Abstract
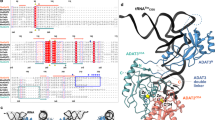
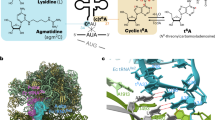
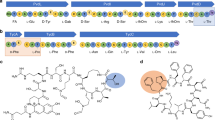
The crystal structure of Escherichia coli cysteinyl-tRNA synthetase (CysRS) bound to tRNACys at a resolution of 2.3 Å reveals base-specific and shape-selective interactions across an extensive protein-RNA recognition interface. The complex contains a mixed α/β C-terminal domain, which is disordered in the unliganded enzyme. This domain makes specific hydrogen bonding interactions with all three bases of the GCA anticodon. The tRNA anticodon stem is bent sharply toward the enzyme as compared with its conformation when bound to elongation factor Tu, providing an essential basis for shape-selective recognition. The CysRS structure also reveals interactions of conserved enzyme groups with the sugar-phosphate backbone in the D loop, adjacent to an unusual G15·G48 tertiary base pair previously implicated in tRNA aminoacylation. A combined mutational analysis of enzyme and tRNA groups at G15·G48 supports the notion that contacts between CysRS and the sugar-phosphate backbone contribute to recognition by indirect readout.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Seeman, N.C., Rosenberg, J.M. & Rich, A. Sequence-specific recognition of double helical nucleic acids by proteins. Proc. Natl. Acad. Sci. USA 73, 804–808 (1976).
Rhodes, D., Schwabe, J.W., Chapman, L. & Fairall, L. Towards an understanding of protein-DNA recognition. Philos. Trans. R. Soc. Lond. B Biol. Sci. 351, 501–509 (1996).
Gromiha, M., Siebers, J.G., Selvaraj, S., Kono, H. & Sarai, A. Intermolecular and intramolecular readout mechanisms in protein-DNA recognition. J. Mol. Biol. 337, 285–294 (2004).
Otwinowski, Z. et al. Crystal structure of trp repressor/operator complex at atomic resolution. Nature 335, 321–329 (1988).
Lesser, D.R., Kurpiewski, M.R., Waters, T., Connolly, B.A. & Jen-Jacobsen, L. Facilitated distortion of the DNA site enhances EcoRI endonuclease-DNA recognition. Proc. Natl. Acad. Sci. USA 90, 7548–7452 (1993).
Martin, A.M., Sam, M.D., Reich, N.O. & Perona, J.J. Structural and energetic origins of indirect readout in site-specific DNA cleavage by a restriction endonuclease. Nat. Struct. Biol. 6, 269–277 (1999).
Lynch, T.W., Read, E.K., Mattis, A.N., Gardner, J.F. & Rice, P.A. Integration host factor: putting a twist on protein-DNA recognition. J. Mol. Biol. 330, 493–502 (2003).
Ibba, M. & Soll, D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 69, 617–650 (2000).
Nissan, T.A. & Perona, J.J. Alternative designs for construction of the class II transfer RNA tertiary core. RNA 6, 1585–1596 (2000).
Avalos, J., Corrochano, L.M. & Brenner, S. Cysteinyl-tRNA synthetase is a direct descendent of the first aminoacyl-tRNA synthetase. FEBS Lett. 286, 176–180 (1991).
Eriani, G., Dirheimer, G. & Gangloff, J. Cysteinyl-tRNA synthetase: determination of the last E. coli aminoacyl-tRNA synthetase primary structure. Nucleic Acids Res. 19, 265–269 (1991).
Hou, Y.M., Shiba, K., Mottes, C. & Schimmel, P. Sequence determination and modeling of structural motifs for the smallest monomeric aminoacyl-tRNA synthetase. Proc. Natl. Acad. Sci. USA 88, 976–980 (1991).
Newberry, K.J., Hou, Y.M. & Perona, J.J. Structural origins of amino acid selection without editing by cysteinyl-tRNA synthetase. EMBO J. 21, 2778–2787 (2002).
Levitt, M. Detailed molecular model for transfer ribonucleic acid. Nature 224, 759–763 (1969).
Nissen, P., Thirup, S., Kjeldgaard, M. & Nyborg, J. The crystal structure of Cys-tRNACys-Ef-TU-GDPNP reveals general and specific features in the ternary complex and in tRNA. Structure Fold Des. 7, 143–156 (1999).
Hamann, C.S. & Hou, Y.M. An RNA structural determinant for tRNA recognition. Biochemistry 36, 7967–7972 (1997).
Hou, Y.M., Westof, E. & Giege, R. An unusual RNA tertiary interaction has a role for the specific aminoacylation of a transfer RNA. Proc. Natl. Acad. Sci. USA 90, 6776–6780 (1993).
Hou, Y.M., Sterner, T. & Bhalla, R. Evidence for a conserved relationship between an acceptor stem and a tRNA for aminoacylation. RNA 1, 707–712 (1995).
