Key Points
-
Population genetics can be applied to investigate how natural selection has shaped the variability of innate immunity genes in the human population, providing an indispensable complement to immunological as well as clinical and epidemiological genetic approaches. Recent studies have shown that the impact of selection on the different families of innate immune receptors and their downstream signalling molecules varies considerably, helping to delineate the relative functional importance of the corresponding pathways.
-
Receptors such as endosomal Toll-like receptors (TLRs) and most NACHT, LRR and pyrin domain-containing proteins (NALPs), adaptors such as myeloid differentiation primary response protein 88 (MYD88) and TIR domain-containing adapter molecule 1 (TRIF), or effector molecules such as a subset of type I interferons (IFNs) and IFNγ have been targeted by strong purifying selection, highlighting the essential and non-redundant nature of the immunological mechanisms involved.
-
Other families of innate immunity molecules, such as the RIG-I-like receptors (RLRs), the cell-surface TLRs, and most nucleotide-binding oligomerization domain-containing (NOD)/Ice protease-activating factor (IPAF)/MHC class II transactivator (CIITA) subfamily members of the NOD-like receptor (NLR) family have evolved under much weaker selective constraints, suggesting greater redundancy in the pathways triggered by these molecules.
-
Events of positive selection have targeted genetic variation at various innate immunity molecules, such as type III IFNs, attesting to the presence of functional variation at these genes and consequently in the immune responses mediated by them, which has allowed for increased host survival under specific environmental pressures, probably related to pathogen presence.
-
Striking differences in selective pressures have been detected in most genes involved in single-gene versus complex diseases, supporting the notion that life-threatening infectious diseases in childhood result mostly from rare single-gene variations, whereas the genetic component of predisposition to secondary infections in adults is more complex.
-
Thus, population genetics helps us to differentiate between innate immune genes that have a high degree of redundancy and those that are essential and non-redundant. The identification of essential genes is particularly important for molecules whose functions remain poorly described, as this helps to prioritize genes to be further studied from an immunological standpoint.
Abstract
Innate immunity involves direct interactions between the host and microorganisms, both pathogenic and symbiotic, so natural selection is expected to strongly influence genes involved in these processes. Population genetics investigates the impact of past natural selection events on the genome of present-day human populations, and it complements immunological as well as clinical and epidemiological genetic studies. Recent data show that the impact of selection on the different families of innate immune receptors and their downstream signalling molecules varies considerably. This Review discusses these findings and highlights how they help to delineate the relative functional importance of innate immune pathways, which can range from being essential to being redundant.
Similar content being viewed by others
Main
Individuals display variable ability to fight infections, as well as variable susceptibility to inflammatory and autoimmune diseases. Evidence that has accumulated since the 1950s suggests that such heterogeneity partly reflects differences in the genetic make-up of the human host1,2,3. Particularly in the past decade, genetic studies have provided numerous examples of genes accounting for differences in susceptibility to rare or common infectious diseases. The advent of new technologies, such as DNA microarrays and next-generation sequencing, has greatly accelerated the field of human genetics, making it possible to evaluate the contribution of genetic diversity to differences in immunity to infection at the level of the entire genome. For example, recent studies have revealed the power of whole-exome sequencing for dissecting the immunological mechanisms that underlie the pathogenesis of severe, rare infectious diseases4,5,6. Likewise, some genome-wide association studies on viral, bacterial and parasitic infections have increased knowledge on the genetic basis of susceptibility to common infectious diseases at the population level3,7. Although these studies have provided the proof-of-concept that increased susceptibility to infectious diseases may result from various types of inborn errors of immunity, only a few cases are fully understood at the genetic level1.
Population genetics can be applied to investigate how natural selection has shaped the variability of host defence genes in the human population8,9,10. The fields of comparative immunology and evolutionary immunobiology have greatly improved our knowledge of the origins and evolution of innate and adaptive immune systems in multiple organisms11,12. Here, we do not attempt to review these topics, but instead focus on how humans have adapted to the pressure imposed by microorganisms through variation in their genomes. Among the different types of adaptation, those affecting immune function are among the most dynamic because the counter-evolution of pathogens drives the need for continuous adaptive change (Box 1). It is precisely because of this strong and ever-renewing demand for adaptation that genes in the immune system acquire unique and clear signatures of natural selection.
Innate immunity is at the front line of host defence against pathogens and is also important for the symbiotic partnership between the host and its microbiota. Thus, innate immunity-associated genes provide an excellent model for the study of the selective pressure that is exerted by microorganisms on the host genome. The innate immune systems of plants, invertebrates and vertebrates involve genes that can be classified into various groups on the basis of their function in immune surveillance, signal transduction or effector response to microorganisms13,14,15,16, and these genes have evolved under the constant environmental pressure of microorganisms. In addition, in vertebrates, innate immunity precedes and shapes adaptive immune responses, so variations in vertebrate innate immunity genes may have important consequences for the quantity and quality of downstream B and T cell responses. Thus, the crucial roles of the proteins encoded by innate immunity genes make them ideal targets of natural selection and valuable tools for population genetics studies.
With the emergence of data sets of genomic variation in an increasing number of human populations17,18, we can now scan genomes for signatures of selection and exploit them to identify the innate immune mechanisms that have a major biological role in host defence. This Review summarizes some of the major findings regarding how selection has shaped the evolution of several families of innate immunity genes in humans and underscores how selection has refined human antimicrobial defences. We also describe how population genetics studies have provided important insights in terms of the biological relevance of the mechanisms triggered by innate immunity molecules, and we discuss the contrasting selection patterns detected in innate immune genes involved in single-gene and complex disease risk.
Natural selection in the human genome
The different modes of selection. There are different forms of selection (Fig. 1), and each of them leaves a distinctive molecular signature in the targeted genomic region. Such signatures can be detected by an increasing number of statistical tests10,19 (Box 2). Selection often maintains the prototype gene by eliminating mutations that compromise organism fitness, as most mutations are more likely to perturb rather than improve protein function. The process by which deleterious mutations are culled from the population is called purifying selection or negative selection and is the most pervasive form of selection acting on genomes. At the population level, genes tend to carry the same number of synonymous mutations (no amino acid change) and non-synonymous mutations (amino acid change), despite the fact that random mutagenesis actually generates nearly twice as many non-synonymous changes. The reduced number of non-synonymous single-nucleotide polymorphisms (SNPs) observed as compared with the non-synonymous mutation rate indicates the elimination of many non-synonymous mutations through purifying selection20.
The evolutionary fate of different types of mutations is represented in a sample of eight chromosomes. Blue circles indicate neutral polymorphisms. a | Purifying selection removes deleterious alleles (indicated by black circles) from the population. The pace at which deleterious mutations are purged from the population depends on their effect on host survival, which can range from lethal (immediately removed from the population) to mildly deleterious (tolerated but kept at low population frequencies). These mutations tend to be associated with rare, severe disorders (for example, Mendelian susceptibility to infection at the individual level). b | Positive selection increases the frequency of an advantageous mutation (indicated by a red circle) in the population. Advantageous mutations can be fixed (completed selective sweep) or polymorphic (ongoing selective sweep; not shown) in the population. Positively selected mutations are often associated with common traits (for example, higher resistance to infection at the population level), which present complex modes of inheritance. c | Balancing selection maintains polymorphism in the population as a result of heterozygote advantage and frequency-dependent advantage (not shown). In the illustrated example, a mutation (indicated by a purple circle) confers a selective advantage at the heterozygote state, so individuals who are heterozygous at this particular position (for example, individuals who possess the anaemia-associated haemoglobin D (HbS) allele sickle-cell variant and are exposed to Plasmodium falciparum) have a greater fitness than homozygous individuals.
Selection may also occur when a novel mutation is favourable in a population, which results in an increase in the frequency of the mutation. This is referred to as positive Darwinian selection, and it is thought to be one of the ways in which adaptive evolution occurs. The spread of the lactase allele, which is associated with the ability to digest milk as adults, in various human populations that adopted dairying practices is an emblematic example of positive selection21. It is also possible that variants have an advantage when rare in the population (frequency-dependent selection) or that heterozygotes for a specific mutation have the highest fitness (heterozygote advantage). These two situations give rise to balancing selection22: a selective regime that maintains variation in the population. A prime example of balancing selection is provided by the MHC region, in which high levels of polymorphism are preserved through various mechanisms, including sexual selection and pathogen diversity23,24,25,26.
The legacy of selection on our genomes. Scanning the human genome for balancing selection events is challenging and, except for the MHC locus, few genes show the molecular signature of this form of selection27.
Interspecies and intraspecies studies of protein-coding sequences have helped to identify proteins that are rapidly evolving as a result of positive selection. One of the first genome-wide selection scans compared protein-coding sequences from the human, chimpanzee and mouse genomes, and found that immunity-related genes are among the functional classes that display the clearest evidence of positive selection28. Subsequent studies have confirmed these early observations and indicated essential immunity-related genes that have undergone strong recurrent changes in both humans and non-human primates29,30,31. Similarly, interspecies studies have established that extensive purifying selection has prevented the accumulation of functional diversity on genes throughout the genome, including many genes involved in innate immunity31.
In contrast to interspecies studies, which detect old selective events, population genetic approaches (that is, intraspecific studies) detect more recent selective events within a species by making use of polymorphism data. The advantage of intraspecific approaches is that they study selection within human populations and pinpoint functionally important genomic regions that may account for phenotypic variation in health and disease. The need to secure a large collection of genome-wide polymorphism data gave rise to the International HapMap Project17 and the more recent launch of the 1000 Genomes Project18,32, which enable the detection of selection with the greatest resolution.
