Key Points
-
This Review discusses natural bacterial transformation, highlighting the common and divergent features that exist in a phylogenetically diverse range of naturally transformable species.
-
Transformation is defined as the uptake of foreign DNA as single strands and its subsequent integration into the bacterial chromosome by homologous recombination. The mechanisms of uptake and integration, which are largely conserved among species, are highlighted and their conservation is explored.
-
In contrast to DNA-uptake mechanisms, the regulation of the ability to transform (which is known as competence) and the signals that induce competence vary widely between species; the range of mechanisms that are involved are discussed.
-
The roles of competence and imported DNA are also considered, and we argue that evidence so far generally points towards a role for transformation in the generation of genetic diversity or in chromosomal repair, rather than a nutritional role.
-
Finally, we explore the future prospects in this field of research, detailing several case studies of species that have recently been shown to be transformable and the potential difficulties in demonstrating transformability in a new species.
Abstract
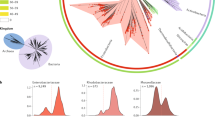
Natural bacterial transformation involves the internalization and chromosomal integration of DNA and has now been documented in ∼80 species. Recent advances have established that phylogenetically distant species share conserved uptake and processing proteins but differ in the inducing cues and regulatory mechanisms that are involved. In this Review, we highlight divergent and common principles that govern the transformation process in different bacteria. We discuss how this cumulative knowledge enables the prediction of new transformable species and supports the idea that the main role of internalized DNA is in the generation of genetic diversity or in chromosome repair rather than in nutrition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Griffith, F. The significance of pneumococcal types. J. Hyg. 27, 113–159 (1928).
Croucher, N. J. et al. Rapid pneumococcal evolution in response to clinical interventions. Science 331, 430–434 (2011).
Lorenz, M. G. & Wackernagel, W. Bacterial gene transfer by natural genetic transformation in the environment. Microbiol. Rev. 58, 563–602 (1994).
Johnsborg, O., Eldholm, V. & Håvarstein, L. S. Natural genetic transformation: prevalence, mechanisms and function. Res. Microbiol. 158, 767–778 (2007).
Averhoff, B. & Friedrich, A. Type IV pili-related natural transformation systems: DNA transport in mesophilic and thermophilic bacteria. Arch. Microbiol. 180, 385–393 (2003).
Chen, I. & Dubnau, D. DNA uptake during bacterial transformation. Nature Rev. Microbiol. 2, 241–249 (2004).
Allemand, J. F. & Maier, B. Bacterial translocation motors investigated by single molecule techniques. FEMS Microbiol. Rev. 33, 593–610 (2009).
Claverys, J. P., Martin, B. & Polard, P. The genetic transformation machinery: composition, localization and mechanism. FEMS Microbiol. Rev. 33, 643–656 (2009).
Allemand, J. F., Maier, B. & Smith, D. E. Molecular motors for DNA translocation in prokaryotes. Curr. Opin. Biotechnol. 23, 503–509 (2012).
Claverys, J. P., Prudhomme, M. & Martin, B. Induction of competence regulons as general stress responses in Gram-positive bacteria. Annu. Rev. Microbiol. 60, 451–475 (2006).
Seitz, P. & Blokesch, M. Cues and regulatory pathways involved in natural competence and transformation in pathogenic and environmental Gram-negative bacteria. FEMS Microbiol. Rev. 37, 336–363 (2012).
Draskovic, I. & Dubnau, D. Biogenesis of a putative channel protein, ComEC, required for DNA uptake: membrane topology, oligomerization and formation of disulphide bonds. Mol. Microbiol. 55, 881–896 (2005).
Mortier-Barrière, I. et al. A key presynaptic role in transformation for a widespread bacterial protein: DprA conveys incoming ssDNA to RecA. Cell 130, 824–836 (2007). This study identifies DprA as a transformation-dedicated loader of RecA onto transforming ssDNA, which is a step that is crucial for the formation of transformation recombinants.
Hobbs, M. & Mattick, J. S. Common components in the assembly of type 4 fimbriae, DNA transfer systems, filamentous phage and protein-secretion apparatus: a general system for the formation of surface-associated protein complexes. Mol. Microbiol. 10, 233–243 (1993).
Dubnau, D. DNA uptake in bacteria. Annu. Rev. Microbiol. 53, 217–244 (1999).
Mann, J. M., Carabetta, V. J., Cristea, I. M. & Dubnau, D. Complex formation and processing of the minor transformation pilins of Bacillus subtilis. Mol. Microbiol. 90, 1201–1215 (2013).
Chen, I., Provvedi, R. & Dubnau, D. A macromolecular complex formed by a pilin-like protein in competent Bacillus subtilis. J. Biol. Chem. 281, 21720–21727 (2006). This paper documents the presence of a short pilus-like structure in B. subtilis . This is the first identification of a competence-specific pseudopilus in Gram-positive bacteria.
Laurenceau, R. et al. A Type IV pilus mediates DNA binding during natural transformation in Streptococcus pneumoniae. PLoS Pathog. 9, e1003473 (2013). By identifying a long pilus that is essential for transformation in S. pneumoniae , this study shows that transformation pili can protrude far beyond the cell wall of Gram-positive bacteria to capture exogenous DNA.
