Abstract
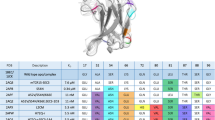
Using NMR spectroscopy, we determined the solution structure of a single-chain T-cell receptor (scTCR) derived from the major histocompatibility complex (MHC) class II-restricted D10 TCR. The conformations of complementarity-determining regions (CDRs) 3β and 1α and surface properties of 2α are different from those of related class I-restricted TCRs. We infer a conserved orientation for TCR Vα domains in complexes with both class I and II MHC–peptide ligands, which implies that small structural variations in Vα confer MHC class preference. High mobility of CDR3 residues relative to other CDR or framework residues (picosecond time scale) provides insight into immune recognition and selection mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Meuer, S.C., Acuto, O., Hercend, T., Schlossman, S.F., Reinherz, E.L. The human T-cell receptor. Annu. Rev. Immunol. 2, 23–50 (1984).
Marrack, P. & Kappler, J. The antigen-specific, major histocompatibility complex-restricted receptor on T cells. Adv. Immunol. 38, 1–30 (1986).
Zinkernagel, R.M. & Doherty, P.C. Restriction of in vitro T cell-mediated cytotoxicity in lymphocytic choriomeningitis within a syngeneic or semiallogeneic system. Nature 248, 701–702 (1974).
Meuer, S.C., Schlossman, S.F. & Reinherz, E.L. Clonal analysis of human cytotoxic T lymphocytes: T4+ and T8+ effector T cells recognize products of different major histocompatibility complex regions. Proc. Natl. Acad. Sci. USA 79, 4395–4399 (1982).
Biddison, W.E. Rao, P.E., Talle, M.A., Goldstein, G. & Shaw, S. Possible involvement of the OKT4 molecule in T cell recognition of class II HLA antigens. Evidence from studies of cytotoxic T lymphocytes specific for SB antigens. J. Exp. Med. 156, 1065–1076 (1982).
Krensky, A.M. Reiss, C.S., Mier, J.W., Strominger, J.L. & Burakoff, S.J. Long-term human cytolytic T-cell lines allospecific for HLA-DR6 antigen are OKT4+. Proc. Natl. Acad. Sci. USA 79, 2365–2369 (1982).
Meuer, S.C. et al. Surface structures involved in target recognition by human cytotoxic T lymphocytes. Science 218, 471–473 (1982).
Utsunomiya, Y. et al. Analysis of a monoclonal rat antibody directed to the alpha-chain variable region (V alpha 3) of the mouse T cell antigen receptor. J. Immunol. 143, 2602–2608 (1989).
Garboczi, D.N. et al. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384, 134–141 (1996).
Garcia, K.C. et al. Structural basis of plasticity in T cell receptor recognition of a self peptide–MHC antigen. Science 279, 1166–1172 (1998).
Ding, Y.-H. et al. Two human T cell receptors bind in a similar diagonal mode to the HLA-A2/Tax peptide complex using different TCR amino acids. Immunity 8, 403–411 (1998).
Teng, M.-K. et al. Identification of a common docking topology with substantial variation among different TCR–peptide–MHC complexes. Curr. Biol. 8, 409–412 (1998).
Kaye, J., Porcelli, S., Tite, J., Jones, B. & Janeway, C.A. Jr. Both a monoclonal antibody and antisera specific for determinants unique to individual cloned helper T cell lines can substitute for antigen and antigen presenting cells in the activation of T cells. J. Exp. Med. 158, 836–856 (1983).
Novotny, J. et al. A soluble, single-chain T-cell receptor fragment endowed with antigen-combining properties. Proc. Natl. Acad. Sci USA 88, 8646–8650 (1991).
Khandekar, S.S. et al. Conformational integrity and ligand binding properties of a single chain T-cell receptor expressed in Escherichia coli. J. Biol. Chem 272, 32190–32197 (1997).
Garcia, K.C. et al. An αβ T cell receptor structure at 2.5 Å and its orientation in the TCR–MHC complex. Science 274, 209–219 (1996).
Wang, J. et al. Atomic structure of an αβ T cell receptor (TCR) heterodimer in complex with an anti-TCR Fab fragment derived from a mitogenic antibody. EMBO J. 17, 10–26 (1998).
Housset, D. et al. The three-dimensional structure of a T-cell antigen receptor VαVβ heterodimer reveals a novel arrangement of the Vβ domain. EMBO J. 16, 4205–4216 (1997).
