Abstract
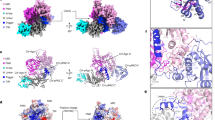
Oligomeric complexes of Trax and Translin proteins, known as C3POs, participate in several eukaryotic nucleic acid metabolism pathways, including RNA interference and tRNA processing. In RNA interference in humans and Drosophila, C3PO activates the RNA-induced silencing complex (RISC) by removing the passenger strand of the small interfering RNA precursor duplex, using nuclease activity present in Trax. How C3POs engage with nucleic acid substrates is unknown. Here we identify a single protein from Archaeoglobus fulgidus that assembles into an octamer highly similar to human C3PO. The structure in complex with duplex RNA reveals that the octamer entirely encapsulates a single 13-base-pair RNA duplex inside a large inner cavity. Trax-like-subunit catalytic sites target opposite strands of the duplex for cleavage separated by 7 base pairs. The structure provides insight into the mechanism of RNA recognition and cleavage by an archaeal C3PO-like complex.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ghildiyal, M. & Zamore, P.D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 10, 94–108 (2009).
Kim, V.N., Han, J. & Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Czech, B. & Hannon, G.J. Small RNA sorting: matchmaking for Argonautes. Nat. Rev. Genet. 12, 19–31 (2011).
Kawamata, T. & Tomari, Y. Making RISC. Trends Biochem. Sci. 35, 368–376 (2010).
Nykänen, A., Haley, B. & Zamore, P.D. ATP requirements and small interfering RNA structure in the RNA interference pathway. Cell 107, 309–321 (2001).
Tomari, Y. et al. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 116, 831–841 (2004).
Pham, J.W., Pellino, J.L., Lee, Y.S., Carthew, R.W. & Sontheimer, E.J.A. Dicer-2-dependent 80s complex cleaves targeted mRNAs during RNAi in Drosophila. Cell 117, 83–94 (2004).
Kawamata, T., Seitz, H. & Tomari, Y. Structural determinants of miRNAs for RISC loading and slicer-independent unwinding. Nat. Struct. Mol. Biol. 16, 953–960 (2009).
Yoda, M. et al. ATP-dependent human RISC assembly pathways. Nat. Struct. Mol. Biol. 17, 17–23 (2010).
Iwasaki, S. et al. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 39, 292–299 (2010).
Miyoshi, T., Takeuchi, A., Siomi, H. & Siomi, M.C. A direct role for Hsp90 in pre-RISC formation in Drosophila. Nat. Struct. Mol. Biol. 17, 1024–1026 (2010).
Johnston, M., Geoffroy, M.C., Sobala, A., Hay, R. & Hutvagner, G. HSP90 protein stabilizes unloaded argonaute complexes and microscopic P-bodies in human cells. Mol. Biol. Cell 21, 1462–1469 (2010).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Lee, Y.S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Liu, X., Jiang, F., Kalidas, S., Smith, D. & Liu, Q. Dicer-2 and R2D2 coordinately bind siRNA to promote assembly of the siRISC complexes. RNA 12, 1514–1520 (2006).
Liu, Y. et al. C3PO, an endoribonuclease that promotes RNAi by facilitating RISC activation. Science 325, 750–753 (2009).
Ye, X. et al. Structure of C3PO and mechanism of human RISC activation. Nat. Struct. Mol. Biol. 18, 650–657 (2011).
Schwarz, D.S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Khvorova, A., Reynolds, A. & Jayasena, S.D. Functional siRNAs and miRNAs exhibit strand bias. Cell 115, 209–216 (2003).
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M.C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005).
Rand, T.A., Petersen, S., Du, F. & Wang, X. Argonaute2 cleaves the anti-guide strand of siRNA during RISC activation. Cell 123, 621–629 (2005).
Matranga, C., Tomari, Y., Shin, C., Bartel, D.P. & Zamore, P.D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Leuschner, P.J., Ameres, S.L., Kueng, S. & Martinez, J. Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep. 7, 314–320 (2006).
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D.J. Structure of the guide-strand-containing argonaute silencing complex. Nature 456, 209–213 (2008).
Schirle, N.T. & MacRae, I.J. The crystal structure of human Argonaute2. Science 336, 1037–1040 (2012).
Nakanishi, K., Weinberg, D.E., Bartel, D.P. & Patel, D.J. Structure of yeast Argonaute with guide RNA. Nature 486, 368–374 (2012).
Elkayam, E. et al. The structure of human Argonaute-2 in complex with miR-20a. Cell 150, 100–110 (2012).
Jaendling, A. & McFarlane, R.J. Biological roles of translin and translin-associated factor-X: RNA metabolism comes to the fore. Biochem. J. 429, 225–234 (2010).