Provvedi, R. & Dubnau, D. ComEA is a DNA receptor for transformation of competent Bacillus subtilis. Mol. Microbiol. 31, 271–280 (1999).
Burrows, L. L. Weapons of mass retraction. Mol. Microbiol. 57, 878–888 (2005).
Puyet, A., Greenberg, B. & Lacks, S. A. Genetic and structural characterization of EndA. A membrane-bound nuclease required for transformation of Streptoccus pneumoniae. J. Mol. Biol. 213, 727–738 (1990).
Bergé, M. et al. Uptake of transforming DNA in Gram-positive bacteria: a view from Streptococcus pneumoniae. Mol. Microbiol. 45, 411–421 (2002).
Seitz, P. & Blokesch, M. DNA-uptake machinery of naturally competent Vibrio cholerae. Proc. Natl Acad. Sci. USA 110, 17987–17992 (2013).
Bergé, M. J. et al. Midcell recruitment of the DNA uptake and virulence nuclease, EndA, for pneumococcal transformation. PLoS Pathog. 9, e1003596 (2013). This study shows that the EndA nuclease is recruited to midcell during pneumococcal competence, which indicates that DNA uptake is likely to occur at this site.
Hahn, J. et al. Transformation proteins and DNA uptake localize to the cell poles in Bacillus subtilis. Cell 122, 59–71 (2005). This study shows that DNA uptake occurs at the cell poles in competent B. subtilis.
Londoño-Vallejo, J. A. & Dubnau, D. Mutation of the putative nucleotide binding site of the Bacillus subtilis membrane protein ComFA abolishes the uptake of DNA during transformation. J. Bacteriol. 176, 4642–4645 (1994).
Yeh, Y. C., Lin, T. L., Chang, K. C. & Wang, J. T. Characterization of a ComE3 homologue essential for DNA transformation in Helicobacter pylori. Infect. Immun. 71, 5427–5431 (2003).
Stingl, K. et al. Composite system mediates two-step DNA uptake into Helicobacter pylori. Proc. Natl Acad. Sci. USA 107, 1184–1189 (2010).
Beernink, H. T. & Morrical, S. W. RMPs: recombination/replication mediator proteins. Trends Biochem. Sci. 24, 385–389 (1999).
Quevillon-Cheruel, S. et al. Structure–function analysis of pneumococcal DprA protein reveals that dimerization is crucial for loading RecA recombinase onto DNA during transformation. Proc. Natl Acad. Sci. USA 109, E2466–E2475 (2012).
Prudhomme, M. et al. Antibiotic stress induces genetic transformability in the human pathogen Streptococcus pneumoniae. Science 313, 89–92 (2006). This paper reports the discovery that pneumococcal competence is induced in response to environmental stresses such as antibiotics and DNA-damaging agents.
Charpentier, X., Polard, P. & Claverys, J. P. Induction of competence for genetic transformation by antibiotics: convergent evolution of stress responses in distant bacterial species lacking SOS? Curr. Opin. Microbiol. 15, 1–7 (2012).
Butala, M., Zgur-Bertok, D. & Busby, S. J. The bacterial LexA transcriptional repressor. Cell. Mol. Life Sci. 66, 82–93 (2009).
Lee, M. S. & Morrison, D. A. Identification of a new regulator in Streptococcus pneumoniae linking quorum sensing to competence for genetic transformation. J. Bacteriol. 181, 5004–5016 (1999).
Peterson, S. et al. Identification of competence pheromone responsive genes in Streptococcus pneumoniae. Mol. Microbiol. 51, 1051–1070 (2004).
Martin, B., Quentin, Y., Fichant, G. & Claverys, J. P. Independent evolution of competence regulatory cascades in streptococci? Trends Microbiol. 14, 339–345 (2006). This paper reports the phylogenetic analysis of the streptococcal ComDE TCS and establishes that S. mutans and several other streptococci lack this master competence regulator, which suggests that there is an alternative competence regulatory circuit in these species.
Fontaine, L. et al. A novel pheromone quorum-sensing system controls the development of natural competence in Streptococcus thermophilus and Streptococcus salivarius. J. Bacteriol. 192, 1444–1454 (2010).
Mashburn-Warren, L., Morrison, D. A. & Federle, M. J. A novel double-tryptophan peptide pheromone controls competence in Streptococcus spp. via an Rgg regulator. Mol. Microbiol. 78, 589–606 (2010).
Morrison, D. A., Guedon, E. & Renault, P. Competence for natural genetic transformation in the Streptococcus bovis Group Streptococci S. infantarius and S. macedonicus. J. Bacteriol. 195, 2612–2620 (2013).
Morikawa, K. et al. Expression of a cryptic secondary sigma factor gene unveils natural competence for DNA transformation in Staphylococcus aureus. PLoS Pathog. 8, e1003003 (2012).
Mohan, S. & Dubnau, D. Transcriptional regulation of comC: evidence for a competence-specific transcription factor in Bacillus subtilis. J. Bacteriol. 172, 4064–4071 (1990).
Maamar, H. & Dubnau, D. Bistability in the Bacillus subtilis K-state (competence) system requires a positive feedback loop. Mol. Microbiol. 56, 615–624 (2005). This paper shows that stochastic variation in comK expression in B. subtilis populations determines which cells become competent.