Numerous genome-wide selection scans in humans have been performed to date (reviewed in Ref. 33). Similarly to interspecies studies, intraspecific approaches have shown that positively selected regions of the genome are often involved in immunity34,35,36,37. More than 300 immunity-related genes were detected as putative targets of positive selection8, although a fraction of them could be spurious signals and further validation is needed33. Nevertheless, these observations suggest that functional variations in a large proportion of immunity genes have conferred a specific selective advantage for host survival, including protection from pathogens and tolerance to microbiota.
Selection and innate immunity in humans
Besides genome-wide scans, another highly effective means of gaining insight into the role of selection on immune processes is to take a targeted approach that focuses on genes with particular immune functions: such an approach would make subsequent functional investigations more feasible. Innate immunity involves the coordinated action of families of receptors, known as pattern-recognition receptors (PRRs) or microbial sensors, that respond to a wide range of microorganisms through the detection of specific, conserved microbial patterns or molecules14,15,38. The biological importance of these microbial sensors is highlighted by the fact that many of them share remarkable structural and functional similarities among vertebrates, invertebrates and plants16 (Box 3). In humans, several receptor families are involved in the cellular arm of innate immunity, including the membrane-bound Toll-like receptors (TLRs) and C-type lectin receptors (CLRs), and the cytosolic RIG-I-like receptors (RLRs), NOD-like receptors (NLRs) and other DNA sensors39,40,41,42,43. In addition, complement receptors and ficolins are circulating proteins that constitute the humoral arm of innate immunity44. Following ligand binding, receptors induce the activation of distinct signalling pathways that involve adaptor molecules, which mediate signalling, and effector molecules, such as interferons (IFNs) or antimicrobial peptides (AMPs), which are required for the eradication of pathogens or danger signals.
In this section, we discuss the major differences in the evolution of some families of innate receptors and their downstream molecules. We focus on the genes that have been most studied by population genetic analyses in humans and closely related primate species. Moreover, we explain how the observed differences in the intensity and form of selection can inform us about the functional relevance of these innate immunity genes8,9,10,45. Notably, genes targeted by strong purifying selection are likely to fulfil functions that are essential. By contrast, genes evolving under more relaxed constraints (such as weaker purifying selection or neutrality) may have variably redundant functions in immune signalling. Finally, genes targeted by positive or balancing selection are possibly involved in mechanisms in which genetic variation has conferred a selective advantage to the population.
Membrane-bound receptors: the case of TLRs. Population genetics studies have provided important insights in terms of the evolution and function of TLRs45,46,47,48,49,50,51,52,53. In humans, TLRs are broadly subdivided into two groups: those primarily expressed at the cell surface (TLR1, TLR2, TLR4, TLR5, TLR6 and TLR10), which detect predominantly pathogen-associated molecular patterns from bacteria, fungi and protozoa, and those expressed within endosomes (TLR3, TLR7, TLR8 and TLR9), which primarily recognize nucleic acids from viruses and bacteria14,38,39.
Comparative interspecies studies have shown that species-wide positive selection has accelerated the divergence of some TLRs among primates, with the strongest evidence obtained for TLR1 and TLR4 (Refs 48, 51, 53). These two TLRs contain the highest number of positively selected codons, several of which are involved in ligand binding or in interaction with the TLR4 co-receptor MD2 (also known as LY96)51. This high interspecies divergence suggests that the presence or absence of pathogens that are sensed by TLR1 and TLR4 may be one of the most important variants in the ecological niches of the different primate species.
Within species, and in humans in particular, balancing selection was initially proposed to be pervasive among innate immunity genes52, including some TLRs. However, most intraspecific studies thus far converge towards the opposite conclusion: TLRs, taken as a set, have mainly evolved under the action of purifying selection46,47,49,51, which highlights their indispensability. More precisely, intracellular TLRs evolve under the strongest purifying selection (Fig. 2), and neither nonsense nor damaging missense mutations are tolerated at these genes46. By contrast, the selective constraints on cell-surface TLRs are much more relaxed, and these TLRs present high levels of genetic and functional diversity across populations46,54. In the human population, up to 23% of individuals have damaging missense mutations and up to 16% present a nonsense mutation in at least one cell-surface TLR46,50, suggesting higher redundancy. The case of TLR5 — the receptor of flagellin — is particularly extreme, as the R392X nonsense variant, which has a dominant-negative effect55, is present at high frequencies (up to 23%) in Europeans and South Asians46,50. This suggests that TLR5 is not crucial for immunity to infection against at least some flagellated bacteria, and that accessory mechanisms of flagellin recognition, such as those involving the cytosolic receptor Ice protease-activating factor (IPAF; also known as NLRC4)56, may provide sufficient protection.
Genes that have undergone purifying selection are shown in red, and those that have evolved under weaker selective constraints are shown in blue. Genes for which no significant evidence of a selective constraint was observed are shown in grey. Colours reflect the intensity of the selective constraints on amino acid-altering variation, as obtained by the McDonald–Kreitman Poisson random field method McDonald-Kretiman Poisson random field method (omega and gamma)31 (Box 2). Genes presenting robust signatures of positive selection in all humans are outlined with a thick black line, whereas genes presenting robust signatures of positive selection that are restricted to specific human populations are outlined with a dashed black line. Endosomal Toll-like receptors (TLRs) show signs of stronger purifying selection than do cell-surface TLRs. Myeloid differentiation primary response protein 88 (MYD88) is the TLR adaptor molecule that has evolved under the strongest purifying selection, which indicates its central role as a pan-adaptor molecule. Furthermore, all adaptors have been targeted by positive selection, either in the entire human lineage (MYD88 and sterile alpha and TIR motif-containing protein (SARM)) or in specific human populations (TIR domain-containing adapter molecule 1 (TRIF), TIR domain-containing adapter molecule 2 (TRAM) and Toll/interleukin-1 receptor domain-containing adapter protein (MAL)), suggesting advantages in terms of immunity that are shared by all humans or that are due to geographically restricted microbial exposure, respectively. Purifying selection has driven the evolution of most NACHT, LRR and pyrin domain-containing proteins (NALPs), whereas other cytosolic microbial sensors, such as the NOD-like receptors (NLRs), Ice protease-activating factor (IPAF) and MHC class II transactivator (CIITA) and most NOD subfamily members, as well as the RIG-I-like receptors (RLRs), have evolved under weaker constraints. NLR family, apoptosis inhibitory protein (NAIP) is not represented in the figure as no population genetic data are available. DAMPs, damage-associated molecular pattern molecules; HA, haemagglutinin; IFNs, interferons; IRFs, IFN-regulatory factors; LAM, lipoarabinomannan; LPS, lipopolysaccharide; LTA, lipoteichoic acid; MAPK, mitogen-activated protein kinase; MDA5, melanoma differentiation-associated protein 5; NF-κB, nuclear factor-κB; NLRC, NLR family CARD domain containing; NLRX1, NLR family member X1; NOD, nucleotide-binding oligomerization domain-containing; PG, peptidoglycan; PLM, phospholipomannan; RIG-I, retinoic acid-inducible gene I protein; tGPI, Trypanosoma cruzi-derived glycosylphosphatidylinositol.
The evolutionary relaxation characterizing some TLRs appears to be specific to humans when compared with other primates51. This observation has been associated with reductions in the size of human populations (during the out-of-Africa migration), which compromised the efficiency of purifying selection and resulted in an increased frequency of deleterious alleles57. However, it is tempting to speculate that such an evolutionary relaxation might also indicate that pathogens recognized by cell-surface TLRs have imposed a milder burden on humans than on other primates. In some cases, genetic variation at cell-surface TLRs may be advantageous for the host, as attested by the signature of positive selection detected in the TLR6–TLR1–TLR10 cluster (Box 4).
Together, the major differences in the intensity of selection between intracellular and cell-surface TLRs indicate that the two groups differ in their biological relevance, with intracellular TLRs being essential and cell-surface TLRs being more redundant. This evolutionary dichotomy is likely to reflect the differences of the two TLR groups in terms of their subcellular localization, the type of microorganisms and ligands that they target, and their self-reactive potential14,38,39. Cell-surface TLRs detect microbial molecules, such as bacterial cell-surface lipopolysaccharides or flagellin, which are usually very distinct from host molecules. By contrast, intracellular TLRs principally sense nucleic acids, such as double-stranded RNA of viruses or the unmethylated CpG islands of bacterial and viral DNA. Although these microbial nucleic acids can be distinguished from host nucleic acids on the basis of specific chemical modifications58, there is still the risk that intracellular TLRs could be stimulated 'inappropriately' by self-derived nucleic acids, which may lead to autoimmunity59,60,61,62. Thus, the extreme conservation of intracellular TLRs might reflect the very narrow window they have to ensure effective pathogen sensing while preventing dangerous recognition of host nucleic acids and thereby minimizing the risk of autoimmunity.
