Abstract
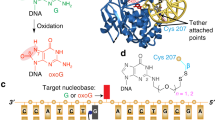
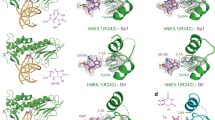
MutM is a bacterial 8-oxoguanine glycosylase responsible for initiating base-excision repair of oxidized guanine residues in DNA. Here we report five different crystal structures of MutM–DNA complexes that represent different steps of the repair reaction cascade catalyzed by the protein and also differ in the identity of the base opposite the lesion (the 'estranged' base). These structures reveal that the MutM active site performs the multiple steps of base-excision and 3′ and 5′ nicking with minimal rearrangement of the DNA backbone.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Michaels, M.L. & Miller, J.H. The GO system protects organisms from the mutagenic effect of the spontaneous lesion 8-hydroxyguanine (7,8-dihydro-8-oxoguanine). J. Bacteriol. 174, 6321–6325 (1992).
Grollman, A.P. & Moriya, M. Mutagenesis by 8-oxoguanine: an enemy within. Trends Genet. 9, 246–249 (1993).
Friedberg, E.C., Walker, G.C. & Siede, W. DNA Repair and Mutagenesis (ASM Press, Washington, D.C.; 1995).
Tajiri, T., Maki, H. & Sekiguchi, M. Functional cooperation of MutT, MutM and MutY proteins in preventing mutations caused by spontaneous oxidation of guanine nucleotide in Escherichia coli. Mutat. Res. 336, 257–267 (1995).
van der Kemp, P.A., Thomas, D., Barbey, R., de Oliveira, R. & Boiteux, S. Cloning and expression in Escherichia coli of the OGG1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine. Proc. Natl. Acad. Sci. USA 93, 5197–5202 (1996).
Nash, H.M. et al. Cloning of a yeast 8-oxoguanine DNA glycosylase reveals the existence of a base-excision DNA-repair protein superfamily. Curr. Biol. 6, 968–980 (1996).
Tchou, J. & Grollman, A.P. The catalytic mechanism of Fpg protein. Evidence for a Schiff base intermediate and amino terminus localization of the catalytic site. J. Biol. Chem. 270, 11671–11677 (1995).
Zharkov, D.O., Rieger, R.A., Iden, C.R. & Grollman, A.P. NH2-terminal proline acts as a nucleophile in the glycosylase/AP-lyase reaction catalyzed by Escherichia coli formamidopyrimidine-DNA glycosylase (Fpg) protein. J. Biol. Chem. 272, 5335–5341 (1997).
Bhagwat, M. & Gerlt, J.A. 3′- and 5′-strand cleavage reactions catalyzed by the Fpg protein from Escherichia coli occur via successive β- and δ-elimination mechanisms, respectively. Biochemistry 35, 659–665 (1996).
Sun, B., Latham, K.A., Dodson, M.L. & Lloyd, R.S. Studies on the catalytic mechanism of five DNA glycosylases. Probing for enzyme-DNA imino intermediates. J. Biol. Chem. 270, 19501–19508 (1995).
Tchou, J. et al. Substrate specificity of Fpg protein. Recognition and cleavage of oxidatively damaged DNA. J. Biol. Chem. 269, 15318–15324 (1994).
Boiteux, S., O'Connor, T.R., Lederer, F., Gouyette, A. & Laval, J. Homogeneous Escherichia coli FPG protein. A DNA glycosylase which excises imidazole ring-opened purines and nicks DNA at apurinic/apyrimidinic sites. J. Biol. Chem. 265, 3916–3922 (1990).
O'Connor, T.R., Boiteux, S. & Laval, J. Repair of imidazole ring-opened purines in DNA: overproduction of the formamidopyrimidine-DNA glycosylase of Escherichia coli using plasmids containing the fpg+ gene. Ann. Ist. Super. Sanita 25, 27–31 (1989).
Tchou, J. et al. 8-oxoguanine (8-hydroxyguanine) DNA glycosylase and its substrate specificity. Proc. Natl. Acad. Sci. USA 88, 4690–4694 (1991).
Hatahet, Z., Kow, Y.W., Purmal, A.A., Cunningham, R.P. & Wallace, S.S. New substrates for old enzymes. 5-Hydroxy-2′-deoxycytidine and 5-hydroxy-2′-deoxyuridine are substrates for Escherichia coli endonuclease III and formamidopyrimidine DNA N-glycosylase, while 5-hydroxy-2′-deoxyuridine is a substrate for uracil DNA N-glycosylase. J. Biol. Chem. 269, 18814–18820 (1994).