Smits, W. K. et al. Stripping Bacillus: ComK auto-stimulation is responsible for the bistable response in competence development. Mol. Microbiol. 56, 604–614 (2005).
Dubnau, D. & Losick, R. Bistability in bacteria. Mol. Microbiol. 61, 564–572 (2006).
Smits, W. K., Kuipers, O. P. & Veening, J. W. Phenotypic variation in bacteria: the role of feedback regulation. Nature Rev. Microbiol. 4, 259–271 (2006).
Maamar, H., Raj, A. & Dubnau, D. Noise in gene expression determines cell fate in Bacillus subtilis. Science 317, 526–529 (2007).
Leisner, M., Stingl, K., Frey, E. & Maier, B. Stochastic switching to competence. Curr. Opin. Microbiol. 11, 553–559 (2008).
Berka, R. M. et al. Microarray analysis of the Bacillus subtilis K-state: genome-wide expression changes dependent on ComK. Mol. Microbiol. 43, 1331–1345 (2002).
Hamoen, L. W. et al. Improving the predictive value of the competence transcription factor (ComK) binding site in Bacillus subtilis using a genomic approach. Nucl. Acids Res. 202, 5517–5528 (2002).
Ogura, M. et al. Whole-genome analysis of genes regulated by the Bacillus subtilis competence transcription factor ComK. J. Bacteriol. 184, 2344–2351 (2002).
Kuzniar, A., van Ham, R. C., Pongor, S. & Leunissen, J. A. The quest for orthologs: finding the corresponding gene across genomes. Trends Genet. 24, 539–551 (2008).
Wise, E. M. Jr., Alexander, S. P. & Powers, M. Adenosine 3′:5′-cyclic monophosphate as a regulator of bacterial transformation. Proc. Natl Acad. Sci. USA 70, 471–474 (1973).
Dorocicz, I. E., Williams, P. M. & Redfield, R. J. The Haemophilus influenzae adenylate cyclase gene: cloning, sequence, and essential role in competence. J. Bacteriol. 175, 7142–7149 (1993).
Chandler, M. S. The gene encoding cAMP receptor protein is required for competence development in Haemophilus influenzae Rd. Proc. Natl Acad. Sci. USA 89, 1626–1630 (1992).
Redfield, R. J. sxy-1, a Haemophilus influenzae mutation causing greatly enhanced spontaneous competence. J. Bacteriol. 173, 5612–5618 (1991).
Macfadyen, L. P. Regulation of competence development in Haemophilus influenzae. J. Theor. Biol. 207, 349–359 (2000).
Redfield, R. J. et al. A novel CRP-dependent regulon controls expression of competence genes in Haemophilus influenzae. J. Mol. Biol. 347, 735–747 (2005).
Meibom, K. L. et al. Chitin induces natural competence in Vibrio cholerae. Science 310, 1824–1827 (2005). This paper reports the discovery that competence in V. cholerae is induced by chitin, which is a sugar polymer that is abundant in the aquatic habitat of this bacterium.
Blokesch, M. A quorum sensing-mediated switch contributes to natural transformation of Vibrio cholerae. Mob. Genet. Elements 2, 224–227 (2012).
Lo Scrudato, M. & Blokesch, M. A transcriptional regulator linking quorum sensing and chitin induction to render Vibrio cholerae naturally transformable. Nucleic Acids Res. 41, 3644–3658 (2013). This paper provides the first documentation of bipartite regulation of competence genes, in which they are directly controlled by two distinct regulators.
Hamoen, L. W., Venema, G. & Kuipers, O. P. Controlling competence in Bacillus subtilis: shared use of regulators. Microbiology 149, 9–17 (2003).
Hamoen, L. W., Van Werkhoven, A. F., Dubnau, D. & Venema, G. The competence transcription factor of Bacillus subtilis recognizes short A/T-rich sequences arranged in a unique, flexible pattern along the DNA helix. Genes Dev. 12, 1539–1550 (1998).
Hamoen, L. W., Van Werkhoven, A. F., Venema, G. & Dubnau, D. The pleiotropic response regulator DegU functions as a priming protein in competence development in Bacillus subtilis. Proc. Natl Acad. Sci. USA 97, 9246–9251 (2000).
Hamoen, L. W. et al. The Bacillus subtilis transition state regulator AbrB binds to the -35 promoter region of comK. FEMS Microbiol. Lett. 218, 299–304 (2003).
Serror, P. & Sonenshein, A. L. CodY is required for nutritional repression of Bacillus subtilis genetic competence. J. Bacteriol. 178, 5910–5915 (1996).
Hoa, T. T., Tortosa, P., Albano, M. & Dubnau, D. Rok (YkuW) regulates genetic competence in Bacillus subtilis by directly repressing comK. Mol. Microbiol. 43, 15–26 (2002).
Turgay, K., Hamoen, L. W., Venema, G. & Dubnau, D. Biochemical characterization of a molecular switch involving the heat shock protein ClpC, which controls the activity of ComK, the competence transcription factor of Bacillus subtilis. Genes Dev. 11, 119–128 (1997). This paper provides in vitro characterization of the interplay between ComK, ClpCP, MecA and ComS and demonstrated that accumulation of B. subtilis ComK is regulated by proteolysis.
