Abstract
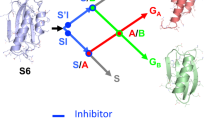
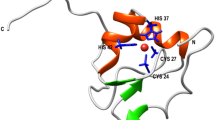
Many proteins populate collapsed intermediate states during folding. In order to elucidate the nature and importance of these species, we have mapped the structure of the on-pathway intermediate of the four-helix protein, Im7, together with the conformational changes it undergoes as it folds to the native state. Kinetic data for 29 Im7 point mutants show that the intermediate contains three of the four helices found in the native structure, packed around a specific hydrophobic core. However, the intermediate contains many non-native interactions; as a result, hydrophobic interactions become disrupted in the rate-limiting transition state before the final helix docks onto the developing structure. The results of this study support a hierarchical mechanism of protein folding and explain why the misfolding of Im7 occurs. The data also demonstrate that non-native interactions can play a significant role in folding, even for small proteins with simple topologies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Fersht, A.R., Matouschek, A. & Serrano, L. The folding of an enzyme.1. Theory of protein engineering analysis of stability and pathway of protein folding. J. Mol. Biol. 224, 771–782 (1992).
Fersht, A.R. Mapping the structures of transition-states and intermediates in folding — delineation of pathways at high-resolution. Phil. Trans. R. Soc. Lond. B 348, 11–15 (1995).
Jackson, S.E. How do small single-domain proteins? Folding Des. 3, R81–R91 (1998).
Grantcharova, V., Alm, E., Baker, D. & Horwich, A. Mechanisms of protein folding. Curr. Opin. Struct. Biol. 11, 70–82 (2001).
Plaxco, K.W., Simons, K.T. & Baker, D. Contact order, transition state placement and the refolding rates of single domain proteins. J. Mol. Biol. 277, 985–994 (1998).
Baker, D. A surprising simplicity to protein folding. Nature 405, 39–42 (2000).
Yeh, S. & Rousseau, D.L. Hierarchical folding of cytochrome c. Nature Struct. Biol. 8, 689–694 (2000).
Roder, H. & Colón, W. Kinetic role of early intermediates in protein folding. Curr. Opin. Struct. Biol. 7, 15–28 (1997).
Gunasekaran, K., Eyles, S.J., Hagler, A.T. & Gierasch, L.M. Keeping it in the family: folding studies of related proteins. Current Opin. Struct. Biol. 11, 83–93 (2001).
Park, S.-H., Shastry, M.C.R. & Roder, H. Folding dynamics of the B1 domain of protein G explored by ultrarapid mixing. Nature Struct. Biol. 6, 943–947 (1999).
Shastry, M.C. & Roder, H. Evidence for barrier-limited protein folding kinetics on the microsecond time scale. Nature Struct. Biol. 5, 385–392 (1998).
Raschke, T.M., Kho, J. & Marqusee, S. Confirmation of the hierarchical folding of RNase H: a protein engineering study. Nature Struct. Biol. 6, 825–831 (1999).
Sosnick, T.R., Shtilerman, M.D., Mayne, L. & Englander, S.W. Ultrafast signals in protein folding and the polypeptide contracted state. Proc. Natl. Acad. Sci. USA 94, 8545–8550 (1997).
Qi, P.X., Sosnick, T.R. & Englander, S.W. The burst phase in ribonuclease A folding and solvent dependence of the unfolded state. Nature Struct. Biol. 5, 882–884 (1998).
Mok, Y.K., Kay, C.M., Kay, L.E. & Forman-Kay, J. NOE data demonstrating a compact unfolded state for an SH3 domain under non-denaturing conditions. J. Mol. Biol. 289, 619–638 (1999).
Baldwin, R.L. Folding consensus? Nature Struct. Biol. 8, 92–94. (2001).
James, R., Kleanthous, C. & Moore, G.R. The biology of E colicins: Paradigms and paradoxes. Microbiology 142, 1569–1580 (1996).
Ferguson, N., Capaldi, A.P., James, R., Kleanthous, C. & Radford, S.E. Rapid folding with and without populated intermediates in the homologous four-helix proteins Im7 and Im9. J. Mol. Biol. 286, 1597–1608 (1999).
Gorski, S.A., Capaldi, A.P., Kleanthous, C. & Radford, S.E. Acidic conditions stabilise intermediates populated during the folding of Im7 and Im9. J. Mol. Biol. 312, 849–863 (2001).
Capaldi, A.P., Shastry, M.C., Kleanthous, C., Roder, H. & Radford, S.E. Ultrarapid mixing experiments reveal that Im7 folds via an on-pathway intermediate. Nature Struct. Biol. 8, 68–72. (2001).
Ozkan, S.B., Bahar, I. & Dill, K.A. Transition states and the meaning of Φ-values in protein folding kinetics. Nature Struct. Biol. 8, 765–769. (2001).
Fersht, A.R. Characterizing transition-states in protein-folding — an essential step in the puzzle. Curr. Opin. Struct. Biol. 5, 79–84 (1995).
Muñoz, V. & Serrano, L. Elucidating the folding problem of helical peptides using empirical parameters. 2. Helix macrodipole effects and rational modification of the helical content of natural peptides. J. Mol. Biol. 245, 275–296 (1995).
Muñoz, V. & Serrano, L. Elucidating the folding problem of helical peptides using empirical parameters. 3. Temperature and pH-dependence. J. Mol. Biol. 245, 297–308 (1995).
