Abstract
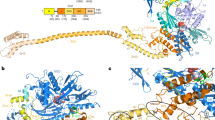
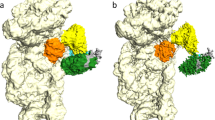
Ribosome anti-association factor eIF6 (originally named according to translation initiation terminology as eukaryotic initiation factor 6) binds to the large ribosomal subunit, thereby preventing inappropriate interactions with the small subunit during initiation of protein synthesis. We have determined the X-ray structures of two IF6 homologs, Methanococcus jannaschii archaeal aIF6 and Sacchromyces cerevisiae eIF6, revealing a phylogenetically conserved 25 kDa protein consisting of five quasi identical α/β subdomains arrayed about a five-fold axis of pseudosymmetry. Yeast eIF6 prevents ribosomal subunit association. Comparative protein structure modeling with other known archaeal and eukaryotic homologs demonstrated the presence of two conserved surface regions, one or both of which may bind the large ribosomal subunit.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Pestova, T.V. & Hellen, C.U. Ribosome recruitment and scanning: what's new? Trends Biochem. Sci. 24, 85–87 (1999).
Trachsel, H., Erni, B., Schreier, M.H. & Staehelin, T. Initiation of mammalian protein synthesis. II. The assembly of the initiation complex with purified initiation factors. J. Mol. Biol. 116, 755–767 (1977).
Russell, D.W. & Spremulli, L.L. Mechanism of action of the wheat germ ribosome dissociation factor: interaction with the 60 S subunit. Arch. Biochem. Biophys. 201, 518–526 (1980).
Russell, D.W. & Spremulli, L.L. Purification and characterization of a ribosome dissociation factor (eukaryotic initiation factor 6) from wheat germ. J. Biol. Chem. 254, 8796–8800 (1979).
Valenzuela, D.M., Chaudhuri, A. & Maitra, U. Eukaryotic ribosomal subunit anti-association activity of calf liver is contained in a single polypeptide chain protein of Mr = 25,500 (eukaryotic initiation factor 6). J. Biol. Chem. 257, 7712–7719 (1982).
Raychaudhuri, P., Stringer, E.A., Valenzuela, D.M. & Maitra, U. Ribosomal subunit antiassociation activity in rabbit reticulocyte lysates. Evidence for a low molecular weight ribosomal subunit antiassociation protein factor (Mr = 25,000). J. Biol. Chem. 259, 11930–11935 (1984).
Si, K., Chaudhuri, J., Chevesich, J. & Maitra, U. Molecular cloning and functional expression of a human cDNA encoding translation initiation factor 6. Proc. Natl. Acad. Sci. USA 94, 14285–14290 (1997).
Si, K. & Maitra, U. The Saccharomyces cerevisiae homologue of mammalian translation initiation factor 6 does not function as a translation initiation factor. Mol. Cell. Biol. 19, 1416–1426 (1999).
Wood, L.C., Ashby, M.N., Grunfeld, C. & Feingold, K.R. Cloning of murine translation initiation factor 6 and functional analysis of the homologous sequence YPR016c in Saccharomyces cerevisiae. J. Biol. Chem. 274, 11653–11659 (1999).
Sanvito, F. et al. The beta4 integrin interactor p27(BBP/eIF6) is an essential nuclear matrix protein involved in 60S ribosomal subunit assembly. J. Cell. Biol. 144, 823–837 (1999).
Sander, C. & Schneider, R. Database of homology-derived protein structures and structural meaning of sequence alignment. Proteins 9, 56–68 (1991).
Altmann, M., Sonenberg, N. & Trachsel, H. Translation in Saccharomyces cerevisiae: initiation factor 4E-dependent cell-free system. Mol. Cell. Biol. 9, 4467–4472 (1989).
Blum, S., Mueller, M., Schmid, S.R., Linder, P. & Trachsel, H. Translation in Saccharomyces cerevisiae: initiation factor 4A-dependent cell-free system. Proc. Natl. Acad. Sci. USA 86, 6043–6046 (1989).
Biffo, S. et al. Isolation of a novel beta4 integrin-binding protein (p27(BBP)) highly expressed in epithelial cells. J. Biol. Chem. 272, 30314–30321 (1997).
Sanvito, F. et al. Expression of a highly conserved protein, p27BBP, during the progression of human colorectal cancer. Cancer Res. 60, 510–516 (2000).
de Pereda, J.M., Wiche, G. & Liddington, R.C. Crystal structure of a tandem pair of fibronectin type III domains from the cytoplasmic tail of integrin alpha6beta4. EMBO J. 18, 4087–4095 (1999).
Burley, S.K. et al. Structural genomics: beyond the human genome project. Nature Genet. 23, 151–157 (1999).
Hendrickson, W. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991).