Turgay, K., Hahn, J., Burghoorn, J. & Dubnau, D. Competence in Bacillus subtilis is controlled by regulated proteolysis of a transcription factor. EMBO J. 17, 6730–6738 (1998).
Nakano, M. M. et al. srfA is an operon required for surfactin production, competence development, and efficient sporulation in Bacillus subtilis. J. Bacteriol. 173, 1770–1778 (1991).
Roggiani, M. & Dubnau, D. ComA, a phosphorylated response regulator protein of Bacillus subtilis, binds to the promoter region of srfA. J. Bacteriol. 175, 3182–3187 (1993).
Hayashi, K. et al. The H2O2 stress-responsive regulator PerR positively regulates srfA expression in Bacillus subtilis. J. Bacteriol. 187, 6659–6667 (2005).
Hui, F. M. & Morrison, D. A. Genetic transformation in Streptococcus pneumoniae: nucleotide sequence analysis shows comA, a gene required for competence induction, to be a member of the bacterial ATP-dependent transport protein family. J. Bacteriol. 173, 372–381 (1991).
Pestova, E. V., Håvarstein, L. S. & Morrison, D. A. Regulation of competence for genetic transformation in Streptococcus pneumoniae by an auto-induced peptide pheromone and a two-component regulatory system. Mol. Microbiol. 21, 853–864 (1996).
Håvarstein, L. S., Coomaraswamy, G. & Morrison, D. A. An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae. Proc. Natl Acad. Sci. USA 92, 11140–11144 (1995).
Martin, B. et al. ComE/ComE∼P interplay dictates activation or extinction status of pneumococcal X-state (competence). Mol. Microbiol. 87, 394–411 (2012).
Gardan, R. et al. The oligopeptide transport system is essential for the development of natural competence in Streptococcus thermophilus strain LMD-9. J. Bacteriol. 191, 4647–4655 (2009). This paper reports the seminal discovery of the involvement of an oligopeptide transporter in a competence regulatory cascade, which paved the way for the discovery of ComRS in S. thermophilus.
Gardan, R. et al. Extracellular life cycle of ComS, the competence-stimulating peptide of Streptococcus thermophilus. J. Bacteriol. 195, 1845–1855 (2013).
Fontaine, L. et al. Mechanism of competence activation by the ComRS signalling system in streptococci. Mol. Microbiol. 87, 1113–1132 (2013).
Sulavik, M. C., Tardif, G. & Clewell, D. B. Identification of a gene, rgg, which regulates expression of glucosyltransferase and influences the Spp phenotype of Streptococcus gordonii Challis. J. Bacteriol. 174, 3577–3586 (1992).
Fleuchot, B. et al. Rgg proteins associated with internalized small hydrophobic peptides: a new quorum-sensing mechanism in streptococci. Mol. Microbiol. 80, 1102–1119 (2011).
de Saizieu, A. et al. Microarray-based identification of a novel Streptococcus pneumoniae regulon controlled by an autoinduced peptide. J. Bacteriol. 182, 4696–4703 (2000).
Federle, M. J. & Morrison, D. A. One if by land, two if by sea: signalling to the ranks with CSP and XIP. Mol. Microbiol. 86, 241–245 (2012).
Lemme, A. et al. Subpopulation-specific transcriptome analysis of competence-stimulating-peptide-induced Streptococcus mutans. J. Bacteriol. 193, 1863–1877 (2011).
Son, M. et al. Microfluidic study of competence regulation in Streptococcus mutans: environmental inputs modulate bimodal and unimodal expression of comX. Mol. Microbiol. 86, 258–272 (2012).
Mirouze, N. et al. Direct involvement of DprA, the transformation-dedicated RecA loader, in the shut-off of pneumococcal competence. Proc. Natl Acad. Sci. USA 110, E1035–E1044 (2013).
Yamamoto, S., Morita, M., Izumiya, H. & Watanabe, H. Chitin disaccharide (GlcNAc)2 induces natural competence in Vibrio cholerae through transcriptional and translational activation of a positive regulatory gene tfoXVC. Gene 457, 42–49 (2010).
Yamamoto, S. et al. Regulation of natural competence by the orphan two-component system sensor kinase ChiS involves a non-canonical transmembrane regulator in Vibrio cholerae. Mol. Microbiol. 91, 326–247 (2013).
Yamamoto, S. et al. Identification of a chitin-induced small RNA that regulates translation of the tfoX gene, encoding a positive regulator of natural competence in Vibrio cholerae. J. Bacteriol. 193, 1953–1965 (2011).
Herriott, R. M., Meyer, E. M. & Vogt, M. Defined nongrowth media for stage II development of competence in Haemophilus influenzae. J. Bacteriol. 101, 517–524 (1970).
Bosse, J. T. et al. Natural competence in strains of Actinobacillus pleuropneumoniae. FEMS Microbiol. Lett. 298, 124–130 (2009).
Sinha, S., Mell, J. & Redfield, R. The availability of purine nucleotides regulates natural competence by controlling translation of the competence activator Sxy. Mol. Microbiol. 6, 1106–1119 (2013).
Cameron, A. D., Volar, M., Bannister, L. A. & Redfield, R. J. RNA secondary structure regulates the translation of sxy and competence development in Haemophilus influenzae. Nucleic Acids Res. 36, 10–20 (2008).
