Abstract
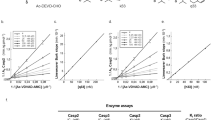
Cysteine proteases related to mammalian interleukin-1β converting enzyme (ICE) and to its Caenorhabditis elegans homologue, CED-3, play a critical role in the biochemical events that culminate in apoptosis. We have determined the three-dimensional structure of a complex of the human CED-3 homologue CPP32/apopain with a potent tetrapeptide-aldehyde inhibitor. The protein resembles ICE in overall structure, but its S4 subsite is strikingly different in size and chemical composition. These differences account for the variation in specificity between the ICE- and CED-3-related proteases and enable the design of specific inhibitors that can probe the physiological functions of the proteins and disease states with which they are associated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Thornberry, N.A. et al. A novel heterodimeric cysteine protease is required for interleukin-1β processing in monocytes. Nature 356, 768–774 (1992).
Cerretti, D.P. et al. Molecular cloning of the interleukin-1β converting enzyme. Science 256, 97–100 (1992).
Munday, N.A. et al. Molecular-cloning and pro-apoptotic activity of ICE(Rel)II and ICE(Rel) III, members of the ICE/CED-3 family of cysteine proteases. J Biol. Chem. 270, 15870–15876 (1995).
Faucheu, C. et al. A novel human protease similar to the interleukin-1β converting- enzyme induces apoptosis in transfected cells. EMBO J. 14, 1914–1922 (1995).
Kamens, J. et al. Identification and characterization of ICH-2 a novel member of the interleukin–1β–converting enzyme family of cysteine proteases. J. Biol. Chem. 270, 15250–15256 (1995).
Kumar, S., Kinoshita, M., Noda, M., Copeland, N.G. & Jenkins, N.A. Induction of apoptosis by the mouse nedd2 gene, which encodes a protein similar to the product of the Caenorhabditis elegans cell-death gene CED-3 and the mammalian IL1-β-converting enzyme. Genes & Development 8, 1613–1626 (1994).
Wang, L., Miura, M., Bergeron, L., Zhu, H. & Yuan, J.Y. ICH-1, an ICE/CED-3-related gene, encodes both positive and negative regulators of programmed cell-death. Cell 78, 739–750 (1994).
Fernandes-Alnemri, T., Litwack, G. & Alnemri, E.S. CPP32, a novel human apoptotic protein with homology to Caenorhabditis elegans cell-death protein CED-3 and mammalian interleukin-1-β- converting enzyme. J. Biol. Chem. 269, 30761–30764 (1994).
Fernandes-Alnemri, T., Litwack, G. & Alnemri, E.S. MCH2, a new member of the apoptotic CED-3/ICE cysteine protease gene family. Cancer Res. 55, 2737–2742 (1995).
Fernandes-Alnemri, T. et al. MCH3, a novel human apoptotic cysteine protease highly related to CPP32. Cancer Res. 55, 6045–6052 (1995).
Duan, H., Chinnaiyan, A.M., Hudson, P.L, Wing, J.P., He, W.-W. & Dixit, V.M. ICE-LAP3, a novel mammalian homolog of the Caednorhabditis elegans cell death protein CED-3 in activated during FAS and tumor necrosis factor-induced apoptosis. J. Biol. Chem. 271, 1621–1625 (1996).
Li, P. et al. Mice deficient in il-1β-converting enzyme are defective in production of mature il-1β and resistant to endotoxic shock. Cell 80, 401–411 (1995).
Kuida, K. et al. Altered cytokine export and apoptosis in mice deficient in interleukin-1β converting-enzyme. Science 267, 2000–2003 (1995).
Ellis, R.E., Yuan, J.Y. & Horvitz, H.R. Mechanisms and functions of cell death. Annu Rev Cell Biol 7, 663–698 (1991).
Miura, M., Zhu, H., Rotello, R., Hartwieg, E.A. & Yuan, J.Y. Induction of apoptosis in fibroblasts by IL-1β-converting enzyme, a mammalian homolog of the C. elegans cell-death gene CED-3. Cell 75, 653–660 (1993).
Yuan, J.Y., Shaham, S., Ledoux, S., Ellis, H.M. & Horvitz, H.R., The, C. The C. elegans cell-death gene CED-3 encodes a protein similar to mammalian interleukin-1-β-converting enzyme. Cell 75, 641–652 (1993).
Kaufmann, S.H., Desnoyers, S., Ottaviano, Y., Davidson, N.E. & Poirier, G.G. Specific proteolytic cleavage of poly(ADP-ribose) polymerase: an early marker of chemotherapy-induced apoptosis. Cancer Res. 53, 3976–3985 (1993).
Lazebnik, Y.A., Kaufmann, S.H., Desnoyers, S., Poirier, G.G. & Earnshaw, W.C. Cleavage of poly(ADP-ribose) polymerase by a proteinase with properties like ICE. Nature 371, 346–347 (1994).
Nicholson, D.W. et al. Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376, 37–43 (1995).
Tewari, M. et al. Yama/CPP32 beta, a mammalian homolog of CED-3, is a CrmA-inhibitable protease that cleaves the death substrate poly(ADP-ribose) polymerase. Cell 81, 801–809 (1995).
