOC5 expression was enhanced in 78% of cytological differentiation grading G2 and G3 tumor in primary HCC. Furthermore, the expression of a putative glycoprotein, Tmed10, is correlated to THOC5 expression level in primary HCCs, suggesting that this protein may be a novel biomarker for HCC. These data imply that the suppression of the multiple THOC5 target genes may represent a novel strategy for HCC therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D . Global cancer statistics. CA Cancer J Clin 2011; 61: 69–90.
Whittaker S, Marais R, Zhu AX . The role of signaling pathways in the development and treatment of hepatocellular carcinoma. Oncogene 2010; 29: 4989–5005.
Ferrin G, Aguilar-Melero P, Rodriguez-Peralvarez M, Montero-Alvarez JL, de la Mata M . Biomarkers for hepatocellular carcinoma: diagnostic and therapeutic utility. Hepat Med 2015; 7: 1–10.
Schulze K, Imbeaud S, Letouze E, Alexandrov LB, Calderaro J, Rebouissou S et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat Genet 2015; 47: 505–511.
Masuda S, Das R, Cheng H, Hurt E, Dorman N, Reed R . Recruitment of the human TREX complex to mRNA during splicing. Genes Dev 2005; 19: 1512–1517.
Rehwinkel J, Herold A, Gari K, Kocher T, Rode M, Ciccarelli FL et al. Genome-wide analysis of mRNAs regulated by the THO complex in Drosophila melanogaster. Nat Struct Mol Biol 2004; 11: 558–566.
Tran DD, Koch A, Tamura T . THOC5, a member of the mRNA export complex: a novel link between mRNA export machinery and signal transduction pathways in cell proliferation and differentiation. Cell Commun Signal 2014; 12: 3.
Chavez S, Garcia-Rubio M, Prado F, Aguilera A . Hpr1 is preferentially required for transcription of either long or G+C-rich DNA sequences in Saccharomyces cerevisiae. Mol Cell Biol 2001; 21: 7054–7064.
Jimeno S, Aguilera A . The THO complex as a key mRNP biogenesis factor in development and cell differentiation. J Biol 2010; 9: 6.
Jimeno S, Rondon AG, Luna R, Aguilera A . The yeast THO complex and mRNA export factors link RNA metabolism with transcription and genome instability. EMBO J 2002; 21: 3526–3535.
Tran DD, Saran S, Williamson AJ, Pierce A, Dittrich-Breiholz O, Wiehlmann L et al. THOC5 controls 3'end-processing of immediate early genes via interaction with polyadenylation specific factor 100 (CPSF100). Nucleic Acids Res 2014; 42: 12249–12260.
Mancini A, Niemann-Seyde SC, Pankow R, El Bounkari O, Klebba-Farber S, Koch A et al. THOC5/FMIP, an mRNA export TREX complex protein, is essential for hematopoietic primitive cell survival in vivo. BMC Biol 2010; 8: 1.
Guria A, Tran DD, Ramachandran S, Koch A, El Bounkari O, Dutta P et al. Identification of mRNAs that are spliced but not exported to the cytoplasm in the absence of THOC5 in mouse embryo fibroblasts. RNA 2011; 17: 1048–1056.
Tran DD, Saran S, Dittrich-Breiholz O, Williamson AJ, Klebba-Farber S, Koch A et al. Transcriptional regulation of immediate-early gene response by THOC5, a member of mRNA export complex, contributes to the M-CSF-induced macrophage differentiation. Cell Death Dis 2013; 4: e879.
Tamura T, Mancini A, Joos H, Koch A, Hakim C, Dumanski J et al. FMIP, a novel Fms-interacting protein, affects granulocyte/macrophage differentiation. Oncogene 1999; 18: 6488–6495.
Pierce A, Carney L, Hamza HG, Griffiths JR, Zhang L, Whetton BA et al. THOC5 spliceosome protein: a target for leukaemogenic tyrosine kinases that affects inositol lipid turnover. Br J Haematol 2008; 141: 641–650.
Griaud F, Pierce A, Gonzalez Sanchez MB, Scott M, Abraham SA, Holyoake TL et al. A pathway from leukemogenic oncogenes and stem cell chemokines to RNA processing via THOC5. Leukemia 2013; 27: 932–940.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2012; 2: 401–404.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013; 6: pl1.
Panagopoulos I, Bjerkehagen B, Gorunova L, Berner JM, Boye K, Heim S . Several fusion genes identified by whole transcriptome sequencing in a spindle cell sarcoma with rearrangements of chromosome arm 12q and MDM2 amplification. Int J Oncol 2014; 45: 1829–1836.
Gubser C, Bergamaschi D, Hollinshead M, Lu X, van Kuppeveld FJ, Smith GL . A new inhibitor of apoptosis from vaccinia virus and eukaryotes. PLoS Pathog 2007; 3: e17.
Mancini A, Koch A, Whetton AD, Tamura T . The M-CSF receptor substrate and interacting protein FMIP is governed in its subcellular localization by protein kinase C-mediated phosphorylation, and thereby potentiates M-CSF-mediated differentiation. Oncogene 2004; 23: 6581–6589.
Corkery DP, Holly AC, Lahsaee S, Dellaire G . Connecting the speckles: Splicing kinases and their role in tumorigenesis and treatment response. Nucleus 2015; 6: 279–288.
Maguire SL, Leonidou A, Wai P, Marchio C, Ng CK, Sapino A et al. SF3B1 mutations constitute a novel therapeutic target in breast cancer. J Pathol 2015; 235: 571–580.
Mancini A, El Bounkari O, Norrenbrock AF, Scherr M, Schaefer D, Eder M et al. FMIP controls the adipocyte lineage commitment of C2C12 cells by downmodulation of C/EBP alpha. Oncogene 2007; 26: 1020–1027.
Koch A, Saran S, Tran D, Klebba-Farber S, Thiesler H, Sewald K et al. Murine precision-cut liver slices (PCLS): a new tool for studying tumor microenvironments and cell signaling ex vivo. Cell Commun Signal 2014; 12: 73.
Saran S, Tran DD, Klebba-Farber S, Moran-Losada P, Wiehlmann L, Koch A et al. THOC5, a member of the mRNA export complex, contributes to processing of a subset of wingless/integrated (Wnt) target mRNAs and integrity of the gut epithelial barrier. BMC Cell Biol 2013; 14: 51.
Acknowledgements
We thank C Bruce Boschek for critically reading the manuscript and Iris Albers and Annika Hamm for technical assistance. SS and DDHT contributed equally to this work. This research was supported by Deutsche Krebshilfe (111153), DFG Ta-111/13-1, and PhD program Molecular Medicine and Leistungsorientierte Mittelvergabe with Frauenfaktor from MHH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Saran, S., Tran, D., Ewald, F. et al. Depletion of three combined THOC5 mRNA export protein target genes synergistically induces human hepatocellular carcinoma cell death. Oncogene 35, 3872–3879 (2016). https://doi.org/10.1038/onc.2015.433
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.433
This article is cited by
-
C20orf204, a hepatocellular carcinoma-specific protein interacts with nucleolin and promotes cell proliferation
Oncogenesis (2021)
-
Myc/Max dependent intronic long antisense noncoding RNA, EVA1A-AS, suppresses the expression of Myc/Max dependent anti-proliferating gene EVA1A in a U2 dependent manner
Scientific Reports (2019)
-
Myc target gene, long intergenic noncoding RNA, Linc00176 in hepatocellular carcinoma regulates cell cycle and cell survival by titrating tumor suppressor microRNAs
Oncogene (2018)