Boutry, C. et al. Adpaptor protein MecA is a negative regulator of the expression of late competence genes in Streptococcus thermophilus. J. Bacteriol. 194, 1777–1788 (2012).
Biornstad, T. J. & Havarstein, L. S. ClpC acts as a negative regulator of competence in Streptococcus thermophilus. Microbiology 157, 1676–1684 (2011).
Sung, C. K. & Morrison, D. A. Two distinct functions of ComW in stabilization and activation of the alternative sigma factor ComX in Streptococcus pneumoniae. J. Bacteriol. 187, 3052–3061 (2005).
Nielsen, K. M., Bones, A. M. & van Elsas, J. D. Induced natural transformation of Acinetobacter calcoaceticus in soil microcosms. Appl. Environ. Microbiol. 63, 3972–3977 (1997).
Lo Scrudato, M. & Blokesch, M. The regulatory network of natural competence and transformation of Vibrio cholerae. PLoS Genet. 8, e1002778 (2012).
Keyhani, N. O. & Roseman, S. Physiological aspects of chitin catabolism in marine bacteria. Biochim. Biophys. Acta 1473, 108–122 (1999).
Murphy, T. F. & Brauer, A. L. Expression of urease by Haemophilus influenzae during human respiratory tract infection and role in survival in an acid environment. BMC. Microbiol. 11, 183 (2011).
Charpentier, X., Kay, E., Schneider, D. & Shuman, H. A. Antibiotics and UV radiation induce competence for natural transformation in Legionella pneumophila. J. Bacteriol. 193, 1114–1121 (2011).
Boutry, C. et al. SOS response activation and competence development are antagonistic mechanisms in Streptococcus thermophilus. J. Bacteriol. 195, 696–707 (2013).
Ratnayake-Lecamwasam, M., Serror, P., Wong, K. W. & Sonenshein, A. L. Bacillus subtilis CodY represses early-stationary-phase genes by sensing GTP levels. Genes Dev. 15, 1093–1103 (2001).
Antonova, E. S., Bernardy, E. E. & Hammer, B. K. Natural competence in Vibrio cholerae is controlled by a nucleoside scavenging response that requires CytR-dependent anti-activation. Mol. Microbiol. 86, 1215–1231 (2012).
Michod, R. E., Bernstein, H. & Nedelcu, A. M. Adaptive value of sex in microbial pathogens. Infect. Genet. Evol. 8, 267–285 (2008).
Vos, M. Why do bacteria engage in homologous recombination? Trends Microbiol. 17, 226–232 (2009).
Redfield, R. J. Genes for breakfast: the have-your-cake-and-eat-it-too of bacterial transformation. J. Hered. 84, 400–404 (1993).
Redfield, R. J. Do bacteria have sex? Nature Rev. Genet. 2, 634–639 (2001).
Bergé, M., Mortier-Barrière, I., Martin, B. & Claverys, J. P. Transformation of Streptococcus pneumoniae relies on DprA- and RecA-dependent protection of incoming single strands. Mol. Microbiol. 50, 527–536 (2003).
Kidane, D. et al. Evidence for different pathways during horizontal gene transfer in competent Bacillus subtilis cells. PLoS. Genet. 5, e1000630 (2009).
Morrison, D. A., Mortier-Barrière, I., Attaiech, L. & Claverys, J. P. Identification of the major protein component of the pneumococcal eclipse complex. J. Bacteriol. 189, 6497–6500 (2007).
Attaiech, L. et al. Role of the single-stranded DNA binding protein SsbB in pneumococcal transformation: maintenance of a reservoir for genetic plasticity. PLoS Genet. 7, e1002156 (2011).
Sinha, S., Mell, J. C. & Redfield, R. J. Seventeen Sxy-dependent cyclic AMP receptor protein site-regulated genes are needed for natural transformation in Haemophilus influenzae. J. Bacteriol. 194, 5245–5254 (2012).
Mortier-Barrière, I., de Saizieu, A., Claverys, J. P. & Martin, B. Competence-specific induction of recA is required for full recombination proficiency during transformation in Streptococcus pneumoniae. Mol. Microbiol. 27, 159–170 (1998).
Lacks, S. A., Ayalew, S., de la Campa, A. G. & Greenberg, B. Regulation of competence for genetic transformation in Streptococcus pneumoniae: expression of dpnA, a late competence gene encoding a DNA methyltransferase of the Dpn II restriction system. Mol. Microbiol. 35, 1089–1098 (2000).
Johnston, C. et al. Programmed protection of foreign DNA from restriction allows pathogenicity island exchange during pneumococcal transformation. PLoS Pathog. 9, e1003178 (2013). This study shows that a competence-induced pneumococcal methylase is crucial for protecting transformant chromosomes from restriction.
Cerritelli, S., Springhorn, S. S. & Lacks, S. A. DpnA, a methylase for single-strand DNA in the Dpn II restriction system, and its biological function. Proc. Natl Acad. Sci. USA 86, 9223–9227 (1989).
Johnston, C., Martin, B., Polard, P. & Claverys, J. P. Post-replication targeting of transformants by bacterial immune systems? Trends Microbiol. 21, 516–521 (2013).
Johnston, C., Polard, P. & Claverys, J. P. The DpnI/DpnII pneumococcal system, defence against foreign attack without compromising genetic exchange. Mob. Genet. Elements 3, e25582 (2013).
