Abstract
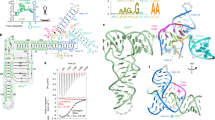
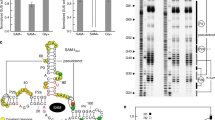
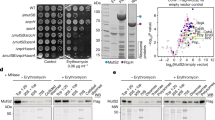
We have identified the SMK box as a conserved RNA motif in the 5′ untranslated leader region of metK (SAM synthetase) genes in lactic acid bacteria, including Enterococcus, Streptococcus and Lactococcus species. This RNA element bound SAM in vitro, and binding of SAM caused an RNA structural rearrangement that resulted in sequestration of the Shine-Dalgarno (SD) sequence. Mutations that disrupted pairing between the SD region and a sequence complementary to the SD blocked SAM binding, whereas compensatory mutations that restored pairing restored SAM binding. The Enterococcus faecalis SMK box conferred translational repression of a lacZ reporter when cells were grown under conditions where SAM pools are elevated, and mutations that blocked SAM binding resulted in loss of repression, demonstrating that the SMK box is functional in vivo. The SMK box therefore represents a new SAM-binding riboswitch distinct from the previously identified S box RNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Grundy, F.J. & Henkin, T.M. Regulation of gene expression by effectors that bind to RNA. Curr. Opin. Microbiol. 7, 126–131 (2004).
Tucker, B.J. & Breaker, R.R. Riboswitches as versatile control elements. Curr. Opin. Struct. Biol. 15, 342–348 (2005).
Nudler, E. & Mironov, A.S. The riboswitch control of bacterial metabolism. Trends Biochem. Sci. 29, 11–17 (2004).
Grundy, F.J. & Henkin, T.M. The S box regulon: a new global transcription termination control system for methionine and cysteine biosynthesis genes in Gram-positive bacteria. Mol. Microbiol. 30, 737–749 (1998).
Grundy, F.J. & Henkin, T.M. The T box and S box transcription termination control systems. Front. Biosci. 8, d20–d31 (2003).
McDaniel, B.A., Grundy, F.J., Artsimovitch, I. & Henkin, T.M. Transcription termination control of the S box system: direct measurement of S-adenosylmethionine by the leader RNA. Proc. Natl. Acad. Sci. USA 100, 3083–3088 (2003).
Epshtein, V., Mironov, A.S. & Nudler, E. The riboswitch-mediated control of sulfur metabolism in bacteria. Proc. Natl. Acad. Sci. USA 100, 5052–5056 (2003).
Winkler, W.C., Nahvi, A., Sudarsen, N., Barrick, J.E. & Breaker, R.R. An mRNA structure that controls gene expression by binding S-adenosylmethionine. Nat. Struct. Biol. 10, 701–707 (2003).
McDaniel, B.A., Grundy, F.J. & Henkin, T.M. A tertiary structural element in S box leader RNAs is required for S-adenosylmethionine-directed transcription termination. Mol. Microbiol. 57, 1008–1021 (2005).
Rodionov, D.A., Vitreschak, A.G., Mironov, A.A. & Gelfand, M.S. Comparative genomics of the methionine metabolism in Gram-positive bacteria: a variety of regulatory systems. Nucleic Acids Res. 32, 3340–3353 (2004).
Grundy, F.J. & Henkin, T.M. Synthesis of serine, glycine, cysteine and methionine. in Bacillus subtilis and Its Closest Relatives: from Genes to Cells (eds. Sonenshein, A.L., Hoch, J.A. & Losick, R.) 245–254 (American Society for Microbiology, Washington, DC, USA 2002).
Wabiko, H., Ochi, K., Nguyen, D.M., Allen, E.R. & Freese, E. Genetic mapping and physiological consequences of metE mutations of Bacillus subtilis. J. Bacteriol. 170, 2705–2710 (1988).
Corbino, K.A. et al. Evidence for a second class of S-adenosylmethionine riboswitches and other regulatory RNA motifs in alpha-proteobacteria. Genome Biol. [online] 6, R70 (2005).
