Abstract
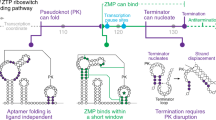
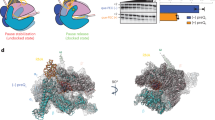
Riboswitches are mRNA domains that bind metabolites and modulate gene expression in cis. We report cocrystal structures of a remarkably compact riboswitch (34 nucleotides suffice for ligand recognition) from Bacillus subtilis that is selective for the essential nucleobase preQ1 (7-aminomethyl-7-deazaguanine). The structures reveal a previously unrecognized pseudoknot fold and suggest a conserved gene-regulatory mechanism whereby ligand binding promotes sequestration of an RNA segment that otherwise assembles into a transcriptional antiterminator.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Iwata-Reuyl, D. Bioorg. Chem. 31, 24–43 (2003).
Roth, A. et al. Nat. Struct. Mol. Biol. 14, 308–317 (2007).
Meyer, M.M., Roth, A., Chervin, S.M., Garcia, G.A. & Breaker, R.R. RNA 14, 685–695 (2008).
Barrick, J.E. et al. Proc. Natl. Acad. Sci. USA 101, 6421–6426 (2004).
Robertson, M.P. & Scott, W.G. Science 315, 1549–1553 (2007).
Xiao, H., Murakami, H., Suga, H. & Ferré-D'Amaré, A.R. Nature 454, 358–361 (2008).
Aalberts, D.P. & Hodas, N.O. Nucleic Acids Res. 33, 2210–2214 (2005).
Klein, D.J. & Ferré-D'Amaré, A.R. Science 313, 1752–1756 (2006).
Gilbert, S.D., Rambo, R.P., Van Tyne, D. & Batey, R.T. Nat. Struct. Mol. Biol. 15, 177–182 (2008).
Egli, M., Minasov, G., Su, L. & Rich, A. Proc. Natl. Acad. Sci. USA 99, 4302–4307 (2002).
Nix, J., Sussman, D. & Wilson, C. J. Mol. Biol. 296, 1235–1244 (2000).
Batey, R.T., Gilbert, S.D. & Montange, R.K. Nature 432, 411–415 (2004).
Edwards, T.E., Klein, D.J. & Ferré-D'Amaré, A.R. Curr. Opin. Struct. Biol. 17, 273–279 (2007).
Leontis, N.B. & Westhof, E. RNA 7, 499–512 (2001).
Acknowledgements
We thank the staff of Advanced Light Source (ALS) beamline 5.0.1 and J. Bolduc for assistance with synchrotron and home laboratory data collection, respectively, R. Huang (University of Illinois at Urbana-Champaign) for his gift of preQ1, and N. Baird, T. Hamma, C. Hoang, N. Kulshina, J. Pitt, J. Posakony, A. Roll-Mecak and H. Xiao for discussions. This work was supported by grants from the Damon Runyon Cancer Research Foundation (DRG-1863-05 to D.J.K. and DRG-1844-04 to T.E.E.), the Howard Hughes Medical Institute (A.R.F.-D.), the US National Institutes of Health (K99 GM084076-01 to D.J.K.) and the W.M. Keck Foundation (A.R.F.-D.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 5258 kb)
Rights and permissions
About this article
Cite this article
Klein, D., Edwards, T. & Ferré-D'Amaré, A. Cocrystal structure of a class I preQ1 riboswitch reveals a pseudoknot recognizing an essential hypermodified nucleobase. Nat Struct Mol Biol 16, 343–344 (2009). https://doi.org/10.1038/nsmb.1563
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1563
This article is cited by
-
A small RNA that cooperatively senses two stacked metabolites in one pocket for gene control
Nature Communications (2022)
-
A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites
Nature Communications (2021)
-
A natural riboswitch scaffold with self-methylation activity
Nature Communications (2021)
-
Structure and functional reselection of the Mango-III fluorogenic RNA aptamer
Nature Chemical Biology (2019)
-
High-affinity recognition of specific tRNAs by an mRNA anticodon-binding groove
Nature Structural & Molecular Biology (2019)