Abstract
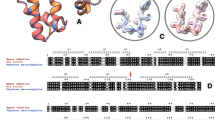
A surface turn position in a four-helix bundle protein, Rop, was selected to investigate the role of turns in protein structure and stability. Although all twenty amino acids can be substituted at this position to generate a correctly folded protein, they produce an unusually large range of thermodynamic stabilities. Moreover, the majority of substitutions give rise to proteins with enhanced thermal stability compared to that of the wild type. By introducing the same twenty mutations at this position, but in a simplified context, we were able to deconvolute intrinsic preferences from local environmental effects. The intrinsic preferences can be explained on the basis of preferred backbone dihedral angles, but local environmental context can significantly modify these effects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Brunet, A.P. et al. The role of turns in the structure of an α-helical protein. Nature 364, 355–358 (1993).
Castagnoli, L., Vetriani, C. & Cesareni, G. Linking an easily detectable phenotype to the folding of a common structural motif: Selection of rare turn mutations that prevent the folding of Rop. J. molec. Biol. 237, 378–387 (1994).
Vlassi, M. et al. Restored heptad pattern continuity does not alter the folding of four-α-helix bundle. Nature struct. Biology 1, 706–716 (1994).
Predki, P.F. & Regan, L. Redesigning the topology of a four-helix-bundle protein: Monomeric Rop. Biochemistry 34, 9834–9839 (1995).
Wright, P.E., Dyson, J.H. & Lerner, R.A. Conformation of peptide fragments of proteins in aqueous solution: Implications for initiation of protein folding. Biochemistry 27, 7167–7175 (1988).
Milburn, P.J., Konishi, Y., Meinwald, Y.C. & Scheraga, H.A. Chain Reversals in model peptides: Studies of cystine-containing cyclic peptides I. Conformational free energies of cyclization of hexapeptides of sequence. Ac-Cys-X-Pro-Gly-Y-Cys-NHMe J. Am. Chem. Soc 109, 4486–4496 (1987).
Dyson, H.J., Ranee, M., Houghten, R.A., Lerner, R.A. & Wright, P.E. Folding of immunogenic peptide fragments of proteins in water solution I. Sequence requirements for the formation of a reverse turn. J. molec. Biol. 201, 161–200 (1988).
Fasman, G.D. Prediction of protein structure and the principles of protein conformation (Plenum, New York, 1989).
Muoz, V., Blanco, F.J. & Serrano, L. The hydrophobic-staple motif and a role for loop-residues in α-helix stability and protein folding. Nature struct Biology 2, 380–385 (1995).
Banner, D.W., Kokkinidis, M. & Tsernoglou, D. Structure of the ColE1 Rop protein at 1.7 Å resolution. J. molec. Biol. 196, 657–675 (1987).
Polisky, B. Col-E-1 replication control circuitry sense from antisense. Cell 55, 929–932 (1988).
Predki, P.F., Nayak, L.M., Gottlieb, M.B.C. & Regan, L. Dissecting RNA-protein interactions: RNA-RNA recognition by Rop. Cell 80, 41–50 (1995).
O-Neil, K.T. & DeGrado, W.F. A thermodynamic scale for the helix-forming tendencies of the commonly occurring amino acids. Science 250, 646–651 (1990).
Horovitz, A., Matthews, J.M. & Fersht, A.R. α-helix stability in proteins II. Factors that influence stability at an internal position J molec. Biol. 227, 560–568 (1992).
Blaber, M. et al. Determination of α-helix propensity within the context of a folded protein: Sites 44 and 131 in bacteriophage T4 lysozyme. J. molec. Biol. 235, 600–624 (1994).
Kim, C.A. & Berg, J.M. Thermodynamic β-sheet propensities measured using a zinc-finger host peptide. Nature 362, 267–270 (1993).
Minor, D.L. Jr. & Kim, P.S. Measurement of the β-sheet-forming propensities of amino acids. Nature 367, 660–663 (1994).
Smith, C.K., Withka, J.M. & Regan, L. A thermodynamic scale for the β-sheet forming tendencies of the amino acids. Biochemistry 33, 5510–5517 (1994).
Minor, D.L. Jr. & Kim, P.S. Context is a major determinant of β-sheet propensity. Nature 371, 264–267 (1994).
Smith, C.K. & Regan, L. Guidelines for protein design: energetics of β-sheet side-chain interactions. Science 270, 980–982 (1995).
Ramachandran, G.N. & Sasisekharan, V. Conformation of polypepties and proteins. Adv. Prot Chem. 23, 283–437 (1968).
Munoz, V., Serrano, L. Intrinsic secondary structure propensities of the amino acids, using statistical phi-psi matrices: Comparison with experimental scales. Proteins 20, 301–311 (1994).
Munson, M., O'Brien, R., Sturtevant, J.M. & Regan, L. Redesigning the hydrophobic core of a four-helix-bundle protein. Prot. Sci. 3, 2015–2022 (1994).
Vriend, G. WHAT IF: A molecular modelling and drug design package. J. molec. Graphics 8, 52–56 (1990).
DeLano, W.L. & Brünger, AT. The direct rotation function: Rotational patterson correlation search applied to molecular replacement. Ada Crystallogr. D, in the press.
Brünger, A.T. Extension of molecular replacement: A new search strategy based on patterson correlation refinement. Acta Crystallogr. A46, 46–57 (1990).
Fuginaga, M. & Read, R.J. Experiences with a new translation-function program. J. appl. Crystallogr. 20, 517–521 (1987).
Brünger, A.T. X-PLOR. Version 3.1 A System for X-ray crystallography and NMR. Yale University Press, New Haven, CT (1992).
Hodel, A., Kim, S.-H. & Brnger, A.T. Model bias in macromolecular crystal structures. Acta Crystallogr. A48, 851–859 (1992).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Brünger, A.T., Krukowski, A. & Erickson, J. Slow-cooling protocols for crystallographic refinement by simulated annealing. Acta Crystallogr A46, 585 (1990).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association. Insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 282–293 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Predki, P., Agrawal, V., Brünger, A. et al. Amino-acid substitutions in a surface turn modulate protein stability. Nat Struct Mol Biol 3, 54–58 (1996). https://doi.org/10.1038/nsb0196-54
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb0196-54
This article is cited by
-
Molecular Cloning and Evolutionary Analysis of Hemoglobin α-Chain Genes in Bats
Biochemical Genetics (2009)
-
Engineering heme binding sites in monomeric rop
JBIC Journal of Biological Inorganic Chemistry (2009)
-
Obligatory steps in protein folding and the conformational diversity of the transition state
Nature Structural & Molecular Biology (1998)
-
Protein alchemy: Changing β-sheet into α-helix
Nature Structural Biology (1997)
-
In vitro evolution of thermodynamically stable turns
Nature Structural Biology (1996)