Gagnon, Y.R. et al. Clustering and co-transcription of the Bacillus subtilis genes encoding the aminoacyl-tRNA synthetase genes specific for glutamate and for cysteine and the first gene for cysteine biosynthesis. J. Biol. Chem. 269, 7473–7482 (1994).
Mansilla, M.C., Albanesi, D. & deMendoza, D. Transcriptional control of the sulfur-regulated cysH operon encoding genes involved in L-cysteine biosynthesis in Bacillus subtilis. J. Bacteriol. 182, 5885–5892 (2000).
Guillouard, I. et al. Identification of Bacillus subtilis CysL, a regulator of the cysJI operon, which encodes sulfite reductase. J. Bacteriol. 184, 4681–4689 (2002).
Fernandez, M., Kleerebezem, M., Kuipers, O.P., Siezen, R.J. & van Kranenburg, R. Regulation of the metC-cysK operon, involved in sulfur metabolism in Lactococcus lactis. J. Bacteriol. 184, 82–90 (2002).
Sperandio, B., Polard, P., Ehrlich, S.D., Renault, P. & Guedon, E. Sulfur amino acid metabolism and its control in Lactococcus lactis IL1403. J. Bacteriol. 187, 3762–3778 (2005).
Shelver, D., Rajagopal, L., Harris, T.O. & Rubens, C.E. MtaR, a regulator of methionine transport, is critical for survival of group B streptococcus in vivo. J. Bacteriol. 185, 6592–6599 (2003).
Greene, R.C. Biosynthesis of methionine. in Escherichia coli and Salmonella: Cellular and Molecular Biology (eds. Neidhardt, F.C. et al.) 542–560 (American Society for Microbiology, Washington, DC, USA, 1996).
Yousef, M.R., Grundy, F.G. & Henkin, T.M. tRNA requirements for glyQS antitermination: a new twist on tRNA. RNA 9, 1148–1156 (2003).
Grundy, F.J., Waters, D.A., Allen, S.H. & Henkin, T.M. Regulation of the Bacillus subtilis acetate kinase gene by CcpA. J. Bacteriol. 175, 7348–7355 (1993).
Donnelly, C.E. & Sonenshein, A.L. Promoter-probe plasmid for Bacillus subtilis. J. Bacteriol. 157, 965–967 (1984).
Anagnostopoulos, C. & Spizizen, J. Requirements for transformation in Bacillus subtilis. J. Bacteriol. 81, 741–746 (1961).
Miller, J. Experiments in Molecular Genetics. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 1972).
Grundy, F.J., Moir, T.R., Haldeman, M.T. & Henkin, T.M. Sequence requirements for terminators and antiterminators in the T box transcription antitermination system: disparity between conservation and functional requirements. Nucleic Acids Res. 30, 1646–1655 (2002).
Acknowledgements
We thank S. Tigert for technical assistance. Preliminary sequence data were obtained from The Institute for Genomic Research website at http://www.tigr.org. This work was supported by the US National Institutes of Health (grant GM63615).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Oligonucleotide primers (PDF 13 kb)
Rights and permissions
About this article
Cite this article
Fuchs, R., Grundy, F. & Henkin, T. The SMK box is a new SAM-binding RNA for translational regulation of SAM synthetase. Nat Struct Mol Biol 13, 226–233 (2006). https://doi.org/10.1038/nsmb1059
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1059
This article is cited by
-
Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution
Nature Communications (2023)
-
SAM-VI riboswitch structure and signature for ligand discrimination
Nature Communications (2019)
-
Unraveling RNA dynamical behavior of TPP riboswitches: a comparison between Escherichia coli and Arabidopsis thaliana
Scientific Reports (2019)
-
Development of a genetically encodable FRET system using fluorescent RNA aptamers
Nature Communications (2018)
-
Secondary structural entropy in RNA switch (Riboswitch) identification
BMC Bioinformatics (2015)