Abstract
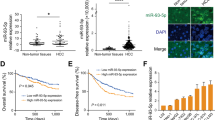
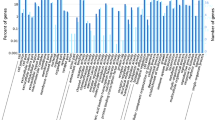
MicroRNAs (miRNAs) have an important role in a wide range of physiological and pathological processes, and their dysregulation has been reported to affect the development and progression of cancers, including hepatocellular carcinoma (HCC). However, in the plethora of dysregulated miRNAs, it is largely unknown which of them have a causative role in the hepatocarcinogenic process. In the present study, we first aimed to determine changes in the expression profile of miRNAs in human HCCs and to compare them with liver tumors generated in a rat model of chemically induced HCC. We found that members of the miR-100 family (miR-100, miR-99a) were downregulated in human HCCs; a similar downregulation was also observed in rat HCCs. Their reduction was paralleled by an increased expression of polo like kinase 1 (PLK1), a target of these miRNAs. The introduction of miR-100 in HCC cells impaired their growth ability and their capability to form colonies in soft agar. Next, we aimed at investigating, in the same animal model, if dysregulation of miR-100 and PLK1 is an early or late event along the multistep process of hepatocarcinogenesis. The obtained results showed that miR-100 downregulation (i) is already evident in very early preneoplastic lesions generated 9 weeks after carcinogenic treatment; (ii) is also observed in adenomas and early HCCs; and (iii) is not simply a marker of proliferating hepatocytes. To our knowledge, this is the first work unveiling the role of a miRNA family along HCC progression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Andersen JB, Loi R, Perra A, Factor VM, Ledda-Columbano GM, Columbano A et al. (2010). Progenitor-derived hepatocellular carcinoma model in the rat. Hepatology 51: 1401–1409.
Cairo S, Wang Y, de RA, Duroure K, Dahan J, Redon MJ et al. (2010). Stem cell-like micro-RNA signature driven by Myc in aggressive liver cancer. Proc Natl Acad Sci USA 107: 20471–20476.
Christodoulou F, Raible F, Tomer R, Simakov O, Trachana K, Klaus S et al. (2010). Ancient animal microRNAs and the evolution of tissue identity. Nature 463: 1084–1088.
De Petro G, Tavian D, Copeta A, Portolani N, Giulini SM, Barlati S . (1998). Expression of urokinase-type plasminogen activator (u-PA), u-PA receptor, and tissue-type PA messenger RNAs in human hepatocellular carcinoma. Cancer Res 58: 2234–2239.
Eckerdt F, Yuan J, Strebhardt K . (2005). Polo-like kinases and oncogenesis. Oncogene 24: 267–276.
Enomoto K, Farber E . (1982). Kinetics of phenotypic maturation of remodeling of hyperplastic nodules during liver carcinogenesis. Cancer Res 42: 2330–2335.
Fu L, Medico E . (2007). FLAME, a novel fuzzy clustering method for the analysis of DNA microarray data. BMC Bioinformatics 8: 3.
Henson BJ, Bhattacharjee S, O'Dee DM, Feingold E, Gollin SM . (2009). Decreased expression of miR-125b and miR-100 in oral cancer cells contributes to malignancy. Genes Chromosomes Cancer 48: 569–582.
Hou J, Lin L, Zhou W, Wang Z, Ding G, Dong Q et al. (2011). Identification of miRNomes in human liver and hepatocellular carcinoma reveals miR-199a/b-3p as therapeutic target for hepatocellular carcinoma. Cancer Cell 19: 232–243.
Imbeaud S, Ladeiro Y, Zucman-Rossi J . (2010). Identification of novel oncogenes and tumor suppressors in hepatocellular carcinoma. Semin Liver Dis 30: 75–86.
Kota J, Chivukula RR, O'Donnell KA, Wentzel EA, Montgomery CL, Hwang HW et al. (2009). Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137: 1005–1017.
Ledda-Columbano GM, Curto M, Piga R, Zedda AI, Menegazzi M, Sartori C et al. (1998). In vivo hepatocyte proliferation is inducible through a TNF and IL-6-independent pathway. Oncogene 17: 1039–1044.
Lee EJ, Gusev Y, Jiang J, Nuovo GJ, Lerner MR, Frankel WL et al. (2007). Expression profiling identifies microRNA signature in pancreatic cancer. Int J Cancer 120: 1046–1054.
Lee JS, Thorgeirsson SS . (2004). Genome-scale profiling of gene expression in hepatocellular carcinoma: classification, survival prediction, and identification of therapeutic targets. Gastroenterology 127: S51–S55.
Leite KR, Sousa-Canavez JM, Reis ST, Tomiyama AH, Camara-Lopes LH, Sanudo A et al. (2009). Change in expression of miR-let7c, miR-100, and miR-218 from high grade localized prostate cancer to metastasis. Urol Oncol 29: 265–269.
Liu W, Gong YH, Chao TF, Peng XZ, Yuan JG, Ma ZY et al. (2009). Identification of differentially expressed microRNAs by microarray: a possible role for microRNAs gene in medulloblastomas. Chin Med J (Engl) 122: 2405–2411.
