Abstract
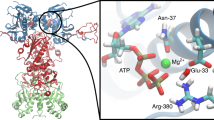
How substrate affinity is modulated by nucleotide binding remains a fundamental, unanswered question in the study of 70 kDa heat shock protein (Hsp70) molecular chaperones. We find here that the Escherichia coli Hsp70, DnaK, lacking the entire α-helical domain, DnaK(1–507), retains the ability to support λ phage replication in vivo and to pass information from the nucleotide binding domain to the substrate binding domain, and vice versa, in vitro. We determined the NMR solution structure of the corresponding substrate binding domain, DnaK(393–507), without substrate, and assessed the impact of substrate binding. Without bound substrate, loop L3,4 and strand β3 are in significantly different conformations than observed in previous structures of the bound DnaK substrate binding domain, leading to occlusion of the substrate binding site. Upon substrate binding, the β-domain shifts towards the structure seen in earlier X-ray and NMR structures. Taken together, our results suggest that conformational changes in the β-domain itself contribute to the mechanism by which nucleotide binding modulates substrate binding affinity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Bukau, B. & Horwich, A.L. Cell 92, 351–366 (1998).
Flaherty, K.M., DeLuca-Flaherty, C. & McKay, D.B. Nature 346, 623– 628 (1990).
Zhu, X. et al. Science 272, 1606–1614 (1996).
Wang, H. et al. Biochemistry 37, 7929–7940 (1998).
Morshauser, R.C. et al. J. Mol. Biol. 289, 1387– 1403 (1999).
Bertelsen, E.B., Zhou, H., Lowry, D.F., Flynn, G.C. & Dahlquist, F.W. Protein Sci. 8, 343– 354 (1999).
McCarty, J.S., Buchberger, A, Reinstein, J. & Bukau, B. J. Mol. Biol. 249, 126–137 (1995).
Misselwitz, B., Staeck, O. & Rapoport, T.A. Mol. Cell. 2, 593– 603 (1998).
Montgomery, D.L., Morimoto, R.I. & Gierasch, L.M. J. Mol. Biol. 286, 915– 932 (1999).
Yochem, J. et al. Mol. Gen. Genet. 164, 9– 14 (1978).
Bukau, B. & Walker, G.C. EMBO J. 9, 4027–4036 (1990).
Pierpaoli, E.V., Gisler, S.M. & Christen, P. Biochemistry 37, 16741– 16748 (1998).
Rüdiger, S., Buchberger, A. & Bukau, B. Nature Struct. Biol. 4, 342– 349 (1997).
Montgomery, D.L., Jordan, R., McMacken, R. & Freire, E. J. Mol. Biol. 232, 680–692 (1993).
Gragerov, A., Zeng, L., Zhao, X., Burkholder, W. & Gottesman, M.E. J. Mol. Biol. 235, 848– 854 (1994).
Buchberger, A. et al. J. Biol. Chem. 270, 16903– 16910 (1995).
Laufen, T. et al. Proc. Natl. Acad. Sci. USA 96, 5452 –5457 (1999).
Flynn, G.C., Pohl, J., Flocco, M.T. & Rothman, J.E. Nature 353, 726–730 (1991).
Rost, B. & Sander, C. J. Mol. Biol. 232, 584–599 (1993).
Burkholder, W.F. et al. Proc. Natl. Acad. Sci. USA 93, 10632 –10637 (1996).
Voisine, C. et al. Cell 97, 565–574 (1999).
Cavanagh, J., Fairbrother, W.J. & Palmer, A.G.III, Skelton, N.J. Protein NMR spectroscopy (Academic Press, San Diego, 1996).
Neri, D., Szyperski, T., Otting, G., Senn, H. & Wüthrich, K. Biochemistry 28 , 7510–7516 (1989).
Güntert, P., Dötsch, V., Wider, G. & Wüthrich, K. J. Biomol. NMR 2, 619–629 (1992).
Bartels, C., Xia, T., Billeter, M., Güntert, P. & Wüthrich, K. J. Biomol. NMR 6, 1–10 (1995).
Güntert, P., Mumenthaler, C. & Wüthrich, K. J. Mol. Biol. 273, 283– 298 (1997).
Schaumann, T., Braun, W. & Wüthrich, K. Biopolymers 29, 679– 694 (1990).
Koradi, R., Billeter, M. & Wüthrich, K. J. Mol. Graphics 14, 52– 55 (1996).
Carrington, A. & McLachlan, A. Introduction to magnetic resonance with applications to chemistry and chemical physics . (Harper & Row, New York, 1967).
Laskowski, R.A., Rullmann, J.A.C., MacArthur, M.W., Kaptein, R. & Thornton, J.M. J. Biomol. NMR 8, 477–486 (1996).
Acknowledgements
This work was supported by NIH grants to E.R.P.Z and to L.M.G., and a NIH fellowship to D.L.M.. The W.M. Keck Foundation, NIH, NSF and Parke-Davis/ Warner Lambert are gratefully acknowledged for financial support towards the 800 MHz NMR instrument. We thank J. Feltham for critical reading of the manuscript, and R. Sivendran for help with the assays of peptide-stimulated ATPase activity.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Pellecchia, M., Montgomery, D., Stevens, S. et al. Structural insights into substrate binding by the molecular chaperone DnaK. Nat Struct Mol Biol 7, 298–303 (2000). https://doi.org/10.1038/74062
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/74062
This article is cited by
-
Direct observation of chaperone-modulated talin mechanics with single-molecule resolution
Communications Biology (2022)
-
Elucidation of transient protein-protein interactions within carrier protein-dependent biosynthesis
Communications Biology (2021)
-
Selective Binding of HSC70 and its Co-Chaperones to Structural Hotspots on CFTR
Scientific Reports (2020)
-
The Hsp70 chaperone network
Nature Reviews Molecular Cell Biology (2019)
-
Activation of the DnaK-ClpB Complex is Regulated by the Properties of the Bound Substrate
Scientific Reports (2018)