Abstract
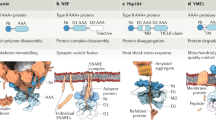
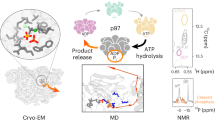
p97 (also called VCP), a member of the AAA ATPase family, is involved in several cellular processes, including membrane fusion and extraction of proteins from the endoplasmic reticulum for cytoplasmic degradation. We have studied the conformational changes that p97 undergoes during the ATPase cycle by cryo-EM and single-particle analysis. Three-dimensional maps show that the two AAA domains, D1 and D2, as well as the N-domains, experience conformational changes during ATP binding, ATP hydrolysis, Pi release and ADP release. The N-domain is flexible in most nucleotide states except after ATP hydrolysis. The rings formed by D1 and D2 rotate with respect to each other, and the size of their axial openings fluctuates. Taken together, our results depict the movements that this and possibly other AAA ATPases can undergo during an ATPase cycle.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Peters, J., Walsh, M.J. & Franke, W.W. An abundant and ubiquitous homo-oligomeric ring-shaped ATPase particle related to the putative vesicle fusion proteins Sec18p and NSF. EMBO J. 9, 1757–1767 (1990).
Rabouille, C. et al. Syntaxin 5 is a common component of the NSF- and p97-mediated reassembly pathways of Golgi cisternae from mitotic Golgi fragments in vitro. Cell 92, 603–610 (1998).
Roy, L. et al. Role of p97 and syntaxin-5 in the assembly of transitional endoplasmic reticulum. Mol. Biol. Cell 11, 2529–2542 (2000).
Hetzer, M. et al. Distinct AAA-ATPase p97 complexes function in discrete steps of nuclear assembly. Nature Cell Biol. 3, 1086–1091 (2001).
Shirogane, T. et al. Synergistic roles for Pim-1 and c-Myc in STAT3-mediated cell cycle progression and antiapoptosis. Immunity 11, 709–719 (1999).
Madeo, F., Fröhlich, E. & Fröhlich, K. A yeast mutant showing diagnostic markers of early and late apoptosis. J. Cell Biol. 139, 729–734 (1997).
Madeo, F., Schlauer, J., Zischka, H., Mecke, D. & Fröhlich, K. Tyrosine phosphorylation regulates cell cycle-dependent nuclear localization of cdc48p. Mol. Biol. Cell 9, 131–141 (1998).
Yamada, T. et al. p97 ATPase, an ATPase involved in membrane fusion, interacts with DNA unwinding factor (DUF) that functions in DNA replication. FEBS Lett. 466, 287–291 (2000).
Dai, R.M., Chen, E., Longo, D.L., Gorbea, C.M. & Li, C.H. Involvement of valosin-containing protein, an ATPase co-purified with IκBα and 26 S proteasome, in ubiquitin-proteasome-mediated degradation of IκBα. J. Biol. Chem. 273, 3562–3573 (1998).
Meyer, H.H., Shorter, J.G., Seemann, J., Pappin, D. & Warren, G. A complex of mammalian ufd1 and npl4 links the AAA-ATPase, p97, to ubiquitin and nuclear transport pathways. EMBO J. 19, 2181–2192 (2000).
Yen, C. et al. Involvement of the ubiquitin-proteasome pathway in the degradation of nontyrosine kinase-type cytokine receptors of IL-9, IL-2, and erythropoietin. J. Immunol. 165, 6372–6380 (2000).
Dai, R.M. & Li, C.H. Valosin-containing protein is a multi-ubiquitin chain-targeting factor required in ubiquitin-proteasome degradation. Nature Cell Biol. 3, 740–744 (2001).
Ye, Y., Meyer, H.H. & Rapoport, T.A. The AAA ATPase cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature 414, 652–656 (2001).
Braun, S., Matuschewski, K., Rape, M., Thoms, S. & Jentsch, S. Role of the ubiquitin-selective CDC48UFD1/NPL4 chaperone (segregase) in ERAD of OLE1 and other substrates. EMBO J. 21, 615–621 (2002).
Jarosch, E. et al. Protein dislocation from the ER requires polyubiquitination and the AAA-ATPase Cdc48. Nature Cell Biol. 4, 134–139 (2002).
Rabinovich, E., Kerem, A., Fröhlich, K.U., Diamant, N. & Bar-Nun, S. AAA-ATPase p97/cdc48p, a cytosolic chaperone required for endoplasmic reticulum-associated protein degradation. Mol. Cell. Biol. 22, 626–634 (2002).
Kondo, H. et al. p47 is a cofactor for p97-mediated membrane fusion. Nature 388, 75–78 (1997).
Meyer, H.H., Kondo, H. & Warren, G. The p47 co-factor regulates the ATPase activity of the membrane fusion protein, p97. FEBS Lett. 437, 255–257 (1998).
Koegl, M.T. et al. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 96, 635–644 (1999).
Ghislain, M., Dohmen, R.J., Lévy, F. & Varshavsky, A. Cdc48p interacts with Ufd3p, a WD repeat protein required for ubiquitin-mediated proteolysis in Saccharomyces cerevisiae. EMBO J. 15, 4884–4899 (1996).
Maurizi, M.R. & Li, C.H. AAA proteins: in search of a common molecular basis. International Meeting on Cellular Functions of AAA Proteins. EMBO Rep. 2, 980–985 (2001).
Patel, S. & Latterich, M. The AAA team: related ATPases with diverse functions. Trends Cell. Biol. 8, 65–71 (1998).
Neuwald, A.F., Aravind, L., Spouge, J.L. & Koonin, E.V. AAA+: a class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 9, 27–43 (1999).