Holm, L. & Sander, C. A review of the use of protein structure comparison in protein classification and function identification. Science 273, 595–602 (1996).
Nicholls, A., Sharp, K. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Lang, D., Thoma, R., Henn-Sax, M., Sterner, R. & Wilmanns, M. Structural evidence for evolution of the beta/alpha barrel scaffold by gene duplication and fusion. Science 289, 1546–1550 (2000).
Fulop, V. & Jones, D.T. Beta propellers: structural rigidity and functional diversity. Curr. Opin. Struct. Biol. 9, 715–721 (1999).
Sprague, E.R., Redd, M.J., Johnson, A.D. & Wolberger, C. Structure of the C-terminal domain of Tup1, a corepressor of transcription in yeast. EMBO J. 19, 3016–3027 (2000).
Sanchez, R. et al. MODBASE, a database of annotated comparative protein structure models. Nucleic Acids Res. 28, 250–253 (2000).
Sanchez, R. & Sali, A. Comparative protein structure modeling in genomics. J. Comp. Phys. 151, 388–401 (1999).
Nikolov, D.B. et al. Crystal structure of TFIID TATA-box binding protein. Nature 360, 40–46 (1992).
Nicholls, A., Sharp, K. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Conte, L.L., Chothia, C. & Janin, J. The atomic structure of protein-protein recognition sites. J. Mol. Biol. 285, 2177–2198 (1999).
Wall, M.A. et al. The structure of the G protein heterotrimer Gi alpha 1 beta 1 gamma 2. Cell 83, 1047–1058 (1995).
Beckmann, R. et al. Alignment of conduits for the nascent polypeptide chain in the ribosome-Sec61 complex. Science 278, 2123–2126 (1997).
Bult, C.J. et al. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273, 1058–1073 (1996).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Otwinowski, Z. Proceedings of the Daresbury CCP4 study weekend (1991).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A. et al. Crystallography and NMR system: a new software system for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Sheldrick, G.M. Isomorphous replacement and anomalous scattering. In Proc. CCP4 Study Weekend (eds, Wolf, W., Evans, P.R. & Leslie, A.G.W.) 23–28 (Daresbury Laboratory, Warrington, UK; 1991).
Laskowski, R.J., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–290 (1993).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Lamzin, V.S. & Wilson, K.S. Automated refinement of protein models. Acta Crystallogr. D 49, 129–149 (1993).
Henikoff, S. & Henikoff, J.G. Amino acid substitution matrices from protein blocks. Proc. Natl. Acad. Sci. USA 89, 10915–10919 (1992).
Gilson, M., Sharp, K. & Honig, B. Calculating the electrostatic potential of molecules in solution: method and error assessment. J. Comput. Chem. 9, 327–335 (1988).
Acknowledgements
At the Brookhaven National Laboratory National Synchrotron Light Source, we thank Z. Dauter and K. R. Rajashankar for their help using beamline X9B. We thank G. Blobel, J. Bonanno, B.T. Chait, M.R. Chance, J. DeAngelis, R.C. Deo, M. Henderson, V. Ilyin, D. Jeruzalmi, J. Kuriyan, C. Kielkopf, C. Lima, J. Marcotrigiano, F. Melo, S. Nair, U. Pieper, G.A. Petsko, S.S. Ray, R. Sanchez, L. Shapiro, F.W. Studier, S. Swaminathan, R.M. Sweet, and W.B. Whitman for many useful discussions. We thank G. He for technical help and T. Niven for editorial assistance. S.K.B. is a member of the Howard Hughes Medical Institute. This work was supported by National Institutes of Health Grants (S.K.B. and A.S.), and by The Rockefeller University (C.M.G.). A.S. is an Alfred P. Sloan Research Fellow and an Irma T. Hirschl Trust Career Scientist, and is also support by The Merck Genome Research Institute, G. Harold and Leila Y. Mathers Charitable Foundation, and the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Groft, C., Beckmann, R., Sali, A. et al. Crystal structures of ribosome anti-association factor IF6. Nat Struct Mol Biol 7, 1156–1164 (2000). https://doi.org/10.1038/82017
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/82017
This article is cited by
-
Distinct genetic pathways define pre-malignant versus compensatory clonal hematopoiesis in Shwachman-Diamond syndrome
Nature Communications (2021)
-
Diverse roles of assembly factors revealed by structures of late nuclear pre-60S ribosomes
Nature (2016)
-
Involvement of eIF6 in external mechanical stretch–mediated murine dermal fibroblast function via TGF-β1 pathway
Scientific Reports (2016)
-
eIF6 over-expression increases the motility and invasiveness of cancer cells by modulating the expression of a critical subset of membrane-bound proteins
BMC Cancer (2015)
-
Structure of the pre-60S ribosomal subunit with nuclear export factor Arx1 bound at the exit tunnel
Nature Structural & Molecular Biology (2012)