Johnston, C. et al. Natural genetic transformation generates a population of merodiploids in Streptococcus pneumoniae. PLoS Genet. 9, e1003819 (2013).
Rabinovich, L. et al. Prophage excision activates Listeria competence genes that promote phagosomal escape and virulence. Cell 150, 792–802 (2012).
Claverys, J. P. & Martin, B. Bacterial “competence” genes: signatures of active transformation, or only remnants? Trends Microbiol. 11, 161–165 (2003).
Hofreuter, D., Odenbreit, S. & Haas, R. Natural transformation competence in Helicobacter pylori is mediated by the basic components of a type IV secretion system. Mol. Microbiol. 41, 379–391 (2001).
Karnholz, A. et al. Functional and topological characterization of novel components of the comB DNA transformation competence system in Helicobacter pylori. J. Bacteriol. 188, 882–893 (2006).
Dorer, M. S., Fero, J. & Salama, N. R. DNA damage triggers genetic exchange in Helicobacter pylori. PLoS Pathog. 6, e1001026 (2010).
Baltrus, D. A. & Guillemin, K. Multiple phases of competence occur during the Helicobacter pylori growth cycle. FEMS Microbiol. Lett. 255, 148–155 (2006).
Higgins, D. A. et al. The major Vibrio cholerae autoinducer and its role in virulence factor production. Nature 450, 883–886 (2007).
Chen, X. et al. Structural identification of a bacterial quorum-sensing signal containing boron. Nature 415, 545–549 (2002).
Stephenson, S., Mueller, C., Jiang, M. & Perego, M. Molecular analysis of Phr peptide processing in Bacillus subtilis. J. Bacteriol. 185, 4861–4871 (2003).
Lanigan-Gerdes, S., Dooley, A. N., Faull, K. F. & Lazazzera, B. A. Identification of subtilisin, Epr and Vpr as enzymes that produce CSF, an extracellular signalling peptide of Bacillus subtilis. Mol. Microbiol. 65, 1321–1333 (2007).
Lanigan-Gerdes, S. et al. Identification of residues important for cleavage of the extracellular signaling peptide CSF of Bacillus subtilis from its precursor protein. J. Bacteriol. 190, 6668–6675 (2008).
Henke, J. M. & Bassler, B. L. Three parallel quorum-sensing systems regulate gene expression in Vibrio harveyi. J. Bacteriol. 186, 6902–6914 (2004).
Neiditch, M. B. et al. Regulation of LuxPQ receptor activity by the quorum-sensing signal autoinducer-2. Mol. Cell 18, 507–518 (2005).
Ng, W. L. & Bassler, B. L. Bacterial quorum-sensing network architectures. Annu. Rev. Genet. 43, 197–222 (2009).
Lenz, D. H. et al. The small RNA chaperone Hfq and multiple small RNAs control quorum sensing in Vibrio harveyi and Vibrio cholerae. Cell 118, 69–82 (2004).
Halfmann, A., Kovacs, M., Hakenbeck, R. & Bruckner, R. Identification of the genes directly controlled by the response regulator CiaR in Streptococcus pneumoniae: five out of 15 promoters drive expression of small non-coding RNAs. Mol. Microbiol. 66, 110–126 (2007).
Schnorpfeil, A. et al. Target evaluation of the non-coding csRNAs reveals a link of the two-component regulatory system CiaRH to competence control in Streptococcus pneumoniae R6. Mol. Microbiol. 2, 334–349 (2013).
Stevens, K. E., Chang, D., Zwack, E. E. & Sebert, M. E. Competence in Streptococcus pneumoniae is regulated by the rate of ribosomal decoding errors. mBio 2 e00071–11 (2011).
Cassone, M. et al. The HtrA protease from Streptococcus pneumoniae digests both denatured proteins and the competence-stimulating peptide. J. Biol. Chem. 287, 38449–38459 (2012).
Foster, P. L. Stress responses and genetic variation in bacteria. Mutat. Res. 569, 3–11 (2005).
Inamine, G. S. & Dubnau, D. ComEA, a Bacillus subtilis integral membrane protein required for genetic transformation, is needed for both DNA binding and transport. J. Bacteriol. 177, 3045–3051 (1995).
Barany, F., Kahn, M. E. & Smith, H. O. Directional transport and integration of donor DNA in Haemophilus influenzae. Proc. Natl Acad. Sci. USA 80, 7274–7278 (1983).
Méjean, V. & Claverys, J. P. Use of a cloned fragment to analyze the fate of donor DNA in transformation of Streptococcus pneumoniae. J. Bacteriol. 158, 1175–1178 (1984).
Mell, J. C., Hall, I. M. & Redfield, R. J. Defining the DNA uptake specificity of naturally competent Haemophilus influenzae cells. Nucleic Acids Res. 40, 8536–8549 (2012).
Claverys, J. P. & Håvarstein, L. S. Cannibalism and fratricide: mechanisms and raisons d'être. Nature Rev. Microbiol. 5, 219–229 (2007).
Claverys, J. P., Martin, B. & Håvarstein, L. S. Competence-induced fratricide in streptococci. Mol. Microbiol. 64, 1423–1433 (2007).