Muñoz, V. & Serrano, L. Development of the multiple sequence approximation within the AGADIR model of α–helix formation: Comparison with Zimm-Bragg and Lifson-Roig formalisms. Biopolymers 41, 495–509 (1997).
Lacroix, E., Viguera, A.R. & Serrano, L. Elucidating the folding problem of α-helices: local motifs, long-range electrostatics, ionic strength dependence and prediction of NMR parameters. J. Mol. Biol. 284, 173–191 (1998).
Kuhlmann, U.C., Pommer, A.J., Moore, G.R., James, R. & Kleanthous, C. Specificity in protein–protein interactions: The structural basis for dual recognition in endonuclease colicin–immunity protein complexes. J. Mol. Biol. 301, 1163–1178 (2000).
Wallis, R. et al. Specificity in protein–protein recognition: conserved Im9 residues are the major determinants of stability in the colicin E9 DNase–Im9 complex. Biochemistry 37, 476–485 (1998).
Li, W. et al. Dual recognition and the role of specificity-determining residues in colicin E9 DNase–immunity protein interactions. Biochemistry 37, 11771–11779 (1998).
Baldwin, R.L. & Rose, G.D. Is protein folding hierarchic? I. Local structure and peptide folding. Trends Biochem. Sci. 24, 26–33 (1999).
Baldwin, R.L. & Rose, G.D. Is protein folding hierarchic? II. Folding intermediates and transition states. Trends Biochem. Sci. 24, 77–83 (1999).
Karplus, M. & Weaver, D.L. Protein folding dynamics — the diffusion-collision model and experimental-data. Protein Sci. 3, 650–668 (1994).
Karplus, M. & Weaver, D.L. Protein-folding dynamics. Nature 260, 404–406 (1976).
Burton, R.E., Myers, J.K. & Oas, T.G. Protein folding dynamics: quantitative comparison between theory and experiment. Biochemistry 37, 5337–5343 (1998).
Myers, J.K. & Oas, T.G. Reinterpretation of GCN4-p1 folding kinetics: partial helix formation precedes dimerization in coiled coil folding. J. Mol. Biol. 289, 205–209 (1999).
Myers, J.K. & Oas, T.G. Preorganized secondary structure as an important determinant of fast protein folding. Nature Struct. Biol. 8, 552–558. (2001).
Pappu, R.V. & Weaver, D.L. The early folding kinetics of apomyoglobin. Protein Sci. 7, 480–490 (1998).
Ko, T.P., Liao, C.C., Ku, W.Y., Chak, K.F. & Yuan, H.S. The crystal structure of the DNase domain of colicin E7 in complex with its inhibitor Im7 protein. Structure Fold. Des . 7, 91–102 (1999).
Li, W., Dennis, C.A., Moore, G.R., James, R. & Kleanthous, C. Protein–protein interaction specificity of Im9 for the endonuclease toxin colicin E9 defined by homologue-scanning mutagenesis. J. Biol. Chem. 272, 22253–22258 (1997).
Wallis, R. et al. Protein–protein interactions in colicin E9 Dnase–immunity protein complexes. Cognate and noncognate interactions that span the mM–fM affinity range. Biochemistry 34, 13751–13759 (1995).
Kuwata, K. et al. Structural and kinetic characterisation of early folding events in β-lactoglobulin. Nature Struct. Biol. 8, 151–154 (2001).
Grantcharova, V.P., Riddle, D.S., Santiago, J.V. & Baker, D. Important role of hydrogen bonds in the structurally polarized transition state for folding of the src SH3 domain. Nature Struct. Biol. 5, 714–720 (1998).
Martinez, J.C. & Serrano, L. The folding transition state between SH3 domains is conformationally restricted and evolutionarily conserved. Nature Struct. Biol. 6, 1010–1016 (1999).
Li, L., Mirny, L.A. & Shakhnovich, E.I. Kinetics, thermodynamics and evolution of non-native interactions in a protein folding nucleus. Nature Struct. Biol. 7, 336–342. (2000).
Mirny, L. & Shakhnovich, E.I. Protein folding theory: from lattice to all-atom models. Annu. Rev. Biophys. Biomol. Struct. 30, 361–396 (2001)
Kraulis, P.J. Molscript — a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merrit, E.A. & Bacon, D.J. Raster3D: Photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Dennis, C.A. et al. A structural comparison of the colicin immunity proteins Im7 and Im9 gives new insights into the molecular determinants of immunity- protein specificity. Biochem. J. 333, 183–191 (1998).
Acknowledgements
We would like to thank A. Ashcroft for analyzing the proteins using mass spectrometry, K. Ainley for technical assistance and A. Berry for help making figures. We are also very grateful to D. Otzen, S. Fonseca and members of the Radford lab for advice and help with molecular biology, and to H. Roder for helpful discussions. We acknowledge the Wellcome Trust, the BBSRC and the University of Leeds for financial support. A.P.C. and S.E.R. are members of the Astbury Centre for Structural Molecular Biology, which is part of the North of England Structural Biology Centre and is funded by the BBSRC.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Capaldi, A., Kleanthous, C. & Radford, S. Im7 folding mechanism: misfolding on a path to the native state. Nat Struct Mol Biol 9, 209–216 (2002). https://doi.org/10.1038/nsb757
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb757