Abstract
In the past decade, an explosion in the number of high-throughput tools for the measurement of different cellular products has occurred. These tools have the potential to further our understanding of human disease and this development has facilitated the identification of new biomarkers in all areas of medicine. In the field of solid organ transplantation, two different areas have developed: the use of biomarkers to predict allograft tolerance for the identification of patients who can be weaned from immunosuppressive therapy, and biomarkers for the prediction of allograft rejection, so that parenchymal damage can be prevented before it becomes irreversible. In this Review, we discuss the development of biomarkers that are indicative of transplant tolerance. Identifying patients in whom donor-specific tolerance has developed would constitute a major advance in the care of organ transplant recipients. This ability would allow the minimization or even the withdrawal of immunosuppressive therapy in selected patients, thus reducing the number of adverse effects and costs, and optimizing long-term graft outcomes. The routine clinical use of these biomarkers, once validated, would bring to the fore the possibility of personalized medicine.
Key Points
-
The use of biomarkers may allow the possibility of personalized medicine in fields such as renal transplantation
-
In the follow-up of solid organ transplant recipients, validated biomarkers of tolerance may be as clinically useful as biomarkers of rejection
-
Biomarkers of tolerance may be used in the future to identify patients in whom immunosuppression can be weaned or even withdrawn
-
Such biomarkers could also be used to establish the success of tolerance-inducing protocols
-
Cross-platform biomarkers may be more effective at identifying tolerance than single biomarkers
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Afzali, B., Lombardi, G. & Lechler, R. I. Pathways of major histocompatibility complex allorecognition. Curr. Opin. Organ Transplant. 13, 438–444 (2008).
Roussey-Kesler, G. et al. Clinical operational tolerance after kidney transplantation. Am. J. Transplant. 6, 736–746 (2006).
Mazariegos, G. V. et al. Weaning of immunosuppression in liver transplant recipients. Transplantation 63, 243–249 (1997).
Lerut, J. & Sanchez-Fueyo, A. An appraisal of tolerance in liver transplantation. Am. J. Transplant. 6, 1774–1780 (2006).
Martínez-Llordella, M. et al. Using transcriptional profiling to develop a diagnostic test of operational tolerance in liver transplant recipients. J. Clin. Invest. 118, 2845–2857 (2008).
Mazariegos, G. V. et al. Cytokine gene polymorphisms in children successfully withdrawn from immunosuppression after liver transplantation. Transplantation 73, 1342–1345 (2002).
Denhaerynck, K. et al. Prevalence, consequences, and determinants of nonadherence in adult renal transplant patients: a literature review. Transpl. Int. 18, 1121–1133 (2005).
Pinsky, B. W. et al. Transplant outcomes and economic costs associated with patient noncompliance to immunosuppression. Am. J. Transplant. 9, 2597–2606 (2009).
Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin. Pharmacol. Ther. 69, 89–95 (2001).
Biomarkers 2011 The 6th Annual Biomarkers Congress [online], (2010).
Trusheim, M. R., Berndt, E. R. & Douglas, F. L. Stratified medicine: strategic and economic implications of combining drugs and clinical biomarkers. Nat. Rev. Drug Discov. 6, 287–293 (2007).
Tol, J. et al. Chemotherapy, bevacizumab, and cetuximab in metastatic colorectal cancer. N. Engl. J. Med. 360, 563–572 (2009).
FDA U.S. Food and Drug Administration Class Labeling Changes to anti-EGFR Monoclonal Antibodies, Cetuximab (Erbitux) and Panitumumab (Vectibix): KRAS Mutations [online], (2010).
Hamburg, M. A. & Collins, F. S. The path to personalized medicine. N. Engl. J. Med. 363, 301–304 (2010).
Hudis, C. A. Trastuzumab—mechanism of action and use in clinical practice. N. Engl. J. Med. 357, 39–51 (2007).
Samani, N. J., Tomaszewski, M. & Schunkert, H. The personal genome—the future of personalised medicine? Lancet 375, 1497–1498 (2010).
Personalized Medicine Coalition The Case for Personalized Medicine [online], (2009).
Bellomo, R., Kellum, J. A. & Ronco, C. Defining acute renal failure: physiological principles. Intensive Care Med. 30, 33–37 (2004).
Khatri, P. & Sarwal, M. M. Using gene arrays in diagnosis of rejection. Curr. Opin. Organ Transplant. 14, 34–39 (2009).
Peltonen, L. & McKusick, V. A. Genomics and medicine. Dissecting human disease in the postgenomic era. Science 291, 1224–1229 (2001).
The United Kingdom and Ireland Renal Transplant Consortium Defining the Genetic Basis of Interactions Between Donor and Recipient DNA that Determine Early and Late Renal Transplant Dysfunction [online], (2010).