Johnsborg, O., Eldholm, V., Bjornstad, M. L. & Håvarstein, L. S. A predatory mechanism dramatically increases the efficiency of lateral gene transfer in Streptococcus pneumoniae and related commensal species. Mol. Microbiol. 69, 245–253 (2008).
Aas, F. E., Lovold, C. & Koomey, M. An inhibitor of DNA binding and uptake events dictates the proficiency of genetic transformation in Neisseria gonorrhoeae: mechanism of action and links to type IV pilus expression. Mol. Microbiol. 46, 1441–1450 (2002).
Redfield, R. J. et al. Evolution of competence and DNA uptake specificity in the Pasteurellaceae. BMC Evol. Biol. 6, 82 (2006).
Frye, S. A., Nilsen, M., Tonjum, T. & Ambur, O. H. Dialects of the DNA uptake sequence in Neisseriaceae. PLoS Genet. 9, e1003458 (2013).
Blokesch, M. & Schoolnik, G. K. The extracellular nuclease Dns and its role in natural transformation of Vibrio cholerae. J. Bacteriol. 190, 7232–7240 (2008).
Finkel, S. E. & Kolter, R. DNA as a nutrient: novel role for bacterial competence gene homologs. J. Bacteriol. 183, 6288–6293 (2001).
Palchevskiy, V. & Finkel, S. E. Escherichia coli competence gene homologs are essential for competitive fitness and the use of DNA as a nutrient. J. Bacteriol. 188, 3902–3910 (2006).
Sun, D. Etude de la transformation plasmidique naturelle d'Escherichia coli et des ses relations éventuelles avec la compétence programmée pour la transformation génétique et la compétence dite nutritionnelle. Thesis, Univ. P. Sabatier, Toulouse, France (2011).
Blomqvist, T., Steinmoen, H. & Håvarstein, L. S. Natural genetic transformation: a novel tool for efficient genetic engineering of the dairy bacterium Streptococcus thermophilus. Appl. Environ. Microbiol. 72, 6751–6756 (2006).
Morikawa, K. et al. A new staphylococcal sigma factor in the conserved gene cassette: functional significance and implication for the evolutionary processes. Genes Cells 8, 699–712 (2003).
Schmid, S., Bevilacqua, C. & Crutz-Le Coq, A. M. Alternative sigma factor σH activates competence gene expression in Lactobacillus sakei. BMC. Microbiol. 12, 32 (2012).
Opdyke, J. A., Scott, J. R. & Moran, C. P. Jr. A secondary RNA polymerase sigma factor from Streptococcus pyogenes. Mol. Microbiol. 42, 495–502 (2001).
Woodbury, R. L., Wang, X. & Moran, C. P. Jr. Sigma X induces competence gene expression in Streptococcus pyogenes. Res. Microbiol. 157, 851–856 (2006).
Mashburn-Warren, L., Morrison, D. A. & Federle, M. J. The cryptic competence pathway in Streptococcus pyogenes is controlled by a peptide pheromone. J. Bacteriol. 194, 4589–4600 (2012).
Thorne, C. B. & Stull, H. B. Factors affecting transformation of Bacillus licheniformis. J. Bacteriol. 91, 1012–1020 (1966).
Koumoutsi, A. et al. Structural and functional characterization of gene clusters directing nonribosomal synthesis of bioactive cyclic lipopeptides in Bacillus amyloliquefaciens strain FZB42. J. Bacteriol. 186, 1084–1096 (2004).
Kovacs, A. T., Smits, W. K., Mironczuk, A. M. & Kuipers, O. P. Ubiquitous late competence genes in Bacillus species indicate the presence of functional DNA uptake machineries. Environ. Microbiol. 11, 1911–1922 (2009).
Mironczuk, A. M., Kovacs, A. T. & Kuipers, O. P. Induction of natural competence in Bacillus cereus ATCC14579. Microb. Biotech. 1, 226–235 (2008).
Cameron, A. D. & Redfield, R. J. Non-canonical CRP sites control competence regulons in Escherichia coli and many other γ-proteobacteria. Nucleic Acids Res. 34, 6001–6014 (2006). This study identifies a subset of CRP-binding sites that depend on the Sxy competence-regulating cofactor for transcriptional activation, which led to the finding that sxy regulons are induced during competence in H. influenzae and Gammaproteobacteria.
Sinha, S., Cameron, A. D. & Redfield, R. J. Sxy induces a CRP-S regulon in Escherichia coli. J. Bacteriol. 191, 5180–5195 (2009).
Sinha, S. & Redfield, R. J. Natural DNA uptake by Escherichia coli. PLoS ONE. 7, e35620 (2012).
Shimodaira, H. & Hasegawa, M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol. Biol. Evol. 16, 1114–1116 (1999).
Acknowledgements
The authors thank Y. Quentin for his help with phylogenetic analysis and the construction of the species tree. The authors apologize to researchers whose work could not be specifically cited owing to space limitations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Supplementary information
Supplementary information S1 (table)
Naturally transformable bacterial species (XLSX 25 kb)
Glossary
- Parasexual
-
A term used to describe a form of reproduction in which genetic recombination occurs in the absence of meiosis, wherein one 'partner' is DNA.
- Homologous recombination
-
The exchange of DNA sequences between identical or similar molecules. In the case of transformation, this involves the host chromosome and internalized single-stranded DNA (ssDNA).
