Key Points
-
Non-coding bacterial regulatory RNAs are key regulators of metabolic, physiological and pathogenic processes, as well as components of bacterial adaptive immunity.
-
Particularly abundant in this group of RNAs are the cis-encoded antisense RNAs (asRNAs), which overlap and are complementary to their target mRNAs encoded on the opposite DNA strand of the same genomic locus.
-
Although little is known about the function of most asRNAs, studied examples demonstrate that asRNAs can affect the expression of their complementary genes at the levels of transcription, mRNA stability or translation.
-
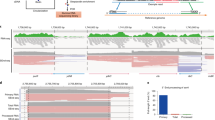
Recent transcriptome-wide studies in Listeria monocytogenes revealed a collection of unusually long asRNAs that extend over adjacent genes with divergent orientations; these genes also often encode proteins with opposing functions. The unique features of these loci have led to the definition of the excludon paradigm.
-
The long asRNA (lasRNA) in the excludon locus serves as both an antisense regulator that negatively regulates the complementary gene, and an mRNA for the adjacent gene, to positively contribute to its expression.
-
A novel concept in bacterial asRNA-mediated gene regulation has recently emerged from these observations: an excludon functions as a genomic toggle to ensure finely tuned regulation of genes encoded at the same genomic locus.
Abstract
In recent years, non-coding RNAs have emerged as key regulators of gene expression. Among these RNAs, the antisense RNAs (asRNAs) are particularly abundant, but in most cases the function and mechanism of action for a particular asRNA remains elusive. Here, we highlight a recently discovered paradigm termed the excludon, which defines a genomic locus encoding an unusually long asRNA that spans divergent genes or operons with related or opposing functions. Because these asRNAs can inhibit the expression of one operon while functioning as an mRNA for the adjacent operon, they act as fine-tuning regulatory switches in bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jacob, F. & Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 3, 318–356 (1961).
Waters, L. S. & Storz, G. Regulatory RNAs in bacteria. Cell 136, 615–628 (2009).
Sorek, R., Kunin, V. & Hugenholtz, P. CRISPR — a widespread system that provides acquired resistance against phages in bacteria and archaea. Nature Rev. Microbiol. 6, 181–186 (2008).
Arraiano, C. M. et al. The critical role of RNA processing and degradation in the control of gene expression. FEMS Microbiol. Rev. 34, 883–923 (2010).
Breaker, R. R. Prospects for riboswitch discovery and analysis. Mol. Cell 43, 867–879 (2011).
Kortmann, J. & Narberhaus, F. Bacterial RNA thermometers: molecular zippers and switches. Nature Rev. Microbiol. 10, 255–265 (2012).
Storz, G., Vogel, J. & Wassarman, K. M. Regulation by small RNAs in bacteria: expanding frontiers. Mol. Cell 43, 880–891 (2011).
Georg, J. & Hess, W. R. cis-antisense RNA, another level of gene regulation in bacteria. Microbiol. Mol. Biol. Rev. 75, 286–300 (2011).
Thomason, M. K. & Storz, G. Bacterial antisense RNAs: how many are there, and what are they doing? Annu. Rev. Genet. 44, 167–188 (2010).
Toledo-Arana, A. et al. The Listeria transcriptional landscape from saprophytism to virulence. Nature 459, 950–956 (2009).
Wurtzel, O. et al. Comparative transcriptomics of pathogenic and non-pathogenic Listeria species. Mol. Syst. Biol. 8, 583 (2012).
Stougaard, P., Molin, S. & Nordström, K. RNAs involved in copy-number control and incompatibility of plasmid R1. Proc. Natl Acad. Sci. USA 78, 6008–6012 (1981).
Tomizawa, J., Itoh, T., Selzer, G. & Som, T. Inhibition of ColE1 RNA primer formation by a plasmid-specified small RNA. Proc. Natl Acad. Sci. USA 78, 1421–1425 (1981).
Simons, R. W. & Kleckner, N. Translational control of IS10 transposition. Cell 34, 683–691 (1983).
Brantl, S. Regulatory mechanisms employed by cis-encoded antisense RNAs. Curr. Opin. Microbiol. 10, 102–109 (2007).
Guell, M. et al. Transcriptome complexity in a genome-reduced bacterium. Science 326, 1268–1271 (2009).
Wurtzel, O. et al. A single-base resolution map of an archaeal transcriptome. Genome Res. 20, 133–141 (2010).
Sharma, C. M. et al. The primary transcriptome of the major human pathogen Helicobacter pylori. Nature 464, 250–255 (2010).
Mitschke, J. et al. An experimentally anchored map of transcriptional start sites in the model cyanobacterium Synechocystis sp. PCC6803. Proc. Natl Acad. Sci. USA 108, 2124–2129 (2011).
Mandin, P., Repoila, F., Vergassola, M., Geissmann, T. & Cossart, P. Identification of new noncoding RNAs in Listeria monocytogenes and prediction of mRNA targets. Nucleic Acids Res. 35, 962–974 (2007).
Mraheil, M. A. et al. The intracellular sRNA transcriptome of Listeria monocytogenes during growth in macrophages. Nucleic Acids Res. 39, 4235–4248 (2011).
Nicolas, P. et al. Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis. Science 335, 1103–1106 (2012).
Kröger, C. et al. The transcriptional landscape and small RNAs of Salmonella enterica serovar Typhimurium. Proc. Natl Acad. Sci. USA 109, 1277–1286 (2012).
Wilms, I., Overlöper, A., Nowrousian, M., Sharma, C. M. & Narberhaus, F. Deep sequencing uncovers numerous small RNAs on all four replicons of the plant pathogen Agrobacterium tumefaciens. RNA Biol. 9, 446–457 (2012).
Wurtzel, O. et al. The single-nucleotide resolution transcriptome of Pseudomonas aeruginosa grown in body temperature. PLoS Pathog. 8, e1002945 (2012).
Kawano, M., Aravind, L. & Storz, G. An antisense RNA controls synthesis of an SOS-induced toxin evolved from an antitoxin. Mol. Microbiol. 64, 738–754 (2007).
Stazic, D., Lindell, D. & Steglich, C. Antisense RNA protects mRNA from RNase E degradation by RNA–RNA duplex formation during phage infection. Nucleic Acids Res. 39, 4890–4899 (2011).
Hernández, J. A. et al. Identification of a furA cis antisense RNA in the Cyanobacterium Anabaena sp. PCC 7120. J. Mol. Biol. 355, 325–334 (2006).
Rasmussen, S., Nielsen, H. B. & Jarmer, H. The transcriptionally active regions in the genome of Bacillus subtilis. Mol. Microbiol. 73, 1043–1057 (2009).
Frohlich, K. S., Papenfort, K., Berger, A. A. & Vogel, J. A conserved RpoS-dependent small RNA controls the synthesis of major porin OmpD. Nucleic Acids Res. 40, 3623–3640 (2012).
Vogel, J. et al. RNomics in Escherichia coli detects new sRNA species and indicates parallel transcriptional output in bacteria. Nucleic Acids Res. 31, 6435–6443 (2003).
Sorek, R. & Cossart, P. Prokaryotic transcriptomics: a new view on regulation, physiology and pathogenicity. Nature Rev. Genet. 11, 9–16 (2010).
Raghavan, R., Sloan, D. B. & Ochman, H. Antisense transcription is pervasive but rarely conserved in enteric bacteria. mBio 3, e00156–12 (2012).
Callen, B. P., Shearwin, K. E. & Egan, J. B. Transcriptional interference between convergent promoters caused by elongation over the promoter. Mol. Cell 14, 647–656 (2004).
Andre, G. et al. S-box and T-box riboswitches and antisense RNA control a sulfur metabolic operon of Clostridium acetobutylicum. Nucleic Acids Res. 36, 5955–5969 (2008).
Adhya, S. & Gottesman, M. Promoter occlusion: transcription through a promoter may inhibit its activity. Cell 29, 939–944 (1982).
Stork, M., Di Lorenzo, M., Welch, T. J. & Crosa, J. H. Transcription termination within the iron transport-biosynthesis operon of Vibrio anguillarum requires an antisense RNA. J. Bacteriol. 189, 3479–3488 (2007).
Giangrossi, M. et al. A novel antisense RNA regulates at transcriptional level the virulence gene icsA of Shigella flexneri. Nucleic Acids Res. 38, 362–3375 (2010).
Dühring, U., Axmann, I. M., Hess, W. R. & Wilde, A. An internal antisense RNA regulates expression of the photosynthesis gene isiA. Proc. Natl Acad. Sci. USA 103, 7054–7058 (2006).
Lee, E.-J. & Groisman, E. A. An antisense RNA that governs the expression kinetics of a multifunctional virulence gene. Mol. Microbiol. 76, 1020–1033 (2010).
Lasa, I. et al. Genome-wide antisense transcription drives mRNA processing in bacteria. Proc. Natl Acad. Sci. USA 108, 20172–20177 (2011).
Lioliou, E. et al. Global regulatory functions of the Staphylococcus aureus endoribonuclease III in gene expression. PLoS Genet. 8, e1002782 (2012).
Opdyke, J. A., Fozo, E. M., Hemm, M. R. & Storz, G. RNase III participates in GadY-dependent cleavage of the gadX-gadW mRNA. J. Mol. Biol. 406, 29–43 (2011).
Opdyke, J. A., Kang, J.-G. & Storz, G. GadY, a small-RNA regulator of acid response genes in Escherichia coli. J. Mol. Biol. 186, 6698–6705 (2004).
Tramonti, A., De Canio, M. & De Biase, D. GadX/GadW-dependent regulation of the Escherichia coli acid fitness island: transcriptional control at the gadY–gadW divergent promoters and identification of four novel 42bp GadX/GadW-specific binding sites. Mol. Microbiol. 70, 965–982 (2008).
Sakurai, I. et al. Positive regulation of psbA gene expression by cis-encoded antisense RNAs in Synechocystis sp. PCC 6803. Plant Physiol. 160, 1000–1010 (2012).
Sayed, N., Jousselin, A. & Felden, B. A cis-antisense RNA acts in trans in Staphylococcus aureus to control translation of a human cytolytic peptide. Nature Struct. Mol. Biol. 19, 105–112 (2011).
Arnvig, K. B. & Young, D. B. Identification of small RNAs in Mycobacterium tuberculosis. Mol. Microbiol. 73, 397–408 (2009).
Mandin, P. & Gottesman, S. Integrating anaerobic/aerobic sensing and the general stress response through the ArcZ small RNA. EMBO J. 29, 3094–3107 (2010).
Papenfort, K. et al. Specific and pleiotropic patterns of mRNA regulation by ArcZ, a conserved, Hfq-dependent small RNA. Mol. Microbiol. 74, 139–158 (2009).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nature Rev. Micro. 6, 613–624 (2008).
Acknowledgements
The authors thank J. Mellin for insightful discussions. This work was supported by the Pasteur–Weizmann programme (grants to P.C. and R.S.), the European Research Council (Advanced Grant 233348 to P.C. and Starting Grant 260432 to R.S.), the French Agence Nationale de la Recherche (grants Bacregrna 09-BLAN-0024-02 to P.C. and BacNet 10-BINF-02-01 to P.C. and N.S.), the Fondation Le Roch Les Mousquetaires (P.C.), the Fondation Louis-Jeantet (P.C.), the German Minerva Foundation (R.S.), the Deutsche Forschungsgemeinschaft (a German–Isaraeli Project Cooperation grant to R.S.), the French Institute National de la Recherche Agronomique (a fellowship to N.S.) and the Azrieli Foundation (a fellowship to O.W.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- CRISPR
-
(Clustered regularly interspaced short palindromic repeats). DNA sequences comprising multiple short direct repeats. These sequences, together with CRISPR-associated (Cas) proteins, constitute an adaptive immune system that is encoded by many bacteria and most archaea, and targets invading bacteriophages and conjugative plasmids.
- Gnotobiotic mice
-
Mice that are born in germ-free conditions and are then experimentally colonized with a defined microbiota or bacterium. Derived from the Greek roots gnostos, meaning known, and bios, meaning life.
- Phosphotransferase system
-
A multiprotein translocation system that is only present in bacteria and catalyses the transport and phosphorylation of numerous monosaccharides, disaccharides, amino sugars, polyols and other sugar derivatives.
- Photosystem
-
A protein complex located in the thylakoid membranes of plants, algae and cyanobacteria and in the cytoplasmic membrane of photosynthetic bacteria. Photosystem complexes carry out the primary photosynthesis reactions: the absorption of light, the transfer of electrons and the production of energy.
- Pleiotropic trans-acting asRNAs
-
Antisense RNAs (asRNAs) that affect the expression of multiple genes located at different loci in the chromosome.
- Riboswitches
-
RNA structures that are found in the 5′ UTRs of mRNAs and directly bind metabolites, metals or tRNAs to regulate the expression of the downstream sequences (which usually encode proteins involved in the synthesis or catabolism of the regulatory metabolite).
- RNA-seq
-
(RNA sequencing). High-throughput sequencing of the cDNA obtained by the reverse transcription of an RNA pool. The sequences obtained can be used to generate a quantitative genome-wide transcriptome map of an organism.
- Tiling array
-
A DNA microarray chip for which the hybridization probes are designed to overlap each other and to cover the whole genome (tiled probes), as opposed to a gene expression array, for which probes are usually designed to represent only the coding sequences of the genome and there are only a few probes per known gene.
- Transposase
-
An enzyme that catalyses the excision of a transposon.
- Transposon
-
A mobile DNA element that can excise and insert into a different locus in the genome.
Rights and permissions
About this article
Cite this article
Sesto, N., Wurtzel, O., Archambaud, C. et al. The excludon: a new concept in bacterial antisense RNA-mediated gene regulation. Nat Rev Microbiol 11, 75–82 (2013). https://doi.org/10.1038/nrmicro2934
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2934
This article is cited by
-
Improved RNA stability estimation indicates that transcriptional interference is frequent in diverse bacteria
Communications Biology (2023)
-
RNase III-CLASH of multi-drug resistant Staphylococcus aureus reveals a regulatory mRNA 3′UTR required for intermediate vancomycin resistance
Nature Communications (2022)
-
Overlapping genes in natural and engineered genomes
Nature Reviews Genetics (2022)
-
Shape of promoter antisense RNAs regulates ligand-induced transcription activation
Nature (2021)
-
Functionally uncoupled transcription–translation in Bacillus subtilis
Nature (2020)