Casciola-Rosen, L.A., Miller, D.K., Anhalt, G.J & Rosen, A. Specific cleavage of the 70-kDa protein-component of the u1 small nuclear ribonucleoprotein is a characteristic biochemical feature of apoptotic cell-death. J. Biol. Chem. 269, 30757–30760 (1994).
Casciola-Rosen, L.A., Anhalt, G.J. & Rosen, A. DNA-dependent protein kinase is one of a subset of autoantigens specifically cleaved early during apoptosis. J. Exp. Med. 182, 1625–1634 (1995).
Casciola-Rosen, L.A. et al. Apopain/CPP32 cleaves proteins that are essential for cellular repair: a fundamental principle of apoptotic death. J. Exp. Med. in press (1996). (AUTHOR: STATUS?)
Martin, S.J. et al. Proteolysis of fodrin (non-erythroid spectrin) during apoptosis. J Biol. Chem. 270, 6425–6428 (1995).
Lazebnik, Y.A. et al. Studies of the lamin proteinase reveal multiple parallel biochemical pathways during apoptotic execution. Proc. Natal. Acad. Sci. USA 92, 9042–9046 (1995).
Brancolini, C., Benedetti, M. & Schneider, C. Microfilament reorganization during apoptosis -the role of gas2, a possible substrate for ICE-like proteases. EMBOJ. 14, 5179–5190 (1995).
Emoto, Y. et al. Proteolyt ic activation of protein kinase Cδ by an ICE-like protease in apoptotic cells. EMBO J. 14, 6148–6156 (1995).
Wilson, K.P. et al. Structure and mechanism of interleukin-1β converting-enzyme. Nature 370, 270–275 (1994).
Walker, N.P.C. et al. Crystal-structure of the cysteine protease interleukin-1β-converting enzyme - a (p20/p10)(2) homodimer. Cell 78, 343–352 (1994).
Thornberry, N.A., Miller, D.K. & Nicholson, D.W. Interleukin-1β converting enzyme and related proteases as potential targets in inflammation and apoptosis. Perspectives in Drug Discovery and Design 2, 389–399 (1995).
Westerik, J.O. & Wolfenden, R. Aldehydes as inhibitors of papain. J. Biol. Chem. 247, 8195–8197 (1972).
Ortiz, C., Tellier, C., Williams, H., Stolowich, N.J. & Scott, A.I. Diastereotopic covalent binding of the natural inhibitor leupeptin to trypsin: detection of two interconverting hemiacetals by solution and solid-state NMRspectroscopy. Biochemistry 30, 10026–10034 (1991).
Delbaere, L.T. & Brayer, G.D. The 1. 8 Å structure of the complex between chymostatin and Streptomyces griseus protease A. A model for serine protease catalytic tetrahedral intermediates. J. Mol. Biol. 183, 89–103 (1985).
Frankfater, A. & Kuppy, T. Mechanism of association of N-acetyl-L-phenylalanylglycinal to papain. Biochemistry 20, 5517–5524 (1981).
Mackenzie, N.E., Grant, S.K., Scott, A.I. & Malthouse, J.P. 13C NMR study of the stereospecificity of the thiohemiacetals formed on inhibition of papain by specific enantiomeric aldehydes. Biochemistry 25, 2293–2298 (1986).
Menard, R. et al. Contribution of the glutamine 19 side chain to transition-state stabilization in the oxyanion hole of papain. Biochemistry 30, 8924–8928 (1991).
SAINT Software Reference Manual (Siemens Analytical Instruments, Madison, Wisconsin, 1995).
Brünger, A.T. X-PLOR: Version 3.1, a System forX-Ray Crystallography and NMR (Yale University Press, New Haven 1992).
Abola, E.E., Bernstein, F.C., Bryant, S.H., Koetzle, T.F. & Weng, J. in Crystallographic Databases - Information Content, Software Systems, Scientific Applications (eds. Alien, F.H., Bergerhoff, G. & Sievers, R.) 107–132 (Data Commission of the International Union of Crystallography, Bonn/Cambridge/Chester, 1987).
Sack, J.S. CHAIN - a crystallographic modeling program. J. Mol. Graphics 6, 224–225 (1988).
Zhang, K.Y.J. SQUASH-combining constraints for macromolecular phase refinement and extension. Acta Crystallogr. D49, 213–222 (1993).
Hodel, A., Kim, S.-H. & Brünger, A.T. Model bias in macromolecular crystal structures. Acta Crystallogr. A48, 851–858 (1992).
Frishman, D. & Argos, P. Knowledge-based protein secondary structure assignment. Proteins Struct. Funct. Genet. 23, 566–579 (1995).
Carson, M. Ribbon models of macromolecules. J. Molec. Graph. 5, 103–106 (1987).
QUANTA User Guide (Molecular Simulations, San Diego, 1996).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Rotonda, J., Nicholson, D., Fazil, K. et al. The three-dimensional structure of apopain/CPP32, a key mediator of apoptosis. Nat Struct Mol Biol 3, 619–625 (1996). https://doi.org/10.1038/nsb0796-619
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0796-619