Abstract
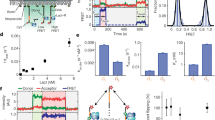
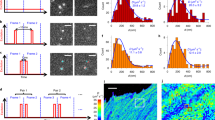
It is known that DNA-binding proteins can slide along the DNA helix while searching for specific binding sites, but their path of motion remains obscure. Do these proteins undergo simple one-dimensional (1D) translational diffusion, or do they rotate to maintain a specific orientation with respect to the DNA helix? We measured 1D diffusion constants as a function of protein size while maintaining the DNA-protein interface. Using bootstrap analysis of single-molecule diffusion data, we compared the results to theoretical predictions for pure translational motion and rotation-coupled sliding along the DNA. The data indicate that DNA-binding proteins undergo rotation-coupled sliding along the DNA helix and can be described by a model of diffusion along the DNA helix on a rugged free-energy landscape. A similar analysis including the 1D diffusion constants of eight proteins of varying size shows that rotation-coupled sliding is a general phenomenon. The average free-energy barrier for sliding along the DNA was 1.1 ± 0.2 kBT. Such small barriers facilitate rapid search for binding sites.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Berg, O.G., Winter, R.B. & Von Hippel, P.H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981).
Gowers, D.M. & Halford, S.E. Protein motion from non-specific to specific DNA by three-dimensional routes aided by supercoiling. EMBO J. 22, 1410–1418 (2003).
Blainey, P.C., van Oijen, A.M., Banerjee, A., Verdine, G.L. & Xie, X.S. A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA. Proc. Natl. Acad. Sci. USA 103, 5752–5757 (2006).
Von Hippel, P.H. & Berg, O.G. Facilitated target location in biological systems. J. Biol. Chem. 264, 675–678 (1989).
Schurr, J.M. The one-dimensional diffusion coefficient of proteins absorbed on DNA. Hydrodynamic considerations. Biophys. Chem. 9, 413–414 (1979).
Sakata-Sogawa, K. & Shimamoto, N. RNA polymerase can track a DNA groove during promoter search. Proc. Natl. Acad. Sci. USA 101, 14731–14735 (2004).
Gnatt, A.L., Cramer, P., Fu, J., Bushnell, D.A. & Kornberg, R.D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292, 1876–1882 (2001).
Kalodimos, C.G. et al. Structure and flexibility adaptation in nonspecific and specific protein-DNA complexes. Science 305, 386–389 (2004).
Von Hippel, P.H. BIOCHEMISTRY: Completing the View of Transcriptional Regulation. Science 305, 350–352 (2004).
Bagchi, B., Blainey, P.C. & Xie, X.S. Diffusion constant of a non-specifically bound protein undergoing curvilinear motion along DNA. J. Phys. Chem. B 112, 6282–6284 (2008).
Zwanzig, R. Diffusion in a rough potential. Proc. Natl. Acad. Sci. USA 85, 2029–2030 (1988).
van Oijen, A.M. et al. Single-molecule kinetics of λ exonuclease reveal base dependence and dynamic disorder. Science 301, 1235–1238 (2003).
Schroeder, C.M., Blainey, P.C., Kim, S.J. & Xie, X.S. Hydrodynamic flow-stretching assay for single-molecule studies of nucleic acid–protein interactions. In: Ha, Taekjip and Selvin, Paul R., editors Single-Molecule Techniques: A Laboratory Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press, 461–492 (2008).
Bruner, S.D., Norman, D.P.G. & Verdine, G.L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Banerjee, A., Yang, W., Karplus, M. & Verdine, G.L. Structure of a repair enzyme interrogating undamaged DNA elucidates recognition of damaged DNA. Nature 434, 612–618 (2005).
Banerjee, A., Santos, W.L. & Verdine, G.L. Structure of a DNA glycosylase searching for lesions. Science 311, 1153–1157 (2006).
Efron, B. & Tibshirani, R.J. An introduction to the bootstrap. Monographs on Statistics and Applied Probability 57. New York: Chapman & Hall, 1993.
Elf, J., Li, G.W. & Xie, X.S. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Slutsky, M. & Mirny, L.A. Kinetics of protein-DNA interaction: facilitated target location in sequence-dependent potential. Biophys. J. 87, 4021–4035 (2004).
Tafvizi, A. et al. Tumor suppressor p53 slides on DNA with low friction and high stability. Biophys. J. 95, L01–L03 (2008).
Liu, S., Abbondanzieri, E.A., Rausch, J.W., Grice, S.F.J. & Zhuang, X. Slide into action: dynamic shuttling of HIV reverse transcriptase on nucleic acid substrates. Science 322, 1092–1097 (2008).
Lin, Y. et al. Using the bias from flow to elucidate single DNA repair protein sliding and interactions with DNA. Biophys. J. 96, 1911–1917 (2009).
Kochaniak, A.B. et al. Proliferating cell nuclear antigen uses two distinct modes to move along DNA. J. Biol. Chem. 284, 17700–17710 (2009).
Ramanathan, S.P. et al. Type III restriction enzymes communicate in 1D without looping between their target sites. Proc. Natl. Acad. Sci. USA 106, 1748–1753 (2009).
Gupta, S. et al. DNA binding provides a molecular strap activating the adenovirus proteinase. Mol. Cell. Proteomics 3, 950–959 (2004).
Acknowledgements
We would like to thank Y. Qi (Harvard Univ.) for providing labeled E. coli MutM, W.J. McGrath and V. Graziano (Brookhaven Natl. Lab.) for a gift of labeled AVP-pVIc, and E.S. Vanamee and A. Aggarwal (Mount Sinai School of Medicine) for a gift of labeled BamHI. We also thank G.W. Li and J. Elf for contributing LacI-Venus diffusion data. B.B. was supported partly by a grant from DST (India) and by a JC Bose Fellowship. Work at Harvard was funded by the NIH Director's Pioneer Award and by NSF.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Methods (or Discussion or Data) and Supplementary Figures 1 and 2 (PDF 779 kb)
Rights and permissions
About this article
Cite this article
Blainey, P., Luo, G., Kou, S. et al. Nonspecifically bound proteins spin while diffusing along DNA. Nat Struct Mol Biol 16, 1224–1229 (2009). https://doi.org/10.1038/nsmb.1716
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1716
This article is cited by
-
Protein-DNA recognition mechanisms and specificity
Biophysical Reviews (2023)
-
MutS functions as a clamp loader by positioning MutL on the DNA during mismatch repair
Nature Communications (2022)
-
Non-flipping DNA glycosylase AlkD scans DNA without formation of a stable interrogation complex
Communications Biology (2021)
-
On the stability of protein–DNA complexes in molecular dynamics simulations using the CUFIX corrections
Journal of the Korean Physical Society (2021)
-
A mini-review of the diffusion dynamics of DNA-binding proteins: experiments and models
Journal of the Korean Physical Society (2021)