Abstract
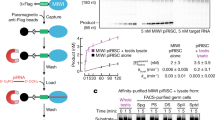
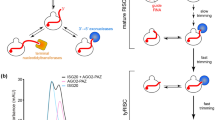
RNA interference (RNAi) is a process in which double-stranded RNA is cleaved into small interfering RNAs (siRNAs) that induce the destruction of homologous single-stranded mRNAs. Argonaute proteins are essential components of this silencing process; they bind siRNAs directly and can cleave RNA targets using a conserved RNase H motif. In Caenorhabditis elegans, the Argonaute protein RDE-1 has a central role in RNAi. In animals lacking RDE-1, the introduction of double-stranded RNA does not trigger any detectable level of RNAi. Here we show that RNase H activity of RDE-1 is required only for efficient removal of the passenger strand of the siRNA duplex and not for triggering the silencing response at the target-mRNA level. These results uncouple the role of the RDE-1 RNase H activity in small RNA maturation from its role in target-mRNA silencing in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Hutvagner, G. & Simard, M.J. Argonaute proteins: key players in RNA silencing. Nat. Rev. Mol. Cell Biol. 9, 22–32 (2008).
Peters, L. & Meister, G. Argonaute proteins: mediators of RNA silencing. Mol. Cell 26, 611–623 (2007).
Lingel, A., Simon, B., Izaurralde, E. & Sattler, M. Structure and nucleic-acid binding of the Drosophila Argonaute 2 PAZ domain. Nature 426, 465–469 (2003).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Ma, J.B., Ye, K. & Patel, D.J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004).
Song, J.J. et al. The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nat. Struct. Biol. 10, 1026–1032 (2003).
Song, J.J., Smith, S.K., Hannon, G.J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Yan, K.S. et al. Structure and conserved RNA binding of the PAZ domain. Nature 426, 468–474 (2003).
Baumberger, N. & Baulcombe, D.C. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl. Acad. Sci. USA 102, 11928–11933 (2005).
Gunawardane, L.S. et al. A Slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007).
Lau, N.C. et al. Characterization of the piRNA complex from rat testes. Science 313, 363–367 (2006).
Saito, K. et al. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes Dev. 20, 2214–2222 (2006).
Irvine, D.V. et al. Argonaute slicing is required for heterochromatic silencing and spreading. Science 313, 1134–1137 (2006).
Rand, T.A., Ginalski, K., Grishin, N.V. & Wang, X. Biochemical identification of Argonaute 2 as the sole protein required for RNA-induced silencing complex activity. Proc. Natl. Acad. Sci. USA 101, 14385–14389 (2004).
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M.C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005).
Rivas, F.V. et al. Purified Argonaute2 and an siRNA form recombinant human RISC. Nat. Struct. Mol. Biol. 12, 340–349 (2005).
Martinez, J., Patkaniowska, A., Urlaub, H., Luhrmann, R. & Tuschl, T. Single-stranded antisense siRNAs guide target RNA cleavage in RNAi. Cell 110, 563–574 (2002).
Okamura, K., Ishizuka, A., Siomi, H. & Siomi, M.C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 18, 1655–1666 (2004).
Buker, S.M. et al. Two different Argonaute complexes are required for siRNA generation and heterochromatin assembly in fission yeast. Nat. Struct. Mol. Biol. 14, 200–207 (2007).
Maiti, M., Lee, H.C. & Liu, Y. QIP, a putative exonuclease, interacts with the Neurospora Argonaute protein and facilitates conversion of duplex siRNA into single strands. Genes Dev. 21, 590–600 (2007).
Leuschner, P.J., Ameres, S.L., Kueng, S. & Martinez, J. Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep. 7, 314–320 (2006).
Matranga, C., Tomari, Y., Shin, C., Bartel, D.P. & Zamore, P.D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Rand, T.A., Petersen, S., Du, F. & Wang, X. Argonaute2 cleaves the anti-guide strand of siRNA during RISC activation. Cell 123, 621–629 (2005).
Kim, K., Lee, Y.S. & Carthew, R.W. Conversion of pre-RISC to holo-RISC by Ago2 during assembly of RNAi complexes. RNA 13, 22–29 (2007).
Pak, J. & Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans. Science 315, 241–244 (2007).
Sijen, T. et al. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107, 465–476 (2001).
Sijen, T., Steiner, F.A., Thijssen, K.L. & Plasterk, R.H. Secondary siRNAs result from unprimed RNA synthesis and form a distinct class. Science 315, 244–247 (2007).
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Aoki, K., Moriguchi, H., Yoshioka, T., Okawa, K. & Tabara, H. In vitro analyses of the production and activity of secondary small interfering RNAs in C. elegans. EMBO J. 26, 5007–5019 (2007).
Steiner, F.A. et al. Structural features of small RNA precursors determine Argonaute loading in Caenorhabditis elegans. Nat. Struct. Mol. Biol. 14, 927–933 (2007).
Tomari, Y., Du, T. & Zamore, P.D. Sorting of Drosophila small RNA silencing RNAs. Cell 130, 299–308 (2007).
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123–132 (1999).
Tabara, H., Yigit, E., Siomi, H. & Mello, C.C. The dsRNA binding protein RDE-4 interacts with RDE-1, DCR-1, and a DExH-box helicase to direct RNAi in C. elegans. Cell 109, 861–871 (2002).
Förstemann, K., Horwich, M.D., Wee, L., Tomari, Y. & Zamore, P.D. Drosophila microRNAs are sorted into functionally distinct Argonaute complexes after production by Dicer-1. Cell 130, 287–297 (2007).
Acknowledgements
We thank C. Mello (University of Massachusetts Medical School) for providing strains and B. Ason, T. Sixma and M. Bühler for help and discussions. The work was supported by a VIDI fellowship from the Dutch Scientific Organization (NWO) to T.S. and the Sixth Framework Programme of the European Commission through the SIROCCO Integrated Project to R.F.K.
Author information
Authors and Affiliations
Contributions
F.A.S., T.S. and R.F.K. designed the experiments; F.A.S., K.L.O., S.W.H. and T.S. performed the experiments; F.A.S. and R.F.K. wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1 (PDF 629 kb)
Rights and permissions
About this article
Cite this article
Steiner, F., Okihara, K., Hoogstrate, S. et al. RDE-1 slicer activity is required only for passenger-strand cleavage during RNAi in Caenorhabditis elegans. Nat Struct Mol Biol 16, 207–211 (2009). https://doi.org/10.1038/nsmb.1541
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1541
This article is cited by
-
Genomes of parasitic nematodes (Meloidogyne hapla, Meloidogyne incognita, Ascaris suum and Brugia malayi) have a reduced complement of small RNA interference pathway genes: knockdown can reduce host infectivity of M. incognita
Functional & Integrative Genomics (2016)
-
Competition between ADAR and RNAi pathways for an extensive class of RNA targets
Nature Structural & Molecular Biology (2011)
-
Two classes of silencing RNAs move between Caenorhabditis elegans tissues
Nature Structural & Molecular Biology (2011)
-
Small RNA sorting: matchmaking for Argonautes
Nature Reviews Genetics (2011)
-
RNAi pathway integration in Caenorhabditis elegans development
Functional & Integrative Genomics (2011)