Li, L. et al. The translin-TRAX complex (C3PO) is a ribonuclease in tRNA processing. Nat. Struct. Mol. Biol. 19, 824–830 (2012).
Tian, Y. et al. Multimeric assembly and biochemical characterization of the Trax-translin endonuclease complex. Nat. Struct. Mol. Biol. 18, 658–664 (2011).
Pascal, J.M., Hart, P.J., Hecht, N.B. & Robertus, J.D. Crystal structure of TB-RBP, a novel RNA-binding and regulating protein. J. Mol. Biol. 319, 1049–1057 (2002).
Sugiura, I. et al. Structure of human translin at 2.2 Å resolution. Acta Crystallogr. D Biol. Crystallogr. 60, 674–679 (2004).
Chennathukuzhi, V. et al. Mice deficient for testis-brain RNA-binding protein exhibit a coordinate loss of TRAX, reduced fertility, altered gene expression in the brain, and behavioral changes. Mol. Cell. Biol. 23, 6419–6434 (2003).
Yang, S. et al. Translin-associated factor X is post-transcriptionally regulated by its partner protein TB-RBP, and both are essential for normal cell proliferation. J. Biol. Chem. 279, 12605–12614 (2004).
Gupta, G.D. et al. Co-expressed recombinant human Translin-Trax complex binds DNA. FEBS Lett. 579, 3141–3146 (2005).
Claussen, M., Koch, R., Jin, Z.Y. & Suter, B. Functional characterization of Drosophila Translin and Trax. Genetics 174, 1337–1347 (2006).
Jaendling, A., Ramayah, S., Pryce, D.W. & McFarlane, R.J. Functional characterisation of the Schizosaccharomyces pombe homologue of the leukaemia-associated translocation breakpoint binding protein translin and its binding partner, TRAX. Biochim. Biophys. Acta 1783, 203–213 (2008).
Aoki, K., Suzuki, K., Ishida, R. & Kasai, M. The DNA binding activity of Translin is mediated by a basic region in the ring-shaped structure conserved in evolution. FEBS Lett. 443, 363–366 (1999).
Chennathukuzhi, V.M., Kurihara, Y., Bray, J.D. & Hecht, N.B. Trax (translin-associated factor X), a primarily cytoplasmic protein, inhibits the binding of TB-RBP (translin) to RNA. J. Biol. Chem. 276, 13256–13263 (2001).
Eliahoo, E. et al. Mapping of interaction sites of the Schizosaccharomyces pombe protein Translin with nucleic acids and proteins: a combined molecular genetics and bioinformatics study. Nucleic Acids Res. 38, 2975–2989 (2010).
Gupta, G.D. & Kumar, V. Identification of nucleic acid binding sites on translin-associated factor X (TRAX) protein. PLoS ONE 7, e33035 (2012).
Heidenreich, O., Pieken, W. & Eckstein, F. Chemically modified RNA: approaches and applications. FASEB J. 7, 90–96 (1993).
Parker, J.S., Roe, S.M. & Barford, D. Crystal structure of a PIWI protein suggests mechanisms for siRNA recognition and slicer activity. EMBO J. 23, 4727–4737 (2004).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P.R. An introduction to data reduction: space-group determination, scaling and intensity statistics. Acta Crystallogr. D Biol. Crystallogr. 67, 282–292 (2011).
Collaborative Computational Project. N. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Blessing, R.H. & Smith, G.D. Difference structure-factor normalization for heavy-atom or anomalous-scattering substructure determinations. J. Appl. Crystallogr. 32, 664–670 (1999).
Weeks, C.M. & Miller, R. The design and implementation of SnB v2.0. J. Appl. Crystallogr. 32, 120–124 (1999).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Bond, C.S. & Schuttelkopf, A.W. ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments. Acta Crystallogr. D Biol. Crystallogr. 65, 510–512 (2009).
Acknowledgements
We thank staff at Diamond Light Source, UK for help with data collection. This work was funded by a UK Medical Research Council Career Development Award to J.S.P. (grant no. G0600097). The pTwo-E vector was a gift from A. Oliver, University of Sussex, Brighton, UK.
Author information
Authors and Affiliations
Contributions
E.A.P. produced and purified the proteins, grew the crystals and performed the biochemical assays. E.D.L. provided advice, maintained facilities and assisted with X-ray data collection. J.S.P. collected X-ray data, determined the structures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 3102 kb)
Rights and permissions
About this article
Cite this article
Parizotto, E., Lowe, E. & Parker, J. Structural basis for duplex RNA recognition and cleavage by Archaeoglobus fulgidus C3PO. Nat Struct Mol Biol 20, 380–386 (2013). https://doi.org/10.1038/nsmb.2487
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2487
This article is cited by
-
Translin: A multifunctional protein involved in nucleic acid metabolism
Journal of Biosciences (2019)