Ogura, T. & Wilkinson, A.J. AAA+ superfamily ATPases: common structure — diverse function. Genes Cells 6, 575–597 (2001).
Vale, R.D. AAA proteins. Lords of the ring. J. Cell Biol. 10, F13–19 (2000).
Babor, S.M. & Fass, D. Crystal structure of the Sec18p N-terminal domain. Proc. Natl. Acad. Sci. USA 96, 14759–14764 (1999).
Coles, M. et al. The solution structure of VAT-N reveals a 'missing link' in the evolution of complex enzymes from a simple βαββ element. Curr. Biol. 9, 1158–1168 (1999).
May, A.P., Misura, K.M.S., Whiteheart, S.W. & Weis, W.I. Crystal structure of the N-terminal domain of N-ethylmaleimide-sensitive fusion protein. Nature Cell Biol. 1, 175–182 (1999).
Yu, R.C., Jahn, R., & Brunger, A.T. NSF N-terminal domain crystal structure: models of NSF function. Mol. Cell 4, 97–107 (1999).
Zhang X., et al. Structure of the AAA ATPase p97. Mol. Cell 6, 1473–1484 (2000).
Peters, J. et al. Ubiquitous soluble Mg2+-ATPase complex. A structural study. J. Mol. Biol. 223, 557–571 (1992).
Rockel, B. et al. Structure of VAT, a CDC48/p97 ATPase homologue from the archaeon Thermoplasma acidophilum as studied by electron tomography. FEBS Lett. 451, 27–32 (1999).
Rouiller, I., Butel, V.M., Latterich, M., Milligan, R.A. & Wilson-Kubalek, E.M. A major conformational change in p97 AAA ATPase upon ATP binding. Mol. Cell 6, 1485–1490 (2000).
Rockel, B., Jakana, J., Chiu, W. & Baumeister, W. Electron cryo-microscopy of VAT, the archaeal p97/CDC48 homologue from Thermoplasma acidophilum. J. Mol. Biol. 317, 673–681 (2002).
Lenzen, C.U., Steinmann, D., Whiteheart, S.W. & Weis, W.I. Crystal structure of the hexamerizatioN-domain of N-ethylmaleimide-sensitive fusion protein. Cell 94, 525–536 (1998).
Yu, R.C., Hanson, P.I., Jahn, R. & Brunger, A.T. Structure of the ATP-dependent oligomerizatioN-domain of N-ethylmaleimide sensitive factor complexed with ATP. Nature Struct. Biol. 5, 803–811 (1998).
Wittinghofer, A. Signaling mechanistics: aluminum fluoride for molecule of the year. Curr. Biol. 7, R682–685 (1997).
Putnam, C.D. et al. Structure and mechanism of the RuvB Holliday junction branch migration motor. J. Mol. Biol. 311, 297–310 (2001).
Sousa, M.C. et al. Crystal and solution structures of an HslUV protease-chaperone complex. Cell 103, 633–643 (2000).
Bochtler, M., Song, H.K., Hartmann, C., Ramachandran, R. & Huber, R. The quaternary arrangement of HslU and HslV in a cocrystal: a response to Wang, Yale. J. Struct. Biol. 135, 281–293 (2001).
Wang, J. A corrected quaternary arrangement of the peptidase HslV and ATPase HslU in a cocrystal structure. J. Struct. Biol. 134, 15–24 (2001).
Wang, J. et al. Nucleotide-dependent conformational changes in a protease-associated ATPase HsIU. Structure 9, 1107–1116 (2001).
Lamb, J.R., Fu, V., Wirtz, E. & Bangs, J.D. Functional analysis of the trypanosomal AAA protein TbVCP with trans-dominant ATP hydrolysis mutants. J. Biol. Chem. 276, 21512–21520 (2001).
Pamnani, V. et al. Cloning, sequencing and expression of VAT, a CDC48/p97 ATPase homologue from the archaeon Thermoplasma acidophilum. FEBS Lett. 404, 263–268 (1997).
Frank, J. & Radermacher, M. SPIDER and WEB: processing and visualization of images in 3D EM and related fields. J. Struct. Biol. 116, 190–199 (1996).
Sorzano, C.O.S. et al. The effect of overabundant projection directions on 3D reconstruction algorithms. J. Struct. Biol. 133, 108–118 (2001).
Wriggers, W., Milligan, R.A. & McCammon, J.A. Situs: a package for docking crystal structures into low-resolution maps from electron microscopy. J. Struct. Biol. 125, 185–195 (1999).
Jones, T.A., Zou, J-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for the building of protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Acknowledgements
This work was supported by National Institutes of Health grants to R.A.M., E.M.W.-K. and W.I.W. A.P.M. was funded by a Wellcome Trust Prize International Traveling Fellowship. We thank P. Chacon for doing the fitting with Situs47 and B. Sheehan for computing help. We also thank S. Kaiser, M. Bowen and C. Moores for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Rouiller, I., DeLaBarre, B., May, A. et al. Conformational changes of the multifunction p97 AAA ATPase during its ATPase cycle. Nat Struct Mol Biol 9, 950–957 (2002). https://doi.org/10.1038/nsb872
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb872
This article is cited by
-
Designing a chemical inhibitor for the AAA protein spastin using active site mutations
Nature Chemical Biology (2019)
-
OsCDC48/48E complex is required for plant survival in rice (Oryza sativa L.)
Plant Molecular Biology (2019)
-
A structure- and chemical genomics-based approach for repositioning of drugs against VCP/p97 ATPase
Scientific Reports (2017)
-
Quantitative interaction mapping reveals an extended UBX domain in ASPL that disrupts functional p97 hexamers
Nature Communications (2016)
-
Molecular snapshots of the Pex1/6 AAA+ complex in action
Nature Communications (2015)