Diamandis, E. P. Mass spectrometry as a diagnostic and a cancer biomarker discovery tool: opportunities and potential limitations. Mol. Cell. Proteomics 3, 367–378 (2004).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Zhang, Y. et al. Analysis of chronic lung transplant rejection by MALDI-TOF profiles of bronchoalveolar lavage fluid. Proteomics 6, 1001–1010 (2006).
Tomizaki, K. Y., Usui, K. & Mihara, H. Protein–protein interactions and selection: array-based techniques for screening disease-associated biomarkers in predictive/early diagnosis. FEBS J. 277, 1996–2005 (2010).
Hofmann, M. & Zerwes, H. G. Identification of organ-specific T cell populations by analysis of multiparameter flow cytometry data using DNA-chip analysis software. Cytometry A 69, 533–540 (2006).
Billingham, L. J. & Abrams, K. R. Simultaneous analysis of quality of life and survival data. Stat. Methods Med. Res. 11, 25–48 (2002).
Bossuyt, P. M. Interpreting diagnostic test accuracy studies. Semin. Hematol. 45, 189–195 (2008).
Bossuyt, P. M. Clinical validity: defining biomarker performance. Scand. J. Clin. Lab. Invest. Suppl. 242, 46–52 (2010).
Lechler, R. I., Garden, O. A. & Turka, L. A. The complementary roles of deletion and regulation in transplantation tolerance. Nat. Rev. Immunol. 3, 147–158 (2003).
Brouard, S. et al. Identification of a peripheral blood transcriptional biomarker panel associated with operational renal allograft tolerance. Proc. Natl Acad. Sci. USA 104, 15448–15453 (2007).
Louis, S. et al. Contrasting CD25hiCD4+T cells/FOXP3 patterns in chronic rejection and operational drug-free tolerance. Transplantation 81, 398–407 (2006).
Velásquez, S. Y., Arias, L. F., García, L. F. & Alvarez, C. M. T cell receptor beta chain (TCR-Vbeta) repertoire of circulating CD4(+) CD25(–), CD4(+) CD25(low) and CD4(+) CD25(high) T cells in patients with long-term renal allograft survival. Transpl. Int. 23, 54–63 (2010).
Indices of Tolerance [online]
Immune Tolerance Network [online].
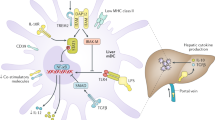
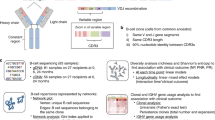
Newell, K. et al. Identification of a B-cell signature associated with renal transplant tolerance in humans. J. Clin. Invest. 120, 1836–1847 (2010).
Sagoo, P. et al. Development of a cross-platform biomarker signature to detect renal transplant tolerance in humans. J. Clin. Invest. 120, 1848–1861 (2010).
Reprogramming the Immune System for Establishment of Tolerance [online]
Mazariegos, G. V. et al. Dendritic cell subset ratio in peripheral blood correlates with successful withdrawal of immunosuppression in liver transplant patients. Am. J. Transplant. 3, 689–696 (2003).
Martínez-Llordella, M. et al. Multiparameter immune profiling of operational tolerance in liver transplantation. Am. J. Transplant. 7, 309–319 (2007).
Koshiba, T. et al. Clinical, immunological, and pathological aspects of operational tolerance after pediatric living-donor liver transplantation. Transpl. Immunol. 17, 94–97 (2007).
Kurian, S. et al. Applying genomics to organ transplantation medicine in both discovery and validation of biomarkers. Int. Immunopharmacol. 7, 1948–1960 (2007).
Li, B. et al. Noninvasive diagnosis of renal-allograft rejection by measurement of messenger RNA for perforin and granzyme B in urine. N. Engl. J. Med. 344, 947–954 (2001).
Muthukumar, T. et al. Serine proteinase inhibitor-9, an endogenous blocker of granzyme B/perforin lytic pathway, is hyperexpressed during acute rejection of renal allografts. Transplantation 75, 1565–1570 (2003).
Ding, R. et al. CD103 mRNA levels in urinary cells predict acute rejection of renal allografts. Transplantation 75, 1307–1312 (2003).
Øzbay, A., Torring, C., Olsen, R. & Carstens, J. Transcriptional profiles in urine during acute rejection, bacteriuria, CMV infection and stable graft function after renal transplantation. Scand. J. Immunol. 69, 357–365 (2009).
Tatapudi, R. R. et al. Noninvasive detection of renal allograft inflammation by measurements of mRNA for IP-10 and CXCR3 in urine. Kidney Int. 65, 2390–2397 (2004).
Peng, W. et al. Urinary fractalkine is a marker of acute rejection. Kidney Int. 74, 1454–1460 (2008).
Seiler, M. et al. Heightened expression of the cytotoxicity receptor NKG2D correlates with acute and chronic nephropathy after kidney transplantation. Am. J. Transplant. 7, 423–433 (2007).
Renesto, P. G., Ponciano, V. C., Cenedeze, M. A., Saraiva Câmara, N. O. & Pacheco-Silva, A. High expression of Tim-3 mRNA in urinary cells from kidney transplant recipients with acute rejection. Am. J. Transplant. 7, 1661–1665 (2007).