Cytosolic receptors: the RLRs and NLRs. Recent population genetics data have increased our understanding of the relative importance of the different members of the RLR and NLR families63,64,65. Similarly to intracellular TLRs, RLRs and NLRs detect signs of infection or danger within the cell14,40,41,42. The three RLR members, retinoic acid-inducible gene I protein (RIG-I; also known as DDX58), melanoma differentiation-associated protein 5 (MDA5; also known as IFIH1) and LGP2 (also known as DHX58), detect viral RNA and induce inflammatory cytokines and type I and type III IFNs41. RIG-I and MDA5 sense directly viral molecules, whereas LGP2 acts as a regulator of RIG-I and MDA5 signalling. The NLR proteins, which have a structure that closely resembles that of the resistance proteins (also known as R proteins) in plants16, are encoded by a family of 22 genes, which include the 14 members of the large NACHT, LRR and pyrin domain-containing protein (NALP) subfamily (NALP1–NALP14), the five nucleotide-binding oligomerization domain-containing (NOD) proteins (NOD1, NOD2, NLR family CARD domain containing 3 (NLRC3), NLRC5 and NLR family member X1 (NLRX1)) and other proteins, such as MHC class II transactivator (CIITA), IPAF and NLR family, apoptosis inhibitory protein (NAIP)40,42. NLRs either activate nuclear factor-κB and mitogen-activated protein kinases to induce inflammatory responses or participate in large cytoplasmic protein complexes called inflammasomes40,42.
RLRs evolve under weaker constraints (weak purifying selection or neutrality) compared with other families of innate receptors, and this suggests that there is some degree of redundancy in their roles (Fig. 2). Within the RLR family, RIG-I displays the most constrained amino acid-altering variation, particularly in the carboxy-terminal and helicase domains (which are involved in RNA recognition)65. This may reflect the fact that RIG-I senses a broader range of RNA viruses than MDA5, which only recognizes picornaviruses. Moreover, the particular structure of RIG-I viral substrates (short double-stranded RNAs and 5′-triphosphate single-stranded RNAs)41 may have stricter binding requirements than MDA5 substrates (long double-stranded RNAs).
Among NLRs, most NALPs have evolved under strong purifying selection (10 of the 14 NALPs)64 (Fig. 2), reflecting the rapid elimination from the population of mutations at these genes as a result of their highly deleterious effects. These observations are consistent with the idea that NALPs have acquired a function that is essential and non-redundant in host survival. By contrast, the weaker selective constraints characterizing IPAF, CIITA and most NOD subfamily members64 testify to their higher degree of redundancy.
In some cases, certain genetic variants at some RLRs or NLRs (for example MDA5, LGP2, NALP1, NALP14 or CIITA) have been targeted by positive selection in specific human populations, suggesting that functional variation at these genes and the resulting changes in the downstream immunological mechanisms have allowed for increased host survival under specific environmental pressures63,64,65. Taken together, population genetics studies have shown that the NALPs have evolved under much stronger selective constraints than other NLR members and RLRs, and the extreme NALP conservation may reflect the type of the ligands that are likely to be sensed by these molecules and/or the important nature of the responses triggered by them. These hypotheses need now to be experimentally validated.
Soluble receptors: the humoral arm. In contrast to the cellular receptors described above, there are only scarce population genetics studies on secreted receptors, such as complement receptors, collectins, ficolins and pentraxins, which are all involved in highly diverse functions (for example, phagocytosis, opsonization and apoptosis)44,66,67. However, the case of the mannose-binding lectin (MBL) — an extensively-studied collectin that binds a broad range of microorganisms and activates the lectin–complement pathway66,68 — neatly illustrates the value of the population genetics approach for determining the ecological relevance of genes and mechanisms in immunity to infection. Conflicting results have been obtained by clinical and epidemiological genetic studies as to the detrimental or beneficial effects of MBL deficiency69,70. MBL deficiency has been associated, although not conclusively, with increased susceptibility to several infectious diseases, such as meningococcal infection or HIV infection71. However, MBL-deficiency alleles are common worldwide (they occur in up to 30% of individuals)69,72,73, suggesting that they may also have a protective effect, as reported for infections with intracellular pathogens such as Mycobacterium tuberculosis and Leishmania spp.74,75.
Some population genetics studies have proposed that balancing selection has driven MBL2 diversity47,72, but others have failed to detect any evidence of selection favouring the increased frequency of deficiency alleles73,76. Indeed, when analysing larger population data sets, the pattern of MBL2 variation appears to be consistent with neutral evolution73. These population genetics observations provide novel insights that inform the long-standing and highly controversial debate as to whether MBL is protective, deleterious or both, and they suggest instead that the immunological mechanisms triggered by this lectin are largely redundant. So, other molecules or pathways, such as the ficolins or the C1q-dependent classical pathway, may compensate for MBL deficiency. Future studies exploring whether the genes involved in these pathways have compensatory mutations in MBL-deficient individuals should help to substantiate this hypothesis.
Adaptor molecules: the TIR domain-containing adaptors. Linking the evolutionary fate of innate immune receptors with their corresponding adaptors, which associate with receptor proteins and initiate downstream signalling14, can further increase our knowledge of the biological relevance of specific immune pathways. This is best exemplified by population genetics data on the five Toll/IL-1R (TIR) domain-containing adaptors: myeloid differentiation primary response protein 88 (MYD88), Toll/interleukin-1 receptor domain-containing adapter protein (MAL; also known as TIRAP), TIR domain-containing adapter molecule 1 (TRIF; also known as TICAM1), TIR domain-containing adapter molecule 2 (TRAM; also known as TICAM2) and sterile alpha and TIR motif-containing protein (SARM), which are involved in the initiation of signalling downstream of TLRs and IL-1R-like receptors14,77. Across primates, the five adaptors have evolved under strong selective constraints48. In humans, MYD88 and TRIF display the highest degree of purifying selection (Fig. 2), indicating that the signals mediated by these two molecules are essential and non-redundant78. Conversely, MAL, TRAM and SARM are subject to more relaxed constraints, which indicates that their functions are more dispensable. The selective regime that has constrained the evolution of adaptor proteins to different extents has not prevented these genes from undergoing some episodes of positive selection78,79.
Collectively, the integration of population genetics data from the TLRs and TIR-containing adaptors has shed light on the mechanisms and pathways triggered by these molecules78 (Fig. 2). MYD88 has a pan-adaptor role77, so a strong constraint is expected, but TRIF has also evolved under strong purifying selection despite not functioning as a pan-adaptor. TRIF is involved in signalling downstream of TLR3 and TLR4 (Ref. 77), and of these two only TLR3 evolves under purifying selection46. Indeed, clinical genetic studies have revealed that rare mutations affecting the TLR3–TRIF pathway underlie herpes simplex virus 1 (HSV-1) encephalitis in childhood45,80,81. This suggests that the entire pathway activated by TLR3 in a TRIF-dependent manner is non-redundant in host defence, at least against HSV-1 (Refs 45, 78). Together with immunological, clinical and epidemiological approaches, future population genetics studies aiming to dissect the selective signatures at the level of entire signalling pathways will allow us to further delineate the most critical mechanisms involved in innate immunity to infection.
Effector molecules: interferons and AMPs. The signalling pathways initiated by adaptor proteins culminate in the expression of cytokines, including interleukins and IFNs, chemokines, members of the tumour necrosis factor (TNF) family and growth factors. Among these effector molecules, IFNs have been extensively studied82, but the biological relevance of the different IFN subtypes for host survival remains largely unknown. Like other families of innate immunity genes, IFNs belong to a multigene family of highly paralogous loci, so gene conversion can markedly alter their genetic diversity, making population genetics data more difficult to interpret83. Recent analyses have shown that the three IFN families and their individual members have followed very different evolutionary trajectories, ranging from highly constrained to redundant and expendable84,85.
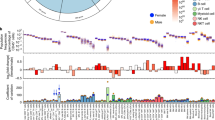
Some type I IFNs, such as IFNα6, IFNα8, IFNα13 and IFNα14 and the type II IFNγ, have evolved under strong purifying selection, displaying very low levels or even a complete absence, as in the case of IFNγ, of amino acid-altering genetic variation (Fig. 3a). Clinical genetic studies have shown that immunodeficiencies due to defects in the type I IFN pathway strongly affect antiviral immunity, and disorders of the IFNγ circuit are associated with Mendelian susceptibility to mycobacterial disease (MSMD)82. The integration of population and clinical genetic data thus indicates that at least IFNγ and a subgroup of type I IFNs have a crucial, non-redundant role in antiviral and anti-mycobacterial immunity, respectively82,84,85.
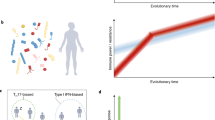
a | Type I, type II and type III interferons (IFNs) display different levels of functional diversity. The circles represent the proportion of chromosomes for each IFN subtype carrying different types of functional variants in the general population. For each circle, the proportion of chromosomes carrying at least one non-synonymous polymorphism is shown in red, and the proportion carrying at least one nonsense polymorphism is shown in black. The blue segment corresponds to the proportion of chromosomes carrying neither non-synonymous nor nonsense polymorphisms. The IFN subtypes in boxes are those for which statistical significance of strong purifying selection was obtained84,85. A schematic representation of the signalling pathways activated by the interaction of type I, type II and type III IFNs with their corresponding receptors is also presented. b | Type III IFNs are the only group of IFNs that have evolved under the action of positive selection, specifically in European and Asian populations. The scheme shows the distribution of the genetic variants under positive selection across the genomic region in which the three type III IFN genes are located. Note that one of the positively selected single-nucleotide polymorphisms (SNPs) in the interleukin-28B (IL28B) region (SNP −3180A>G, rs12979860) has been recently found to lay within the newly discovered IFNL4 gene, which is located upstream of IL28B123. Highlighted mutations result in amino acid changes, whereas the rest are non-coding SNPs. Signatures of positive selection were detected in Asia for the variants in IL28A and IL28B, and in both Asia and Europe for that in IL29. Figure is modified from Ref. 84. GAS, IFNγ-activated site; IFNAR, IFNα/β receptor; IFNγR, IFNγ receptor; IFNλR1, IFNλ receptor 1 (also known as IL-28Rα); IL-10Rβ, IL-10 receptor-β; IRF9, IFN regulatory factor 9; ISRE, interferon-stimulated response element; JAK, Janus kinase; STAT, signal transducer and activator of transcription; TYK, tyrosine kinase.
By contrast, other type I IFN subtypes (mainly IFNα10 and IFNɛ, but also IFNα1, IFNα4, IFNα7, IFNα16 and IFNα17) have accumulated missense or nonsense mutations at high population frequencies (Fig. 3a), suggesting that their functions largely overlap with those of other IFN subtypes84. The varying degrees of genetic diversity and redundancy displayed by type I IFNs suggest that there may be variability in the antiviral potencies and/or expression pattern of the different IFN subtypes. Finally, genetic variants at type III IFNs have been targeted by positive selection, conferring a selective advantage to Eurasian populations84 (Fig. 3b).
Another set of effector molecules that have been thoroughly studied with population genetics approaches are the AMPs. In particular, the defensins are an AMP family with broad antimicrobial activities and non-immune functions, such as in reproduction or cell signalling86. In humans, the α- and β-defensin multigene families contain multiple paralogous genes, with some genes displaying copy-number variability87,88. Comparative analyses of α- and β-defensins in human and non-human primates have revealed that episodes of purifying, positive and balancing selection have driven the evolution of these gene families88,89,90,91. These complex patterns may reflect the need to preserve the functional integrity of these molecules while favouring, in several cases, functional diversity to provide responses to a wide range of pathogens. An interesting example of increased functional diversity is provided by the human β-defensin gene DEFB1, which has evolved under the action of long-term balancing selection91, a process that is thought to be extremely rare outside the MHC genes. The balancing selective event has been localized to the promoter region of DEFB1, where increased functional variation (through heterozygosity) appears to have conferred a selective advantage, possibly against severe sepsis91,92.
Overall, population genetics studies of effector molecules such as IFNs or AMPs emphasize the complex events of selection characterizing these molecules. Although the nature of such variable pressures is not totally clear, the identification of individual genes, such as specific IFN subtypes, subject to strong constraints or to positive evolution paves the way for additional studies to evaluate the potential of these molecules for use in vaccination, diagnosis and treatment.
Selection patterns and immunological relevance
The examples discussed above illustrate how the delineation of the selection mode that has driven the diversity of innate immunity genes can complement immunological and clinical and epidemiological genetic studies. Receptors such as endosomal TLRs and most NALPs, adaptors such as MYD88 and TRIF or effector molecules such as a subset of type I IFNs and IFNγ have been targeted by strong purifying selection46,48,51,64,78,84,85, highlighting the essential and non-redundant nature of the immunological mechanisms involved. Moreover, the fact that selective constraints acting on RLRs are weaker than those acting on endosomal TLRs46,51,65, even though both types of receptors detect nucleic acids, might reflect some degree of redundancy for RLR-mediated antiviral immunity. Similarly, the weaker constraints acting on most NOD, IPAF and CIITA subfamily members and cell-surface TLRs with respect to NALPs46,64 suggest greater redundancy in the pathways triggered by the former molecules in response to microbial products and stress signals (Fig. 2). Extreme cases of redundancy are provided by molecules such as MBL or TLR5, for which the frequency of loss-of-function alleles can increase in the population by genetic drift46,50,73,76. This contrasts with other situations in which gene loss has conferred an advantage to the host. For example, the increased frequencies of loss-of-function alleles of DARC (which encodes Duffy antigen/chemokine receptor) or CASP12 (which encodes inactive caspase 12) is a result of positive selection that is likely to be due to their protective effects against Plasmodium vivax infection and sepsis, respectively93,94.
The occurrence of positive selection attests to more dynamic immunological mechanisms, variation in which has been beneficial for the host8,9,10,45. These positive selection events can be ancient and shared by all humans. Such is the case for the adaptor MYD88 (Ref. 78): a functional change in this protein may have favoured the survival of the entire human species. The human MYD88 differs from its primate orthologue by a single amino acid substitution (H80Q), which might represent the target of selection48,78. This variant is located within the DEATH domain, which is crucial for downstream protein–protein interactions, and mutations in this domain have been associated with defective immunity to infection95,96. Conversely, other events of positive selection seem to be more recent and restricted to specific populations, such as those detected at some RLRs, NLRs, TIR-containing adaptors and type III IFNs63,64,65,78,79,84. So, the advantage conferred by such events is more dependent on environmental variables, possibly related to exposure to pathogens. For example, a variant in MDA5 (R460H) has been targeted by positive selection in Europeans and Asians63,65, suggesting an advantage to host defence that could be related to the sensing of particular RNA viruses by this sensor. Population genetics thus helps us to differentiate between genes with a high degree of redundancy and genes that are essential and non-redundant. The identification of essential genes is particularly important for molecules with poorly described functions, such as most NALPs, as this helps to prioritize which genes should be studied from an immunological standpoint.
Does the evolution of innate immunity genes and their signatures of selection reflect a quest for improved host defence against pathogenic microorganisms? The answer is yes, but not exclusively. It is obvious that pathogens are likely to be a major force exerting pressure on host genes26,97. However, we coexist with millions of symbiotic, generally non-pathogenic microorganisms, and these can also be recognized by innate immunity receptors98,99,100,101,102. Furthermore, these receptors are not only involved in the mere sensing of microorganisms but also in functions that ensure tissue development and tissue homeostasis, including inflammatory control, autophagy and apoptosis. For example, some TLRs appear to be involved in central nervous system development, and some NALPs seem to have roles in intestinal homeostasis, early development and reproduction40,103,104. Moreover, innate receptors, including TLRs and RLRs, have also been implicated in autoimmune pathogenesis60,61,105,106,107. In light of this, the traditional view of pathogens as the only force driving the evolution of immunity genes may be too simplistic, as non-infection-related factors may have further contributed to the patterns of selection observed at these genes. For example, given the increasing range of functions attributed to NALP3, which acts as both a microbial sensor and a regulator of intestinal homeostasis40,104, its extreme conservation may not only attest to a major role in controlling infection but also reflect the evolutionary equilibrium reached by the host to maintain a peaceful coexistence with the symbiotic microbiota.
Selection and disease susceptibility
Unequal selective pressures are expected to be exerted on genes associated with Mendelian, single-gene disorders or with complex disease risks31,108. At the genome-wide level, genes associated with Mendelian disease are enriched in signs of purifying selection31,34,109, as their deleterious mutations are usually not transmitted to the next generation as a result of early death, and thus have low (<1%) population frequencies110 (Fig. 1).
In the context of innate immunity, for example, mutations in the strongly constrained TIR–MYD88 and TLR3–TRIF pathways have been associated with increased susceptibility to life-threatening infections by pyogenic bacteria96,111 and HSV-1 (Refs 80, 81), respectively. Likewise, mutations in NALP3, which is subject to the highest degree of purifying selection among NALPs64, have been associated with severe inflammatory diseases104,112,113. Finally, mutations in genes affecting the production or activity of IFNγ, which is subject to the strongest purifying selection of all IFNs84,85, have been associated with MSMD82,114. These clinical examples further support the notion that immunity-related genes that evolve under purifying selection are of major biological relevance in host survival, and their mutations are likely to predispose individuals to early-onset, life-threatening disease115.
Mutations associated with complex disease risk generally have a lower penetrance and are therefore able to reach higher frequencies in the population (>5–10%) than single-gene disease susceptibility alleles20,108. Genes with alleles that contribute to complex diseases in adults display signs of less pervasive purifying selection109,115. For example, although missense mutations are tolerated in some innate immunity genes (for example, TLR1, TLR4 and TLR10), weak negative selection keeps them at low population frequency (Fig. 2), attesting to their non-negligible impact on host fitness46,49. Likewise, mutations in weakly constrained or even neutrally evolving genes might subtly modulate complex susceptibility to disease, as illustrated by the case of MBL, variation in which may have an effect in particular, narrow conditions, such as co-existent morbidity116. Thus, weakly constrained genes generally have a more modest impact on host survival and may be involved in complex susceptibility to infection at the population level.
Finally, genes associated with complex disease susceptibility are generally enriched for signals of positive selection31,37,109,117. Specifically, positively selected variants in TLR1, MDA5, CIITA or NALP1 (Refs 46, 63, 64, 65) have been associated with infectious, autoimmune or inflammatory diseases106,118,119,120,121,122. Likewise, variation in type III IFNs, including five SNPs in IL28B (also known as IFNL3), two in IL28A (also known as IFNL2) and one in IL29 (also known as IFNL1), has been targeted by positive selection84 (Fig. 3b). Interestingly, the positively selected SNPs in the IL28B region, one of which now lies within the newly discovered IFNL4 gene123, have been associated with spontaneous clearance of hepatitis C virus (HCV) and a better response to treatment for chronic HCV infection123,124,125,126,127. This suggests that the nature of the selective advantage conferred by these IFNs is increased resistance to viral infection. Collectively, the overlap between the positive selection events detected at some genes and their association with susceptibility to complex disease provides an important proof-of-concept for the added, predictive value of population genetics. This is particularly important for evaluating the potential implications in human disease of other, as-yet-uncharacterized variants targeted by positive selection.
Furthermore, there is increasing evidence to suggest that, in some cases, positively selected mutations lead, directly or indirectly, to maladaptation and are associated with immune dysfunction, such as autoimmunity and inflammation8,128. For example, the positively selected MDA5 R460H variant seems to increase the risk of psoriasis106. Similarly, positive selection has increased the frequency of the V1059M mutation in NALP1 among Europeans64, despite also increasing the frequency of the linked L155H NALP1 mutation, which is associated with autoimmune disorders including Addison's disease, type 1 diabetes and vitiligo118,129. Likewise, DEFB1 haplotypes that have been proposed to be maintained by balancing selection because they confer protection against sepsis91,92 seem to predispose to asthma and atopy130. Although further clinical and epidemiological genetics work is needed, these examples provide additional evidence in favour of the hygiene hypothesis, according to which the current increased incidence of autoimmune and inflammatory disorders may result, at least partially, from past events of selection that increased host resistance to infection128.
The distinction between genes involved in single-gene disorders and complex diseases, as well as between the type of selective patterns that are expected to characterize them, is not always clear-cut. For example, genes associated with inflammatory bowel disease (IBD), the genetic basis of which is complex, are enriched in genes involved in Mendelian primary immunodeficiencies, including MSMD131. Likewise, genes can display complex signatures of selection, such as those detected at NOD2, variation in which has been associated with Crohn's disease131,132. At the gene level, the diversity of NOD2 is largely consistent with neutrality64, but some NOD2 Crohn's disease-risk alleles have been proposed to be maintained in the European population by balancing or recent positive selection131,133, although this remains highly controversial. Such conflicting signatures of selection may reflect the complex, multiple phenotypes with which NOD2, or its interacting gene networks, have been associated, including Crohn's disease, ulcerative colitis, Blau syndrome and mycobacterial infections131.
Future perspectives
Here, we have described specific examples of how population genetics studies can offer important functional insights into innate immunity. The evolutionary dissection of innate immunity genes allows us to rank them with respect to their biological relevance and also informs us about their respective contribution to immunity-related disorders. However, immune systems do not evolve one gene at a time; instead, selective pressures are likely to affect multiple interacting genes, resulting in polygenic adaptation134. Furthermore, such pressures can be numerous and related to various diseases simultaneously, as in the case of NOD2 alleles. Thus, genes in functional networks might show coordinated signatures of selection, as recently proposed for a group of antibacterial innate immunity genes (that is, some TLR pathways), in which the position of each gene in the network conditions its evolvability and therefore its adaptability135. The recent advent of whole-genome sequencing data sets from human populations provides us with tools to test these hypotheses and also enables the analysis of additional families of innate immunity molecules, including CLRs, cytosolic DNA sensors, chemokines and members of the TNF family. For this, improved statistical methods for detecting epistatic interactions or selection in gene networks are needed.
Population genetics studies of selection have traditionally followed a gene-centric view, focusing on qualitative changes at the protein level. However, selection can also target non-coding regions of the genome, such as regulatory elements136. For example, changes in gene expression levels can indeed constitute a target for adaptive evolution, as illustrated by the fact that polymorphisms in regulatory elements (that is, expression quantitative trait loci (eQTLs)) are enriched for signs of positive selection137. Furthermore, regulatory variation is increasingly documented as being associated with phenotypic variation, both benign and disease-associated136,138,139. Multidisciplinary approaches (such as those used by the ENCODE Consortium136) that combine whole-genome sequencing data, expression and proteomic profiles, chromatin accessibility and DNA methylation marks from different cell types challenged with different immune stimuli should facilitate an unbiased assessment of the immunological mechanisms favouring our past and present survival in the natural setting.
Finally, future population genetics studies may help us to understand diverse aspects of immune function, including interaction with the gut microbiota, interplay with viruses and transposable elements, and gene–environment interactions. Indeed, the extent to which host genetic diversity drives, or co-evolves with, changes in the microbiota remains to be explored in much further detail. Similarly, detailed genotype-to-phenotype studies of large population cohorts from different ethnic backgrounds and exposed to different environments will help us to delineate the relative contribution of the modern environment to the present-day natural variation of immune responses. Notably, the change in dietary habits and the use of domesticated animals since the advent of agriculture 10,000 years ago (which is an instant in evolutionary time), life in large cities and the extended use of modern antibiotics have radically altered our exposure to microorganisms. As our immune systems evolved in a totally different context of diet and microbial exposure, interaction of our genetic make-up with these new environmental factors may lead to phenotypes associated with disease states, such as inappropriate inflammatory responses or autoimmunity. So, the integration of all these data into a population genetics framework will help us to identify immunological mechanisms with adaptive or maladaptive roles in the immunity of modern human populations.
The challenge that lies ahead now is to apply knowledge of population genetics to infer molecular details of immune responses and design effective therapies.
References
Alcais, A., Abel, L. & Casanova, J. L. Human genetics of infectious diseases: between proof of principle and paradigm. J. Clin. Invest. 119, 2506–2514 (2009).
Casanova, J. L. & Abel, L. Human genetics of infectious diseases: a unified theory. EMBO J. 26, 915–922 (2007).
Chapman, S. J. & Hill, A. V. Human genetic susceptibility to infectious disease. Nature Rev. Genet. 13, 175–188 (2012).
Bolze, A. et al. Whole-exome-sequencing-based discovery of human FADD deficiency. Am. J. Hum. Genet. 87, 873–881 (2010).
Byun, M. et al. Whole-exome sequencing-based discovery of STIM1 deficiency in a child with fatal classic Kaposi sarcoma. J. Exp. Med. 207, 2307–2312 (2010).
Liu, L. et al. Gain-of-function human STAT1 mutations impair IL-17 immunity and underlie chronic mucocutaneous candidiasis. J. Exp. Med. 208, 1635–1648 (2011). References 4–6 report the first identifications of Mendelian defects related to an infectious disease using whole-exome approaches.
Vannberg, F. O., Chapman, S. J. & Hill, A. V. Human genetic susceptibility to intracellular pathogens. Immunol. Rev. 240, 105–116 (2011).
Barreiro, L. B. & Quintana-Murci, L. From evolutionary genetics to human immunology: how selection shapes host defence genes. Nature Rev. Genet. 11, 17–30 (2010).
Quintana-Murci, L., Alcais, A., Abel, L. & Casanova, J. L. Immunology in natura: clinical, epidemiological and evolutionary genetics of infectious diseases. Nature Immunol. 8, 1165–1171 (2007).
Sabeti, P. C. et al. Positive natural selection in the human lineage. Science 312, 1614–1620 (2006).
Cooper, M. D. & Alder, M. N. The evolution of adaptive immune systems. Cell 124, 815–822 (2006).
Kimbrell, D. A. & Beutler, B. The evolution and genetics of innate immunity. Nature Rev. Genet. 2, 256–267 (2001).
Lemaitre, B. & Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 25, 697–743 (2007).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Medzhitov, R. Recognition of microorganisms and activation of the immune response. Nature 449, 819–826 (2007).
Spoel, S. H. & Dong, X. How do plants achieve immunity? Defence without specialized immune cells. Nature Rev. Immunol. 12, 89–100 (2012).
The International HapMap 3 Consortium. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Nielsen, R., Hellmann, I., Hubisz, M., Bustamante, C. & Clark, A. G. Recent and ongoing selection in the human genome. Nature Rev. Genet. 8, 857–868 (2007). This study and reference 10 are excellent reviews of how natural selection acts. They list the signatures and implications of natural selection, along with the different methods used for detecting selection on the basis of population genetic data.
Kryukov, G. V., Pennacchio, L. A. & Sunyaev, S. R. Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am. J. Hum. Genet. 80, 727–739 (2007).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nature Genet. 39, 31–40 (2007). An elegant example of a second independent mutation in the same gene resulting in a parallel adaptation in humans to metabolize milk throughout life.
Charlesworth, D. Balancing selection and its effects on sequences in nearby genome regions. PLoS Genet. 2, e64 (2006).
Chaix, R., Cao, C. & Donnelly, P. Is mate choice in humans MHC-dependent? PLoS Genet. 4, e1000184 (2008).
Hedrick, P. W., Whittam, T. S. & Parham, P. Heterozygosity at individual amino acid sites: extremely high levels for HLA-A and -B genes. Proc. Natl Acad. Sci. USA 88, 5897–5901 (1991).
Hughes, A. L. & Nei, M. Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335, 167–170 (1988).
Prugnolle, F. et al. Pathogen-driven selection and worldwide HLA class I diversity. Curr. Biol. 15, 1022–1027 (2005).
Andres, A. M. et al. Targets of balancing selection in the human genome. Mol. Biol. Evol. 26, 2755–2764 (2009).
Clark, A. G. et al. Inferring nonneutral evolution from human-chimp-mouse orthologous gene trios. Science 302, 1960–1963 (2003). The first genome-wide analysis of genes evolving under positive selection in the human evolutionary lineage.
Nielsen, R. et al. A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol. 3, e170 (2005).
Kosiol, C. et al. Patterns of positive selection in six Mammalian genomes. PLoS Genet. 4, e1000144 (2008).
Bustamante, C. D. et al. Natural selection on protein-coding genes in the human genome. Nature 437, 1153–1157 (2005).
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012). This study and reference 18 report the first large-scale population study based on complete human genome sequences, as part of the 1000 Genomes Project.
Akey, J. M. Constructing genomic maps of positive selection in humans: where do we go from here? Genome Res. 19, 711–722 (2009).
Barreiro, L. B., Laval, G., Quach, H., Patin, E. & Quintana-Murci, L. Natural selection has driven population differentiation in modern humans. Nature Genet. 40, 340–345 (2008). A clear demonstration that natural selection, often pathogen mediated, drives adaptive differences among geographically distinct human populations.
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Pickrell, J. K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Sabeti, P. C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–918 (2007).
Beutler, B. et al. Genetic analysis of host resistance: toll-like receptor signaling and immunity at large. Annu. Rev. Immunol. 24, 353–389 (2006).
Kawai, T. & Akira, S. Toll-like receptors and their crosstalk with other innate receptors in infection and immunity. Immunity 34, 637–650 (2011).
Kufer, T. A. & Sansonetti, P. J. NLR functions beyond pathogen recognition. Nature Immunol. 12, 121–128 (2011).
Loo, Y. M. & Gale, M. Jr. Immune signaling by RIG-I-like receptors. Immunity 34, 680–692 (2011).
Schroder, K. & Tschopp, J. The inflammasomes. Cell 140, 821–832 (2010).
Sancho, D. & Reis e Sousa, C. Signaling by myeloid C-type lectin receptors in immunity and homeostasis. Annu. Rev. Immunol. 30, 491–529 (2012).
Bottazzi, B., Doni, A., Garlanda, C. & Mantovani, A. An integrated view of humoral innate immunity: pentraxins as a paradigm. Annu. Rev. Immunol. 28, 157–183 (2010).
Casanova, J. L., Abel, L. & Quintana-Murci, L. Human TLRs and IL-1Rs in host defense: natural insights from evolutionary, epidemiological, and clinical genetics. Annu. Rev. Immunol. 29, 447–491 (2011).
Barreiro, L. B. et al. Evolutionary dynamics of human Toll-like receptors and their different contributions to host defense. PLoS Genet. 5, e1000562 (2009).
Mukherjee, S., Sarkar-Roy, N., Wagener, D. K. & Majumder, P. P. Signatures of natural selection are not uniform across genes of innate immune system, but purifying selection is the dominant signature. Proc. Natl Acad. Sci. USA 106, 7073–7078 (2009).
Nakajima, T. et al. Natural selection in the TLR-related genes in the course of primate evolution. Immunogenetics 60, 727–735 (2008).
Smirnova, I., Hamblin, M. T., McBride, C., Beutler, B. & Di Rienzo, A. Excess of rare amino acid polymorphisms in the Toll-like receptor 4 in humans. Genetics 158, 1657–1664 (2001).
Wlasiuk, G., Khan, S., Switzer, W. M. & Nachman, M. W. A history of recurrent positive selection at the toll-like receptor 5 in primates. Mol. Biol. Evol. 26, 937–949 (2009).
Wlasiuk, G. & Nachman, M. W. Adaptation and constraint at Toll-like receptors in primates. Mol. Biol. Evol. 27, 2172–2186 (2010).
Ferrer-Admetlla, A. et al. Balancing selection is the main force shaping the evolution of innate immunity genes. J. Immunol. 181, 1315–1322 (2008).
Ortiz, M. et al. The evolutionary history of the CD209 (DC-SIGN) family in humans and non-human primates. Genes Immun. 9, 483–492 (2008).
Ferwerda, B. et al. TLR4 polymorphisms, infectious diseases, and evolutionary pressure during migration of modern humans. Proc. Natl Acad. Sci. USA 104, 16645–16650 (2007).
Hawn, T. R. et al. A common dominant TLR5 stop codon polymorphism abolishes flagellin signaling and is associated with susceptibility to Legionnaires' disease. J. Exp. Med. 198, 1563–1572 (2003).
Franchi, L. et al. Cytosolic flagellin requires Ipaf for activation of caspase-1 and interleukin 1β in salmonella-infected macrophages. Nature Immunol. 7, 576–582 (2006).
Lohmueller, K. E. et al. Proportionally more deleterious genetic variation in European than in African populations. Nature 451, 994–997 (2008). A well-documented case that the out-of-Africa migration resulted in a bottleneck that increased the amount of deleterious mutations among non-African populations.
Kariko, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Deane, J. A. & Bolland, S. Nucleic acid-sensing TLRs as modifiers of autoimmunity. J. Immunol. 177, 6573–6578 (2006).
Marshak-Rothstein, A. Toll-like receptors in systemic autoimmune disease. Nature Rev. Immunol. 6, 823–835 (2006).
Pisitkun, P. et al. Autoreactive B cell responses to RNA-related antigens due to TLR7 gene duplication. Science 312, 1669–1672 (2006).
Krieg, A. M. & Vollmer, J. Toll-like receptors 7, 8, and 9: linking innate immunity to autoimmunity. Immunol. Rev. 220, 251–269 (2007).
Fumagalli, M. et al. Population genetics of IFIH1: ancient population structure, local selection and implications for susceptibility to type 1 diabetes. Mol. Biol. Evol. 27, 2555–2566 (2010).
Vasseur, E. et al. The evolutionary landscape of cytosolic microbial sensors in humans. Am. J. Hum. Genet. 91, 27–37 (2012).
Vasseur, E. et al. The selective footprints of viral pressures at the human RIG-I-like receptor family. Hum. Mol. Genet. 20, 4462–4474 (2011).
Fujita, T., Matsushita, M. & Endo, Y. The lectin-complement pathway — its role in innate immunity and evolution. Immunol. Rev. 198, 185–202 (2004).
Holmskov, U., Thiel, S. & Jensenius, J. C. Collections and ficolins: humoral lectins of the innate immune defense. Annu. Rev. Immunol. 21, 547–578 (2003).
Garred, P., Larsen, F., Seyfarth, J., Fujita, R. & Madsen, H. O. Mannose-binding lectin and its genetic variants. Genes Immun. 7, 85–94 (2006).
Casanova, J. L. & Abel, L. Human mannose-binding lectin in immunity: friend, foe, or both? J. Exp. Med. 199, 1295–1299 (2004).
Heitzeneder, S., Seidel, M., Forster-Waldl, E. & Heitger, A. Mannan-binding lectin deficiency — good news, bad news, doesn't matter? Clin. Immunol. 143, 22–38 (2012).
Eisen, D. P. & Minchinton, R. M. Impact of mannose-binding lectin on susceptibility to infectious diseases. Clin. Infect. Dis. 37, 1496–1505 (2003).
Bernig, T. et al. Sequence analysis of the mannose-binding lectin (MBL2) gene reveals a high degree of heterozygosity with evidence of selection. Genes Immun. 5, 461–476 (2004).
Verdu, P. et al. Evolutionary insights into the high worldwide prevalence of MBL2 deficiency alleles. Hum. Mol. Genet. 15, 2650–2658 (2006).
Hoal-Van Helden, E. G. et al. Mannose-binding protein B allele confers protection against tuberculous meningitis. Pediatr. Res. 45, 459–464 (1999).
Santos, I. K. et al. Mannan-binding lectin enhances susceptibility to visceral leishmaniasis. Infect. Immun. 69, 5212–5215 (2001).
Walsh, E. C. et al. Searching for signals of evolutionary selection in 168 genes related to immune function. Hum. Genet. 119, 92–102 (2006).
O'Neill, L. A. & Bowie, A. G. The family of five: TIR-domain-containing adaptors in Toll-like receptor signalling. Nature Rev. Immunol. 7, 353–364 (2007).
Fornarino, S. et al. Evolution of the TIR domain-containing adaptors in humans: swinging between constraint and adaptation. Mol. Biol. Evol. 28, 3087–3097 (2011).
Ferwerda, B. et al. Functional and genetic evidence that the Mal/TIRAP allele variant 180L has been selected by providing protection against septic shock. Proc. Natl Acad. Sci. USA 106, 10272–10277 (2009).
Casrouge, A. et al. Herpes simplex virus encephalitis in human UNC-93B deficiency. Science 314, 308–312 (2006).
Zhang, S. Y. et al. TLR3 deficiency in patients with herpes simplex encephalitis. Science 317, 1522–1527 (2007).
Zhang, S. Y. et al. Inborn errors of interferon (IFN)-mediated immunity in humans: insights into the respective roles of IFN-α/β, IFN-γ, and IFN-λ in host defense. Immunol. Rev. 226, 29–40 (2008).
Ohta, T. Gene conversion and evolution of gene families: an overview. Genes 1, 349–356 (2010).
Manry, J. et al. Evolutionary genetic dissection of human interferons. J. Exp. Med. 208, 2747–2759 (2011). The first population genetic study of human IFNs that demonstrated that the different IFN families, and individual IFN subtypes, differ widely in their biological relevance for host defence.
Manry, J. et al. Evolutionary genetics evidence of an essential, nonredundant role of the IFN-γ pathway in protective immunity. Hum. Mutat. 32, 633–642 (2011).
Klotman, M. E. & Chang, T. L. Defensins in innate antiviral immunity. Nature Rev. Immunol. 6, 447–456 (2006).
Abu Bakar, S., Hollox, E. J. & Armour, J. A. Allelic recombination between distinct genomic locations generates copy number diversity in human β-defensins. Proc. Natl Acad. Sci. USA 106, 853–858 (2009).
Aldred, P. M., Hollox, E. J. & Armour, J. A. Copy number polymorphism and expression level variation of the human α-defensin genes DEFA1 and DEFA3. Hum. Mol. Genet. 14, 2045–2052 (2005).
Hollox, E. J. & Armour, J. A. Directional and balancing selection in human β-defensins. BMC Evol. Biol. 8, 113 (2008).
Das, S. et al. Comparative genomics and evolution of the α-defensin multigene family in primates. Mol. Biol. Evol. 27, 2333–2343 (2010).
Cagliani, R. et al. The signature of long-standing balancing selection at the human defensin β-1 promoter. Genome Biol. 9, R143 (2008).
Chen, Q. X. et al. Genomic variations within DEFB1 are associated with the susceptibility to and the fatal outcome of severe sepsis in Chinese Han population. Genes Immun. 8, 439–443 (2007).
Hamblin, M. T., Thompson, E. E. & Di Rienzo, A. Complex signatures of natural selection at the Duffy blood group locus. Am. J. Hum. Genet. 70, 369–383 (2002).
Xue, Y. et al. Spread of an inactive form of caspase-12 in humans is due to recent positive selection. Am. J. Hum. Genet. 78, 659–670 (2006).
Isnardi, I. et al. IRAK-4- and MyD88-dependent pathways are essential for the removal of developing autoreactive B cells in humans. Immunity 29, 746–757 (2008).
von Bernuth, H. et al. Pyogenic bacterial infections in humans with MyD88 deficiency. Science 321, 691–696 (2008).
Fumagalli, M. et al. Signatures of environmental genetic adaptation pinpoint pathogens as the main selective pressure through human evolution. PLoS Genet. 7, e1002355 (2011). This study demonstrates that the selective pressures imposed by pathogens have been among the strongest influences on the patterns of diversity of the human genome.
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Round, J. L. & Mazmanian, S. K. The gut microbiota shapes intestinal immune responses during health and disease. Nature Rev. Immunol. 9, 313–323 (2009).
Eberl, G. A new vision of immunity: homeostasis of the superorganism. Mucosal Immunol. 3, 450–460 (2010).
Rakoff-Nahoum, S., Paglino, J., Eslami-Varzaneh, F., Edberg, S. & Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118, 229–241 (2004).
Bouskra, D. et al. Lymphoid tissue genesis induced by commensals through NOD1 regulates intestinal homeostasis. Nature 456, 507–510 (2008).
Ma, Y. et al. Toll-like receptor 8 functions as a negative regulator of neurite outgrowth and inducer of neuronal apoptosis. J. Cell Biol. 175, 209–215 (2006).
Strowig, T., Henao-Mejia, J., Elinav, E. & Flavell, R. Inflammasomes in health and disease. Nature 481, 278–286 (2012).
Jin, Y. et al. Genome-wide association analyses identify 13 new susceptibility loci for generalized vitiligo. Nature Genet. 44, 676–680 (2012).
Li, Y. et al. Carriers of rare missense variants in IFIH1 are protected from psoriasis. J. Invest. Dermatol. 130, 2768–2772 (2010).
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J. A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009).
Di Rienzo, A. Population genetics models of common diseases. Curr. Opin. Genet. Dev. 16, 630–636 (2006).
Blekhman, R. et al. Natural selection on genes that underlie human disease susceptibility. Curr. Biol. 18, 883–889 (2008).
Nielsen, R. et al. Darwinian and demographic forces affecting human protein coding genes. Genome Res. 19, 838–849 (2009).
Picard, C. et al. Pyogenic bacterial infections in humans with IRAK-4 deficiency. Science 299, 2076–2079 (2003).
Aganna, E. et al. Association of mutations in the NALP3/CIAS1/PYPAF1 gene with a broad phenotype including recurrent fever, cold sensitivity, sensorineural deafness, and AA amyloidosis. Arthritis Rheum. 46, 2445–2452 (2002).
Hoffman, H. M., Mueller, J. L., Broide, D. H., Wanderer, A. A. & Kolodner, R. D. Mutation of a new gene encoding a putative pyrin-like protein causes familial cold autoinflammatory syndrome and Muckle-Wells syndrome. Nature Genet. 29, 301–305 (2001).
Filipe-Santos, O. et al. Inborn errors of IL-12/23- and IFN-γ-mediated immunity: molecular, cellular, and clinical features. Semin. Immunol. 18, 347–361 (2006).
Alcais, A. et al. Life-threatening infectious diseases of childhood: single-gene inborn errors of immunity? Ann. NY Acad. Sci. 1214, 18–33 (2010). An elaborated view of the genetic architecture of infectious diseases, whereby life-threatening infectious diseases in childhood result mostly from rare single-gene variations of large effect, whereas the genetic component of predisposition to secondary infections in adults is genetically more complex.
Peterslund, N. A., Koch, C., Jensenius, J. C. & Thiel, S. Association between deficiency of mannose-binding lectin and severe infections after chemotherapy. Lancet 358, 637–638 (2001).
Torgerson, D. G. et al. Evolutionary processes acting on candidate cis-regulatory regions in humans inferred from patterns of polymorphism and divergence. PLoS Genet. 5, e1000592 (2009).
Jin, Y. et al. NALP1 in vitiligo-associated multiple autoimmune disease. N. Engl. J. Med. 356, 1216–1225 (2007).
Johnson, C. M. et al. Cutting edge: a common polymorphism impairs cell surface trafficking and functional responses of TLR1 but protects against leprosy. J. Immunol. 178, 7520–7524 (2007).
Ma, X. et al. Full-exon resequencing reveals Toll-like receptor variants contribute to human susceptibility to tuberculosis disease. PLoS ONE 2, e1318 (2007).
Schuring, R. P. et al. Polymorphism N248S in the human Toll-like receptor 1 gene is related to leprosy and leprosy reactions. J. Infect. Dis. 199, 1816–1819 (2009).
Swanberg, M. et al. MHC2TA is associated with differential MHC molecule expression and susceptibility to rheumatoid arthritis, multiple sclerosis and myocardial infarction. Nature Genet. 37, 486–494 (2005).
Prokunina-Olsson, L. et al. A variant upstream of IFNL3 (IL28B) creating a new interferon gene IFNL4 is associated with impaired clearance of hepatitis C virus. Nature Genet. 45, 164–171 (2013).
Ge, D. et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature 461, 399–401 (2009).
Suppiah, V. et al. IL28B is associated with response to chronic hepatitis C interferon-α and ribavirin therapy. Nature Genet. 41, 1100–1104 (2009).
Tanaka, Y. et al. Genome-wide association of IL28B with response to pegylated interferon-α and ribavirin therapy for chronic hepatitis C. Nature Genet. 41, 1105–1109 (2009).
Thomas, D. L. et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature 461, 798–801 (2009). References 123–127 report one of the most convincing genome-wide associations of a host genetic factor with complex susceptibility to infection and response to treatment.
Sironi, M. & Clerici, M. The hygiene hypothesis: an evolutionary perspective. Microbes Infect. 12, 421–427 (2010).
Magitta, N. F. et al. A coding polymorphism in NALP1 confers risk for autoimmune Addison's disease and type 1 diabetes. Genes Immun. 10, 120–124 (2009).
Leung, T. F. et al. Asthma and atopy are associated with DEFB1 polymorphisms in Chinese children. Genes Immun. 7, 59–64 (2006).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012). The most comprehensive analysis of the genetic architecture of IBD. The study showed an extensive overlap between IBD loci and loci that are linked to other disorders, such as ankylosing spondylitis, psoriasis and mycobacterial infections.
Cho, J. H. The genetics and immunopathogenesis of inflammatory bowel disease. Nature Rev. Immunol. 8, 458–466 (2008).
Nakagome, S. et al. Crohn's disease risk alleles on the NOD2 locus have been maintained by natural selection on standing variation. Mol. Biol. Evol. 29, 1569–1585 (2012).
Pritchard, J. K., Pickrell, J. K. & Coop, G. The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation. Curr. Biol. 20, R208–R215 (2010).
Casals, F. et al. Genetic adaptation of the antibacterial human innate immunity network. BMC Evol. Biol. 11, 202 (2011).
Consortium, T. E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kudaravalli, S., Veyrieras, J. B., Stranger, B. E., Dermitzakis, E. T. & Pritchard, J. K. Gene expression levels are a target of recent natural selection in the human genome. Mol. Biol. Evol. 26, 649–658 (2009).
Schaub, M. A., Boyle, A. P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759 (2012).
Vernot, B. et al. Personal and population genomics of human regulatory variation. Genome Res. 22, 1689–1697 (2012).
Landolt, G. A. & Olsen, C. W. Up to new tricks — a review of cross-species transmission of influenza A viruses. Anim. Health Res. Rev. 8, 1–21 (2007).
Greenbaum, B. D., Levine, A. J., Bhanot, G. & Rabadan, R. Patterns of evolution and host gene mimicry in influenza and other RNA viruses. PLoS Pathog. 4, e1000079 (2008).
Rabadan, R., Levine, A. J. & Robins, H. Comparison of avian and human influenza A viruses reveals a mutational bias on the viral genomes. J. Virol. 80, 11887–11891 (2006).
Takeshita, F. et al. Signal transduction pathways mediated by the interaction of CpG DNA with Toll-like receptor 9. Semin. Immunol. 16, 17–22 (2004).