Lu J, Getz G, Miska EA, varez-Saavedra E, Lamb J, Peck D et al. (2005). MicroRNA expression profiles classify human cancers. Nature 435: 834–838.
Nam EJ, Yoon H, Kim SW, Kim H, Kim YT, Kim JH et al. (2008). MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 14: 2690–2695.
Oneyama C, Ikeda J, Okuzaki D, Suzuki K, Kanou T, Shintani Y et al. (2011). MicroRNA-mediated downregulation of mTOR/FGFR3 controls tumor growth induced by Src-related oncogenic pathways. Oncogene 30: 3489–3501.
Parkin DM . (2001). Global cancer statistics in the year 2000. Lancet Oncol 2: 533–543.
Pellegrino R, Calvisi DF, Ladu S, Ehemann V, Staniscia T, Evert M et al. (2010). Oncogenic and tumor suppressive roles of polo-like kinases in human hepatocellular carcinoma. Hepatology 51: 857–868.
Satoh K, Kitahara A, Soma Y, Inaba Y, Hatayama I, Sato K . (1985). Purification, induction, and distribution of placental glutathione transferase: a new marker enzyme for preneoplastic cells in the rat chemical hepatocarcinogenesis. Proc Natl Acad Sci USA 82: 3964–3968.
Shi W, Alajez NM, Bastianutto C, Hui AB, Mocanu JD, Ito E et al. (2010). Significance of Plk1 regulation by miR-100 in human nasopharyngeal cancer. Int J Cancer 126: 2036–2048.
Smith MR, Wilson ML, Hamanaka R, Chase D, Kung H, Longo DL et al. (1997). Malignant transformation of mammalian cells initiated by constitutive expression of the polo-like kinase. Biochem Biophys Res Commun 234: 397–405.
Solt DB, Medline A, Farber E . (1977). Rapid emergence of carcinogen-induced hyperplastic lesions in a new model for the sequential analysis of liver carcinogenesis. Am J Pathol 88: 595–618.
Su H, Yang JR, Xu T, Huang J, Xu L, Yuan Y et al. (2009). MicroRNA-101, down-regulated in hepatocellular carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer Res 69: 1135–1142.
Tarantino C, Paolella G, Cozzuto L, Minopoli G, Pastore L, Parisi S et al. (2010). miRNA 34a, 100, and 137 modulate differentiation of mouse embryonic stem cells. FASEB J 24: 3255–3263.
Thorgeirsson SS . (2011). The almighty MYC: orchestrating the micro-RNA universe to generate aggressive liver cancer. J Hepatol 55: 486–487.
Ueda T, Volinia S, Okumura H, Shimizu M, Taccioli C, Rossi S et al. (2010). Relation between microRNA expression and progression and prognosis of gastric cancer: a microRNA expression analysis. Lancet Oncol 11: 136–146.
Varnholt H, Drebber U, Schulze F, Wedemeyer I, Schirmacher P, Dienes HP et al. (2008). MicroRNA gene expression profile of hepatitis C virus-associated hepatocellular carcinoma. Hepatology 47: 1223–1232.
Wang FZ, Weber F, Croce C, Liu CG, Liao X, Pellett PE . (2008). Human cytomegalovirus infection alters expression of cellular microRNA species that affect its replication. J Virol 82: 9065–9074.
Wienholds E, Kloosterman WP, Miska E, varez-Saavedra E, Berezikov E, de BE et al. (2005). MicroRNA expression in zebrafish embryonic development. Science 309: 310–311.
Wong TS, Liu XB, Wong BY, Ng RW, Yuen AP, Wei WI . (2008). Mature miR-184 as potential oncogenic microRNA of squamous cell carcinoma of tongue. Clin Cancer Res 14: 2588–2592.
Acknowledgements
We thank L Tamagnone, S Corso, MM Angioni and our colleagues for helpful discussions; C Manca for the preparation of rat samples and C Isella for statistical analysis; L Palmas for technical assistance. Work was supported by the Associazione Italiana per la Ricerca sul Cancro (AIRC grants to SG, AC), Regione Piemonte (grant to SG) and MIUR (grants to SG, AC, GMLC, GDP).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Petrelli, A., Perra, A., Schernhuber, K. et al. Sequential analysis of multistage hepatocarcinogenesis reveals that miR-100 and PLK1 dysregulation is an early event maintained along tumor progression. Oncogene 31, 4517–4526 (2012). https://doi.org/10.1038/onc.2011.631
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.631
Keywords
This article is cited by
-
CircCASC15-miR-100-mTOR may influence the cervical cancer radioresistance
Cancer Cell International (2022)
-
HIF1A activates the transcription of lncRNA RAET1K to modulate hypoxia-induced glycolysis in hepatocellular carcinoma cells via miR-100-5p
Cell Death & Disease (2020)
-
MicroRNA miR-509 Regulates ERK1/2, the Vimentin Network, and Focal Adhesions by Targeting Plk1
Scientific Reports (2018)
-
Regulation of PLK1 through competition between hnRNPK, miR-149-3p and miR-193b-5p
Cell Death & Differentiation (2017)
-
PLK1-associated microRNAs are correlated with pediatric medulloblastoma prognosis
Child's Nervous System (2017)