- Transduction
-
A mechanism of horizontal gene transfer, in which DNA is accidentally transferred to a new host by a bacteriophage vector.
- Conjugation
-
A mechanism of horizontal gene transfer by cell-to-cell contact that is primarily used for plasmid transfer but occasionally leads to chromosomal transfer.
- Regulons
-
Sets of genes that are under coordinated control by dedicated regulatory circuits.
- Transformation heteroduplex
-
A DNA duplex that is comprised of one strand of host chromosomal DNA and a complementary strand of internalized DNA.
- Pili
-
Filamentous extracellular appendages that are present in some bacteria and that participate in different processes. Transformation pili promote the capture of exogenous DNA for uptake.
- Alternative sigma factors
-
(alternative σ factors). Alternative RNA polymerase cofactors that direct the RNA polymerase to specific promoters.
- Transcription co-regulators
-
Proteins that lack DNA-binding activity but can interact with and stimulate the activity of a transcription activator, thus indirectly promoting transcription.
- SOS response
-
A global response to DNA damage that enables DNA repair in bacteria.
- Orthologue
-
A homologous gene that is derived by a speciation event from a single ancestral sequence. Orthologues typically carry out equivalent functions in closely related species.
- Two-component signal-transduction system
-
(TCS). A system that comprises a histidine kinase and a response regulator; TCSs enable bacteria to sense signals (including those in the extracellular environment) and to regulate genes accordingly.
- Bacteriocins
-
Small peptides that are produced by bacteria to inhibit the growth of other species (sometimes closely related species) to which the producer possesses an immunity mechanism.
- Non-coding small RNA
-
A functional RNA molecule that is not translated into a protein.
- CAI-1
-
(Cholerae autoinducer 1).The main quorum-sensing signalling molecule of the human pathogen Vibrio cholerae; it has been identified as (S)-3-hydroxytridecan-4-one.
- AI-2
-
(Autoinducer 2). An inter-genera signalling molecule that is involved in quorum-sensing; it has been identified as the furanyl borate diester (2S,4S)-2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran borate.
- CSF
-
(Competence and sporulation factor). A signalling molecule that contributes to quorum-sensing in Bacillus subtilis. CSF is a pentapeptide that is produced from Phr precursor peptide, which is re-imported by the oligopeptide permease Opp.
- Pherotypes
-
Streptococci that produce the same competence-stimulating peptide (CSP) belong to the same pherotype.
- Quorum-sensing
-
The regulation of gene expression in response to cell density; secreted inducing molecules are sensed and induction occurs only when a critical cell density is reached.
- Cell–cell signalling
-
A mechanism of cell–cell communication in which an inducing molecule is produced and can be sensed by neighbouring cells, resulting in coordinated gene expression. This mechanism is not necessarily dependent on cell density owing to the fact that inducer expression can be regulated by external signals.
- Alarmone
-
A signalling molecule that is produced by bacteria in response to stress, which stimulates the expression of proteins involved in cellular processes that counteract the stress.
- Autoinducers
-
Non-proteinaceous chemical signalling molecules that are involved in cell–cell signalling and quorum-sensing.
- Carbon catabolite repression
-
(CCR). A regulatory mechanism in which the regulation of phosphotransferase systems enables the sequential utilization of carbon sources.
- Mitomycin C
-
A DNA-damaging agent that crosslinks target DNA and is toxic to bacterial cells.
- Hydroxyurea
-
A synthetic compound that promotes the stalling of bacterial replication forks by depleting nucleotide pools.
- Fratricide
-
The ability of competent pneumococcal cells to promote lysis of non-competent neighbouring pneumococci and closely related streptococci, liberating DNA for transformation and virulence factors.
- Paralogue
-
One of a pair of homologous genes that are derived by a duplication event from a single sequence. Paralogous relationships occur both within and between genomes, and paralogues can evolve to have novel functions.
- Restriction–modification system
-
(R–M system). A bacterial immune system that protects cells from invading foreign DNA, such as that injected by bacteriophages. Most of these systems encode a restriction enzyme that cleaves specific sequences in unmethylated DNA and a methylase that methylates the host genome, thereby protecting it from restriction.
- K-state
-
An alternative for competence in ComK-possessing bacteria (Bacilli), representing induction of the ComK regulon.
- X-state
-
An alternative for competence in σX-possessing bacteria (Streptococci), representing induction of the σX regulon.
Rights and permissions
About this article
Cite this article
Johnston, C., Martin, B., Fichant, G. et al. Bacterial transformation: distribution, shared mechanisms and divergent control. Nat Rev Microbiol 12, 181–196 (2014). https://doi.org/10.1038/nrmicro3199
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3199
This article is cited by
-
The priority of yeast to select among various DNA options to repair genome breaks by homologous recombination
Molecular Biology Reports (2024)
-
Tumor-derived cell-free DNA and circulating tumor cells: partners or rivals in metastasis formation?
Clinical and Experimental Medicine (2024)
-
An updated review on how biochar may possess potential in soil ARGs control on aspects of source, fate and elimination
Biochar (2024)
-
Programming bacteria for multiplexed DNA detection
Nature Communications (2023)
-
Lysogenic bacteriophages encoding arsenic resistance determinants promote bacterial community adaptation to arsenic toxicity
The ISME Journal (2023)