Richardson, J.S. The anatomy and taxonomy of protein structure. Adv. Protein Chem. 34, 167–339 (1981).
Bentley, G.A., Boulot, G., Karjalainen, K. & Mariuzza, R.A. Crystal structure of the β chain of a T cell antigen receptor. Science 267, 1984–1987 (1995).
Acuto, O., Hussey, R.E. & Reinherz, E.L. Multiple class I and class II major histocompatibility complex allospecificities are generated with T cell receptor variable (V) domains created by a single Ti β V gene family. J. Exp. Med. 162, 1387–1392 (1985).
Stern, L.J. & Wiley, D.C. Antigenic peptide binding by class I and class II histocompatibility proteins. Structure 15, 245–251 (1994).
Jorgenson, J.L., Esser, U., Fazekas de St. Groth, B., Reay, P.A. & Davis, M.M. Mapping T-cell receptor-peptide contacts by variant peptide immunization of single-chain transgenics. Nature 355, 224–230 (1992).
Sant'Angelo, D.B. et al. The specificity and orientation of a TCR to its peptide–MHC class II ligands. Immunity 4, 367–376 (1996).
Rudensky, A.Y., Preston-Hurlburt, P., Hong, S.C., Barlow, A. & Janeway, C.A. Sequence analysis of peptides bound to MHC class II molecules. Nature 353, 622–627 (1991).
Brown, J.H. et al. Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 364, 33–39 (1993).
Vignali, D.A.A. & Strominger, J.L. Amino acid residues that flank core peptide epitopes and the extracellular domains of CD4 modulate differential signaling through the T cell receptor. J. Exp. Med. 179, 1945–1956 (1994).
Arden, B., Clark, S.P., Kabelitz, D. & Mak, T.W. Mouse T-cell receptor variable gene segments. Immunogenetics 42, 501–530 (1995).
Sim, B.-C., Zerva, L., Greene, M.I. & Gascoigne, N.R.J. Control of MHC Restriction by TCR Vα CDR1 and CDR2. Science 273, 963–966 (1996).
Rupp, F et al. T-cell antigen receptors with identical variable regions but different diversity and joining region gene segments have distinct specificities but cross-reactive idiotypes. Proc. Natl. Acad. Sci. USA 84, 219–222 (1987).
Cammarota, G. et al. Identification of a CD4 binding site on the β2 domain of HLA-DR molecules. Nature 356, 799–801 (1992).
König, R., Huang, L.-Y. & Germain, R.N. MHC class II interaction with CD4 mediated by a region analogous to the MHC class I binding site for CD8. Nature 356:796–798 (1992).
Norment, A.M., Salter, R.D., Parham, P., Engelhard, V.H. & Littman, D.R. Cell–cell adhesion mediated by CD8 and MHC class I molecules. Nature 336:79–81.
Farrow, N.A., Zhang, O., Forman-Kay, J.D. & Kay, L.E. Comparison of the backbone dynamics of a folded and an unfolded SH3 domain existing in equilibrium in aqueous buffer. Biochemistry 34, 868–878 (1995).
Markus, M.A., Dayie, K.T., Matsudaira, P. & Wagner, G. Local mobility within villin 14T probed via heteronuclear relaxation measurements and a reduced spectral density mapping. Biochemistry 35, 1722–1732 (1996).
Wyss, D.F. Dayie, K.T. & Wagner, G. The counterreceptor binding site of human CD2 exhibits an extended surface patch with multiple conformations fluctuating with millisecond to microsecond motions. Protein Sci. 6, 534–542 (1997).
Takahashi, H., Suzuki, E., Shimada, I. & Arata, Y. Dynamical structure of the antibody combining site as studied by 1H-15N shift correlation NMR spectroscopy. Biochemistry 31, 2464–2468 (1992).
Abergel, C. & Claverie, J-M. A strong propensity toward loop formation characterizes the expressed reading frames of the D segments at the Ig H and T cell receptor loci. Eur. J. Immunol. 21, 3021–3025 (1991).
Gagné, S.M., Tsuda, S., Spyracopoulos, L., Kay, L.E. & Sykes, B.D. Backbone and methyl dynamics of the regulatory domain of troponin C: anisotropic rotational diffusion and contribution of conformational entropy to calcium affinity. J. Mol. Biol. 278, 667–686 (1998).