Lipman, R.S. & Hou, Y.M. Aminoacylation of tRNA in the evolution of an aminoacyl-tRNA synthetase. Proc. Natl. Acad. Sci. USA 95, 13495–13500 (1998).
Cavarelli, J., Delagoutte, B., Eriani, G., Gangloff, J. & Moras, D. L-arginine recognition by yeast arginyl-tRNA synthetase. EMBO J. 17, 5438–5448 (1998).
Cusack, S., Yaremchuk, A. & Tukalo, M. The 2 Å crystal structure of leucyl-tRNA synthetase and its complex with a leucyl-adenylate analogue. EMBO J. 19, 2351–2361 (2000).
Fukai, S. et al. Structural basis for double-sieve discrimination of L-valine from L-isoleucine and L-threonine by the complex of tRNA(Val) and valyl-tRNA synthetase. Cell 103, 793–803 (2000).
Silvian, L.F., Wang, J. & Steitz, T.A. Insights into editing from an ile-tRNA structure with tRNAIle and mupirocin. Science 285, 1074–1077 (1999).
Sugiura, I. et al. The 2.0 Å crystal structure of Thermus thermophilus methionyl-tRNA synthetase reveals two RNA-binding modules. Structure Fold Des. 8, 197–208 (2000).
Nakama, T., Nureki, O. & Yokoyama, S. Structural basis for the recognition of isoleucyl-adenylate and an antibiotic, mupirocin, by isoleucyl-tRNA synthetase. J. Biol. Chem. 276, 47387–47393 (2001).
Fukai, S. et al. Mechanism of molecular interactions for tRNA(Val) recognition by valyl-tRNA synthetase. RNA 9, 100–111 (2003).
Sherlin, L.D. & Perona, J.J. tRNA-dependent active site assembly in a class I aminoacyl-tRNA synthetase. Structure 11, 591–603 (2003).
Williamson, J.R. Induced fit in RNA-protein recognition. Nat. Struct. Biol. 7, 834–837 (2000).
Komatsoulis, G.A. & Abelson, J. Recognition of tRNA(Cys) by Escherichia coli cysteinyl-tRNA synthetase. Biochemistry 32, 7435–7444 (1993).
Burkard, M.E., Kierzek, R. & Turner, D.H. Thermodynamics of unpaired terminal nucleotides on short RNA helixes correlates with stacking at helix termini in larger RNAs. J. Mol. Biol. 290, 967–982 (1999).
Ming, X., Smith, K., Suga, H. & Hou, Y.M. Recognition of tRNA backbone for aminoacylation with cysteine: evolution from Escherichia coli to human. J. Mol. Biol. 318, 1207–1220 (2002).
Hamann, C.S. & Hou, Y.M. Probing a tRNA core that contributes to aminoacylation. J. Mol. Biol. 295, 777–789 (2000).
Hamann, C.S. & Hou, Y.M. A strategy of tRNA recognition that includes determinants of RNA structure. Bioorg. Med. Chem. 5, 1011–1019 (1997).
Hou, Y.M. Structural elements that contribute to an unusual tertiary interaction in a transfer RNA. Biochemistry 33, 4677–4681 (1994).
Christian, T., Lipman, R.S., Evilia, C. & Hou, Y.M. Alternative design of a tRNA core for aminoacylation. J. Mol. Biol. 303, 503–514 (2000).
Delagoutte, B., Moras, D. & Cavarelli, J. tRNA aminoacylation by arginyl-tRNA synthetase: induced conformations during substrates binding. EMBO J. 19, 5599–5610 (2000).
Fersht, A.R. & Dingwall, C. Cysteinyl-tRNA synthetase from Escherichia coli does not need an editing mechanism to reject serine and alanine. High energy of small groups in specific molecular interactions. Biochemistry 18, 1245–1249 (1979).
Zhang, C.M., Christian, T., Newberry, K.J., Perona, J.J. & Hou, Y.M. Zinc-mediated amino acid discrimination in cysteinyl-tRNA synthetase. J. Mol. Biol. 327, 911–917 (2003).
Zhang, C.M., Perona, J.J. & Hou, Y.M. Amino acid discrimination by a highly differentiated metal center of an aminoacyl-tRNA synthetase. Biochemistry 42, 10931–10937 (2003).
Brick, P., Bhat, T.N. & Blow, D.M. Structure of tyrosyl-tRNA synthetase refined at 2.3 Å resolution. Interaction of the enzyme with the tyrosyl adenylate intermediate. J. Mol. Biol. 208, 83–98 (1989).
Sekine, S. et al. ATP binding by glutamyl-tRNA synthetase is switched to the productive mode by tRNA binding. EMBO J. 22, 676–688 (2003).
Rould, M.A., Perona, J.J., Soll, D. & Steitz, T.A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA(Gln) and ATP at 2.8 Å resolution. Science 246, 1135–1142 (1989).