Bruner, S.D., Norman, D.P. & Verdine, G.L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Norman, D.P.G., Bruner, S.D. & Verdine, G.L. Coupling of substrate recognition and catalysis by a human base-excision DNA repair protein. J. Am. Chem. Soc. 123, 359–360 (2001).
Boiteux, S., O'Connor, T.R. & Laval, J. Formamidopyrimidine-DNA glycosylase of Escherichia coli: cloning and sequencing of the fpg structural gene and overproduction of the protein. Embo J. 6, 3177–3183 (1987).
Michaels, M.L., Pham, L., Cruz, C. & Miller, J.H. MutM, a protein that prevents G-C–T-A transversions, is formamidopyrimidine-DNA glycosylase. Nucleic Acids Res. 19, 3629–3632 (1991).
Sugahara, M. et al. Crystal structure of a repair enzyme of oxidatively damaged DNA, MutM (Fpg), from an extreme thermophile, Thermus thermophilus HB8. EMBO J. 19, 3857–3869 (2000).
Zharkov, D.O. et al. Structural analysis of an Escherichia coli endonuclease VIII covalent reaction intermediate. Embo J. 21, 789–800 (2002).
Castaing, B., Boiteux, S. & Zelwer, C. DNA containing a chemically reduced apurinic site is a high affinity ligand for the E.coli formamidopyrimidine-DNA glycosylase. Nucleic Acids Res. 20, 389–394 (1992).
Tchou, J., Michaels, M.L., Miller, J.H. & Grollman, A.P. Function of the zinc finger in Escherichia coli Fpg protein. J. Biol. Chem. 268, 26738–26744 (1993).
Sidorkina, O.M. & Laval, J. Role of lysine-57 in the catalytic activities of Escherichia coli formamidopyrimidine-DNA glycosylase (Fpg protein). Nucleic Acids Res. 26, 5351–5357 (1998).
Saparbaev, M. et al. Repair of oxidized purines and damaged pyrimidines by E. coli Fpg protein: different roles of proline 2 and lysine 57 residues. Environ. Mol. Mutagen. 39, 10–17 (2002).
Eschenmoser, A. & Dobler, M. Why pentose and not hexose nucleic acids? Part I. Helv. Chim. Acta 75, 218–259 (1992).
Plum, G.E., Grollman, A.P., Johnson, F. & Breslauer, K.J. Influence of the oxidatively damaged adduct 8-oxodeoxyguanosine on the conformation, energetics, and thermodynamic stability of a DNA duplex. Biochemistry 34, 16148–16160 (1995).
Sidorkina, O.M. & Laval, J. Role of the N-terminal proline residue in the catalytic activities of the Escherichia coli Fpg protein. J. Biol. Chem. 275, 9924–9929 (2000).
Guan, Y. et al. MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nature Struct. Biol. 5, 1058–1064 (1998).
Kuo, C.F. et al. Atomic structure of the DNA repair [4Fe-4S] enzyme endonuclease III. Science 258, 434–440 (1992).
Hollis, T., Ichikawa, Y. & Ellenberger, T. DNA bending and a flip-out mechanism for base excision by the helix-hairpin-helix DNA glycosylase. Escherichia coli AlkA. Embo J. 19, 758–766 (2000).
Bjoras, M., Seeberg, E., Luna, L., Pearl, L.H. & Barrett, T.E. Reciprocal 'flipping' underlies substrate recognition and catalytic activation by the human 8-oxo-guanine DNA glycosylase. J. Mol. Biol. 317, 171–177 (2002).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1996).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Brünger, A.T. Assessment of phase accuracy by cross validation: the free R value. Methods and applications. Acta Crystallogr. D 49, 24–36 (1993).
Read, R.J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A 42 140–149 (1986).
Laskowski, R.J., Macarthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–290 (1993).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Gilboa, R. et al. Structure of a formamidopyrimidine DNA glycosylase-DNA complex. J. Biol. Chem., 23 March 2002 (10.1074/jbc.M202058200).
Acknowledgements
We are grateful to B. Roe and S. Lewis of the OU Advanced Center for Genome Technology for their help with B. stearothermophilus. The entire staff of MacCHESS, especially C. Heaton and B. Miller, provided valuable assistance with data collection and processing. We thank M. Spong for assistance with the crystallization screens, and members of the Verdine, Harrison and Wiley labs for thoughtful discussions. J.C.F is funded by an NSF predoctoral fellowship and an NIH training grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Fromme, J., Verdine, G. Structural insights into lesion recognition and repair by the bacterial 8-oxoguanine DNA glycosylase MutM. Nat Struct Mol Biol 9, 544–552 (2002). https://doi.org/10.1038/nsb809
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb809