Abstract
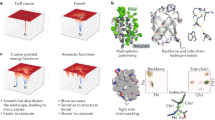
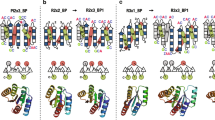
For most proteins the amino acid sequence determines the tertiary structure. The relative importance of the individual amino acids in specifying the fold, however, remains unclear. To highlight this. Creamer and Rose put forth the ‘Paracelsus challenge’: Design a protein with 50% sequence identity to a protein with a different fold. We have met this challenge by designing a sequence which retains 50% identity to a predominantly β-sheet protein, but which now adopts a four helix bundle conformation and possesses the attributes of a native protein. Our results emphasize that a subset of the amino acid sequence is sufficient to specify a fold, and have implications both for structure prediction and design.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Anfinsen, C. Principles that govern the folding of protein chains. Science 181, 223–230 (1973).
Lattman, E.E. & Rose, G.D. Protein folding-what's the question? Proc. Natl. Acad. Sci. USA 90, 439–441 (1993).
Rose, G.D. & Creamer, T.P. Protein folding: predicting predicting. Proteins: Struct. Funct. Genet. 19, 1–3 (1994).
Jones, D.T. et al. Towards Meeting the Paracelsus challenge: The design, synthesis, and characterization of paracelsin-43, an α-helical protein with over 50% sequence identity to an all-β protein. Proteins: Struct. Funct. Genet. 24, 502–513 (1996).
Gronenborn, A.M. et al. a novel, highly stable fold of the immunoglobulin binding domain of streptococcal protein G. Science 253, 657–661 (1991).
Banner, D., Kokkinidis, M. & Tsernoglou, D. Structure of the ColE1 Rop protein at 1.7 Å resolution. J. Mol. Biol. 196, 657–675 (1987).
Smith, C. & Regan, L. Guidelines for protein design: the energetics of β sheet side chain interactions. Science 270, 980–982 (1995).
Chakrabartty, A. & Baldwin, R.L. Stability of α-helices. Adv. Prot. Chem. 46, 141–76 (1995).
Kim, C.A. & Berg, J.M. Thermodynamic β-sheet propensities measured using a zinc-finger host peptide. Nature 362, 267–270 (1993).
Smith, C.K., Withka, J.M. & Regan, LA Thermodynamic scale for the beta-sheet forming tendencies of the amino acids. Biochemistry 33, 5510–5517 (1994).
Minor, Jr., D.L. & Kim, P.S. Measuring the β-sheet forming propensities of amino acids. Nature 367, 660–663 (1994).
Munson, M. et al. What makes a protein a protein? Hydrophobic core designs that specify stability and structural properties. Prot. Sci. 5, 1584–1593 (1996).
Predki, P.F., Nayak, L.M., Gottlieb, M.B. & Regan, L. Dissecting RNA-protein Interactions: RNA-RNA Recognition by Rop Cell 80, 41–50 (1995).
Gamier, J., Osguthorpe, D.J. & Robson, B. Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. J. Mol. Biol. 120, 97–120 (1978).
Altieri, A.S., Hinton, D.P. & Byrd, R.A. Association of bimolecular systems via pulsed field gradient NMR self-diffusion measurements. J. Am. Chem. Soc. 117, 7566–7567 (1995).
Stejskal, E.O. & Tanner, J.E. Spin Diffusion measurements: spin echoes in the presence of a time-dependent field gradient. J. Chem. Phys. 42, 288–292 (1965).
Regan, L. & DeGrado, W.F. Characterization of a helical protein designed from first principles. Science 241, 976–978 (1988).
Hecht, M.H., Richardson, J.S., Richardson, D.C. & Ogden, R.C. De novo design, expression, and characterization of Felix: a four-helix bundle protein of native-like sequence. Science 249, 884–891 (1990).
Kamtekar, S., Schiffer, J.M., Xiong, H., Babik, J.M. & Hecht, M.H. Protein design by binary patterning of polar and nonpolar amino acids. Science 262, 1680–1685 (1993).
Piantinin, U., Sorensen, O.W. & Ernst, R.R. Multiple quantum filters for elucidating NMR coupling networks. J. Am. Chem. Soc. 104, 6800–6801 (1982).
Braunschweiler, L. & Ernst, R.R. Coherence transfer by isotropic mixing: application to proton correlation spectroscopy. J. Magn. Reson. 53, 521–528 (1983).
Kabsch, W. & Sander, C. On the use of sequence homologies to predict protein structure: identical pentapeptides can have completely different conformations. Proc. Natl. Acad. Sci USA 81, 1075–1078 (1984).
Bowie, J.U., Reidhaar-Olson, J.F., Lim, W.A., Sauer, R.T. Deciphering the message in protein sequences: tolerance to amino acid substitutions. Science 247, 1306–1310 (1990).
Bowie, J.U., Luthy, R. & Eisenberg, D.A. Method to identify protein sequences that fold into a known three-dimensional structure. Science 253, 164–170 (1991).
Munson, M., Predki, P.F. & Regan, L. ColE1-compatible vectors for high-level expression of cloned DNAs from the T7 promoter. Gene 144, 59–62 (1994).
Alexander, P., Fahnestock, S., Lee, T., Orban, J. & Bryan, P. Thermodynamic analysis of the folding of the streptococcal protein G IgG-binding domains B1 and B2: Why small proteins tend to have high denaturation temperatures. Biochemistry 31, 3597–3603 (1992).
Tanner, J.E. Use of the stimulated echo in NMR diffusion studies. J. Chem. Phys. 52, 2523–2526 (1970).
Kay, L.E., Keifer, P., Saarinen, T. Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J. Am. Chem. Soc. 114, 10663–10665 (1992).
Kraulis, P.J. Molscript: A program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Predki, P.F., Agrawal, V., Brünger, AT. & Regan, L. Amino-acid substitutions in a surface turn modulate protein stability. Nature Struct. Biol. 3, 54–58 (1996).
Smith, C.K., Munson, M. & Regan, L. Studying α-helix and β-sheet formation in small proteins. Techniques in Protein Chemistry, Vol VI (Academic Press, San Diego, CA, 1995).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Dalal, S., Balasubramanian, S. & Regan, L. Protein alchemy: Changing β-sheet into α-helix. Nat Struct Mol Biol 4, 548–552 (1997). https://doi.org/10.1038/nsb0797-548
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0797-548
This article is cited by
-
An in silico approach to study the role of epitope order in the multi-epitope-based peptide (MEBP) vaccine design
Scientific Reports (2022)
-
Examination of the quality of various force fields and solvation models for the equilibrium simulations of GA88 and GB88
Journal of Molecular Modeling (2016)
-
Replica exchange molecular dynamics simulation of structure variation from α/4β-fold to 3α-fold protein
Journal of Molecular Modeling (2012)
-
Why similar protein sequences encode similar three-dimensional structures?
Theoretical Chemistry Accounts (2010)
-
A Reexamination of Correlations of Amino Acids with Particular Secondary Structures
The Protein Journal (2009)