Yang, X.L. et al. Crystal structures that suggest late development of genetic code components for differentiating aromatic side chains. Proc. Natl. Acad. Sci. USA 100, 15376–15380 (2003).
Perona, J.J., Swanson, R.N., Rould, M.A., Steitz, T.A. & Soll, D. Structural basis for misaminoacylation by mutant E. coli glutaminyl-tRNA synthetase enzymes. Science 246, 1152–1154 (1989).
Doublie, S., Bricogne, G., Gilmore, C. & Carter, C.W. Jr. Tryptophanyl-tRNA synthetase crystal structure reveals an unexpected homology to tyrosyl-tRNA synthetase. Structure 3, 17–31 (1995).
Pallanck, L., Li, S. & Schulman, L.H. The anticodon and discriminator base are major determinants of cysteine tRNA identity in vivo. J. Biol. Chem. 267, 7221–7223 (1992).
Hamann, C.S. & Hou, Y.M. Enzymatic aminoacylation of tRNA acceptor stem helices with cysteine is dependent on a single nucleotide. Biochemistry 34, 6527–6532 (1995).
Biou, V., Yaremchuk, A., Tukalo, M. & Cusack, S. The 2.9 Å crystal structure T. thermophilus seryl-tRNA synthetase complexed with tRNA(Ser). Science 263, 1404–1410 (1994).
Sekine, S. et al. Major identity determinants in the “augmented D helix” of tRNA(Glu) from Escherichia coli. J. Mol. Biol. 256, 685–700 (1996).
Shimada, A., Nureki, O., Goto, M., Takahashi, S. & Yokoyama, S. Structural and mutational studies of the recognition of the arginine tRNA-specific major identity element, A20, by arginyl-tRNA synthetase. Proc. Natl. Acad. Sci. USA 98, 13537–13542 (2001).
Sherlin, L.D. et al. Influence of transfer RNA tertiary structure on aminoacylation efficiency by glutaminyl and cysteinyl-tRNA synthetases. J. Mol. Biol. 299, 431–446 (2000).
Giege, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998).
Grodberg, J.D., Jr. OmpT encodes the Escherichia coli outer membrane protease that cleaves T7 RNA polymerase during purification. J. Bacteriol. 170, 1245–1258 (1988).
Lyakhov, D. et al. Site-specific mutagenesis of the Lys-172 residue in phage T7 RNA polymerase: characterization of the transcriptional properties of mutant proteins. Mol. Biol. 26, 679–687 (1992).
Newberry, K.J., Kohn, J., Hou, Y.M. & Perona, J.J. Crystallization and preliminary diffraction analysis of Escherichia coli cysteinyl-tRNA synthetase. Acta Crystallogr. D 55, 1046–1047 (1999).
Sherlin, L.D. et al. Chemical and enzymatic synthesis of tRNAs for high-throughput crystallization. RNA 7, 1671–1678 (2001).
Bullock, T.L., Uter, N., Nissan, T.A. & Perona, J.J. Amino acid discrimination by a class I aminoacyl-tRNA synthetase specified by negative determinants. J. Mol. Biol. 328, 395–408 (2003).
Chen, Z. & Ruffner, D.E. Modified crush-and-soak method for recovering oligodeoxynucleotides from polyacrylamide gel. Biotechniques 21, 820–822 (1996).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brunger, A.T. et al. Crystallography & NMR system. A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta. Crystallogr. D 50, 760–763 (1994).
Fersht, A.R. et al. Active site titration and aminoacyl adenylate binding stoichiometry of aminoacyl-tRNA synthetases. Biochemistry 14, 1–4 (1975).
Acknowledgements
We thank A. Héroux for data collection at the National Synchrotron Light Source, Brookhaven National Laboratory, and T. Christian for assisting with preparation of the T7 transcript of tRNACys. This work was supported by US National Institutes of Health grants GM63713 (to J.J.P.) and GM56662 (to Y.M.H.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Sequence alignment of CysRS enzymes. (PDF 178 kb)
Rights and permissions
About this article
Cite this article
Hauenstein, S., Zhang, CM., Hou, YM. et al. Shape-selective RNA recognition by cysteinyl-tRNA synthetase. Nat Struct Mol Biol 11, 1134–1141 (2004). https://doi.org/10.1038/nsmb849
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb849
This article is cited by
-
Relaxed sequence constraints favor mutational freedom in idiosyncratic metazoan mitochondrial tRNAs
Nature Communications (2020)
-
RNAfitme: a webserver for modeling nucleobase and nucleoside residue conformation in fixed-backbone RNA structures
BMC Bioinformatics (2018)
-
Structural basis for tRNA-dependent cysteine biosynthesis
Nature Communications (2017)
-
Ligand co-crystallization of aminoacyl-tRNA synthetases from infectious disease organisms
Scientific Reports (2017)
-
Plant plasma membrane-bound staphylococcal-like DNases as a novel class of eukaryotic nucleases
BMC Plant Biology (2012)