Kobasa, D. et al. Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 445, 319–323 (2007).
Huang, K. J. et al. An interferon-γ-related cytokine storm in SARS patients. J. Med. Virol. 75, 185–194 (2005).
Haque, A., Hober, D. & Kasper, L. H. Confronting potential influenza A (H5N1) pandemic with better vaccines. Emerg. Infect. Dis. 13, 1512–1518 (2007).
Ferrandon, D., Imler, J. L., Hetru, C. & Hoffmann, J. A. The Drosophila systemic immune response: sensing and signalling during bacterial and fungal infections. Nature Rev. Immunol. 7, 862–874 (2007).
Jiggins, F. M. & Kim, K. W. Contrasting evolutionary patterns in Drosophila immune receptors. J. Mol. Biol. 63, 769–780 (2006).
Sackton, T. B. et al. Dynamic evolution of the innate immune system in Drosophila. Nature Genet. 39, 1461–1468 (2007).
Lazzaro, B. P. Elevated polymorphism and divergence in the class C scavenger receptors of Drosophila melanogaster and D. simulans. Genetics 169, 2023–2034 (2005).
Schlenke, T. A. & Begun, D. J. Natural selection drives Drosophila immune system evolution. Genetics 164, 1471–1480 (2003).
Waterhouse, R. M. et al. Evolutionary dynamics of immune-related genes and pathways in disease-vector mosquitoes. Science 316, 1738–1743 (2007).
Begun, D. J. & Whitley, P. Adaptive evolution of Relish, a Drosophila NF-κB/IκB protein. Genetics 154, 1231–1238 (2000).
Jiggins, F. M. & Kim, K. W. The evolution of antifungal peptides in Drosophila. Genetics 171, 1847–1859 (2005).
Lazzaro, B. P. & Clark, A. G. Molecular population genetics of inducible antibacterial peptide genes in Drosophila melanogaster. Mol. Biol. Evol. 20, 914–923 (2003).
Candille, S. I. et al. A β-defensin mutation causes black coat color in domestic dogs. Science 318, 1418–1423 (2007).
Hawn, T. R. et al. A common human TLR1 polymorphism regulates the innate immune response to lipopeptides. Eur. J. Immunol. 37, 2280–2289 (2007).
Misch, E. A. et al. Human TLR1 deficiency is associated with impaired mycobacterial signaling and protection from leprosy reversal reaction. PLoS Negl. Trop. Dis. 2, e231 (2008).
Enard, D., Depaulis, F. & Roest Crollius, H. Human and non-human primate genomes share hotspots of positive selection. PLoS Genet. 6, e1000840 (2010).
Acknowledgements
We would like to thank M. Albert, L. Barreiro, J.-L. Casanova, G. Eberl, E. Patin and J. Pothlichet for helpful comments and discussions. This work was supported by Institut Pasteur, the Centre National de la Recherche Scientifique, the Agence Nationale de la Recherche (ANR-08-MIEN-009-01), the Fondation pour la Recherche Médicale, and an École Polytechnique Federale de Lausanne-Debiopharm Life Sciences Award to L.Q.-M. The laboratory of L.Q.-M. has received funding from the French Government's Investissement d'Avenir program, Laboratoire d'Excellence “Integrative Biology of Emerging Infectious Diseases” (grant no. ANR-10-LABX-62-IBEID), and from the European Research Council under the European Union's Seventh Framework Programme (FP/2007–2013)/ERC Grant Agreement No. 281297. A.G.C. receives funding from the US National Institutes of Health (R01 AI064950).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Next-generation sequencing
-
Non-Sanger-based, high-throughput methods that are able to produce thousands or millions of sequence reads at once. Next-generation sequencing can be used for the direct sequencing of genomes and transcripts, but also to learn more about genome-wide variation of regulatory mechanisms: for example, variation in transcription factor binding sites or epigenetic modifications such as DNA methylation.
- Whole-exome sequencing
-
A technology that involves capturing the exonic portion of the genome (roughly 2–3% of the human genome) using microarrays and then applying next-generation sequencing. This approach remains more affordable than whole-genome sequencing and retains the most likely sources of genetic disease risk. This technology is increasingly being used in medical genetics studies.
- Genome-wide association studies
-
(GWAS). Unbiased genome-wide screens in which associations between genetic variation and a phenotype or trait of interest are identified by genotyping cases (for example, diseased individuals) and controls (for example, healthy individuals). The dominant technology used so far has been genome-wide single-nucleotide polymorphism arrays.
- Fitness
-
A measure of the capacity of an organism to survive and reproduce.
- Synonymous mutations
-
Substitutions of one nucleotide for another in the DNA sequence of an exon that do not alter the corresponding amino acid sequence. Synonymous mutations that occur outside protein-coding genes are broadly known as silent mutations. Synonymous and silent mutations are often assumed to be neutral.
- Non-synonymous mutations
-
Nucleotide substitutions in an exon that, in contrast to synonymous changes, alter the amino acid sequence of a protein. Depending on how radical the amino acid change is, the impact of a non-synonymous mutation on protein function is variable and subject to natural selection to different extents.
- Single-nucleotide polymorphisms
-
(SNPs). Bi-allelic (typically) base-pair substitutions, which are the most common forms of genetic polymorphism.
- International HapMap Project
-
The International HapMap Project has built a haplotype map of the human genome and reports the common patterns of human genetic variation based on the results of genotyping analyses. This freely available data set reports information on the allelic frequencies of up to 3 million single-nucleotide polymorphisms distributed throughout the genome, across different human populations.
- 1000 Genomes Project
-
The 1000 Genomes Project aims to provide a large number of complete human genome sequences from individuals from different ethnic backgrounds. The advantage of this project is that it provides information on all forms of DNA polymorphism as well as on low-frequency and rare variants, which are absent in the HapMap Project.
- Complement receptors
-
The complement system is a family of serum proteins and cell-surface receptors that participate in innate and adaptive immunity, and is one of the main effector mechanisms of antibody-mediated immunity. They act in concert to mediate inflammation, enhance B and T cell immunity, and regulate self-reactive B cells.
- Ficolins
-
A group of humoral proteins that contain a collagen-like domain and a fibrinogen-like domain. They can bind carbohydrate molecules on pathogens, apoptotic and necrotic cells to activate the lectin–complement pathway.
- Collectins
-
C-type lectins that have a collagen-like domain. One group of collectins, the secreted lectins, consists of mannose-binding lectin (MBL), bovine conglutinin (BKg) and collectin 43 (CL43) in blood, and the two mucosal-associated proteins surfactant protein A (SPA) and SPD. The other group of collectins consists of the newly discovered non-secreted-type collectin liver 1 (CL-L1) and membrane-type collectin placenta 1 (CL-P1).
- Pentraxins
-
Pentraxins constitute a superfamily of evolutionarily conserved proteins characterized by a cyclic multimeric structure and the presence in the carboxyl terminus of a pentraxin domain. They are prototypic components of the humoral arm of innate immunity.
- Lectin–complement pathway
-
The lectin–complement pathway involves carbohydrate recognition by pattern-recognition receptors, such as mannose binding lectin (MBL) and ficolins, and the subsequent activation of associated unique enzymes that are known as MBL-associated serine proteases (MASPs). Other complement pathways include the classical-complement pathway and the alternative-complement pathway.
- Defensins
-
A class of antimicrobial peptides that have activity against Gram-positive and Gram-negative bacteria, fungi and viruses. Defensins are classified into two main categories on the basis of the position of conserved cysteine and hydrophobic residues and the linkages of disulphide bonds: α-defensins are produced by intestinal Paneth cells and neutrophils, and β-defensins are expressed by most epithelial cells. A third category, the θ-defensins, arises from the splicing of two α-defensin-related peptides into a circular molecule; at present, these defensins have been detected only in the neutrophils of rhesus macaques.
- Genetic drift
-
The random fluctuations in allele frequencies over time that are due to chance alone.
- DEATH domain
-
A protein domain that is found in many proteins that are involved in cellular signalling processes, including apoptosis, inflammation and development. This domain mediates protein–protein interactions.
- Expression quantitative trait loci
-
(eQTLs). Genomic loci in which genetic variants alter individual differences in quantitative levels of gene expression.
- ENCODE Consortium
-
The goal of the Encyclopedia of DNA Elements (ENCODE) Consortium is to systematically map regions of transcription, transcription factor association, chromatin structure and histone modification. In doing so, it provides new insights into the organization and regulation of the human genome and constitutes an expansive resource of functional annotations for biomedical research.
Rights and permissions
About this article
Cite this article
Quintana-Murci, L., Clark, A. Population genetic tools for dissecting innate immunity in humans. Nat Rev Immunol 13, 280–293 (2013). https://doi.org/10.1038/nri3421
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri3421
This article is cited by
-
Non-MHC immunity genes do not affect parasite load in European invasive populations of common raccoon
Scientific Reports (2023)
-
Regulatory regions in natural transposable element insertions drive interindividual differences in response to immune challenges in Drosophila
Genome Biology (2021)
-
Inherited GATA2 Deficiency Is Dominant by Haploinsufficiency and Displays Incomplete Clinical Penetrance
Journal of Clinical Immunology (2021)
-
The human genetic determinism of life-threatening infectious diseases: genetic heterogeneity and physiological homogeneity?
Human Genetics (2020)
-
Evolutionary and population (epi)genetics of immunity to infection
Human Genetics (2020)