Manning, T.C. et al. Alanine scanning mutagenesis of an alpha beta T cell receptor: mapping the energy of antigen recognition. Immunity 8, 413–425 (1998).
Ghendler, Y. et al. Differential thymic selection outcomes stimulated by focal structural alteration in peptide/major histocompatibility complex ligands. Proc. Natl. Acad. Sci USA 95, 10061–10066 (1998).
Bevan, M.J. In thymic selection, peptide diversity gives and takes away. Immunity 7, 175–178 (1997).
Liu, C.-P., Parker, D., Kappler, J. & Marrack, P. Selection of antigen-specific T cells by a single IEk peptide combination. J. Exp. Med. 186, 1441–1450 (1997).
Evavold, B.D., Sloan-Lancaster, J., Wilson, K.J., Rothbard, J.B. & Allen, P.M. Specific T cell recognition of minimally homologous peptides: evidence for multiple endogenous ligands. Immunity 2, 655–663 (1995).
Piotto, M, Saudek, V. & Sklenar, V. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J. Biomol. NMR 2, 661–665 (1992).
Yamazaki, T. et al. An HNCA pulse scheme for the backbone assignment of 15N, 13C, 2H-labeled proteins: application to a 37-kDa trp repressor–DNA complex. J. Am. Chem. Soc. 116, 6464–6465 (1994).
Talluri, S. & Wagner, G. An optimized 3D NOESY-HSQC. J. Magn. Reson. 112B, 200–205 (1996).
Del Rio-Portilla, F., Blechta, V. & Freeman, R.M. Measurement of poorly resolved splittings by J doubling in the frequency domain. J. Magn. Reson. 111A, 132–135 (1994).
Güntert, P., Braun, W., Billeter, M. & Wüthrich, K. Automated stereospecific 1H NMR assignments and their impact on the precision of protein structure determination in solution. J. Am. Chem. Soc. 111, 3997–4004 (1989).
Nilges, M., Clore, G.M. & Gronenborn, A.M. Determination of three-dimensional structures of proteins from interproton distance data by dynamical simulated annealing from a random array of atoms. FEBS Lett. 239, 129–136 (1988).
Brünger, A.T. X-PLOR version 3.1: a system for X-ray crystallography and NMR. (Yale University Press, New Haven, Connecticut; 1993).
Dayie, K.T. & Wagner, G. Relaxation-rate measurements for 15N-1H groups with pulsed-field gradients and preservation of coherence pathways. J. Magn. Reson. 111A, 121–126 (1994).
Peng, J.W. & Wagner, G. Mapping of spectral density functions using heteronuclear NMR relaxation measurements. J. Magn. Reson. 98, 308–322 (1992).
Kay, L.E., Torchia, D.A. & Bax, A. Backbone dynamics of proteins as studied by 15N inverse detected heteronuclear NMR spectroscopy: application to staphylococcal nuclease. Biochemistry 28, 8972–8979 (1989).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
This work was supported by the Cancer Research Institute (fellowship to B.J.H.) and the National Institutes of Health (G.W. and E.L.R.). P.S.K. acknowledges support from the Schweizerischer Nationalfond and Schweizerische Krebsliga. We thank D. Austen and K. Huestis for technical assistance. We thank S. Khandekar and B. Bettencourt for helpful discussions. We thank F. Del Rio-Portilla for assistance with coupling constant measurements.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hare, B., Wyss, D., Osburne, M. et al. Structure, specificity and CDR mobility of a class II restricted single-chain T-cell receptor. Nat Struct Mol Biol 6, 574–581 (1999). https://doi.org/10.1038/9359
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/9359
This article is cited by
-
T-Cell Receptor Variable β Domains Rigidify During Affinity Maturation
Scientific Reports (2020)
-
NMR: an essential structural tool for integrative studies of T cell development, pMHC ligand recognition and TCR mechanobiology
Journal of Biomolecular NMR (2019)
-
Backbone resonance assignment of N15, N30 and D10 T cell receptor β subunits
Biomolecular NMR Assignments (2016)
-
Chaperone-assisted thermostability engineering of a soluble T cell receptor using phage display
Scientific Reports (2013)
-
NMR spectroscopy reveals unexpected structural variation at the protein–protein interface in MHC class I molecules
Journal of Biomolecular NMR (2013)