Hartono, C., Muthukumar, T. & Suthanthiran, M. Noninvasive diagnosis of acute rejection of renal allografts. Curr. Opin. Organ Transplant. 15, 35–41 (2010).
Shin, G. T., Kim, S. J., Lee, T. S., Oh, C. K. & Kim, H. Gene expression of perforin by peripheral blood lymphocytes as a marker of acute rejection. Nephron Clin. Pract. 100, c63–c70 (2005).
Simon, T., Opelz, G., Wiesel, M., Ott, R. C. & Süsal, C. Serial peripheral blood perforin and granzyme B gene expression measurements for prediction of acute rejection in kidney graft recipients. Am. J. Transplant. 3, 1121–1127 (2003).
Simon, T. et al. Serial peripheral blood interleukin-18 and perforin gene expression measurements for prediction of acute kidney graft rejection. Transplantation 77, 1589–1595 (2004).
Alakulppi, N. S., Kyllönen, L. E., Partanen, J., Salmela, K. T. & Laine, J. T. Diagnosis of acute renal allograft rejection by analyzing whole blood mRNA expression of lymphocyte marker molecules. Transplantation 83, 791–798 (2007).
Aquino-Dias, E. C. et al. Non-invasive diagnosis of acute rejection in kidney transplants with delayed graft function. Kidney Int. 73, 877–884 (2008).
Solez, K. et al. Banff 07 classification of renal allograft pathology: updates and future directions. Am. J. Transplant. 8, 753–760 (2008).
Hoffmann, S. C. et al. Functionally significant renal allograft rejection is defined by transcriptional criteria. Am. J. Transplant. 5, 573–581 (2005).
Sarwal, M. et al. Molecular heterogeneity in acute renal allograft rejection identified by DNA microarray profiling. N. Engl. J. Med. 349, 125–138 (2003).
Mueller, T. F. et al. Microarray analysis of rejection in human kidney transplants using pathogenesis-based transcript sets. Am. J. Transplant. 7, 2712–2722 (2007).
Famulski, K. S. et al. Defining the canonical form of T-cell-mediated rejection in human kidney transplants. Am. J. Transplant. 10, 810–820 (2010).
Einecke, G. et al. A molecular classifier for predicting future graft loss in late kidney transplant biopsies. J. Clin. Invest. 120, 1862–1872 (2010).
Anglicheau, D. et al. MicroRNA expression profiles predictive of human renal allograft status. Proc. Natl Acad. Sci. USA 106, 5330–5335 (2009).
King's College London Nephrology & Transplantation Genetic Analysis and Monitoring of Biomarkers of Immunological Tolerance [online], (2010).
Näther, B. J. et al. Modified ELISPOT technique--highly significant inverse correlation of post-Tx donor-reactive IFNgamma-producing cell frequencies with 6 and 12 months graft function in kidney transplant recipients. Transpl. Immunol. 16, 232–237 (2006).
Gagliani, N. et al. Antigen-specific dependence of Tr1-cell therapy in preclinical models of islet transplant. Diabetes 59, 433–439 (2010).
Issa, F., Schiopu, A. & Wood, K. J. Role of T cells in graft rejection and transplantation tolerance. Expert Rev. Clin. Immunol. 6, 155–169 (2010).
Kawai, T. et al. HLA-mismatched renal transplantation without maintenance immunosuppression. N. Engl. J. Med. 358, 353–361 (2008).
Acknowledgements
M. P. Hernandez-Fuentes and R. I. Lechler acknowledge financial support from the EU (QLRT–2002–02127, from FP5), Immune Tolerance Network (ITN503ST), RISET consortium (512090 IP, from FP6), MRC (G0801537/ID: 88245) and Guy's & St Thomas' Charity (Grant 080530). M. P. Hernandez-Fuentes acknowledges financial support from the Department of Health via the NIH Research (NIHR) comprehensive Biomedical Research Center award to Guy's & St Thomas' NHS Foundation Trust in partnership with King's College London and King's College Hospital NHS Foundation Trust.
Author information
Authors and Affiliations
Contributions
M. P. Hernandez-Fuentes and R. I. Lechler substantially contributed to the discussion of content and reviewed/edited the manuscript before submission. M. P. Hernandez-Fuentes also researched data for the article and wrote the article.
Corresponding author
Ethics declarations
Competing interests
M. P. Hernandez-Fuentes and R. I. Lechler have a patent application with King's College London.
Rights and permissions
About this article
Cite this article
Hernandez-Fuentes, M., Lechler, R. A 'biomarker signature' for tolerance in transplantation. Nat Rev Nephrol 6, 606–613 (2010). https://doi.org/10.1038/nrneph.2010.112
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneph.2010.112
This article is cited by
-
Tolerance: an overview and perspectives
Nature Reviews Nephrology (2010)