Abstract
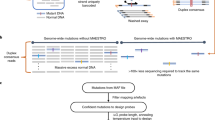
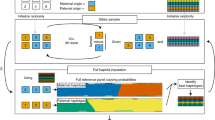
The goal of DNA sequencing and genotyping is to efficiently generate accurate high-throughput digital genetic information that unambiguously identifies sources of genetic variation and clearly distinguishes heterozygous from homozygous variants. Recent advances in mass-spectrometry-based DNA sequencing and genotyping bode well for meeting these criteria. Pilot studies show that these recently developed approaches allow unambiguous multiplex detection of heterozygous variants and the identification of deletion and insertion variants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Collins, F. S., Green, E. D., Guttmacher, A. E. & Guyer, M. S. A vision for the future of genomics research. Nature 422, 835–847 (2003).
Smith, L. M. et al. Fluorescence detection in automated DNA sequence analysis. Nature 321, 674–679 (1986).
Ju, J., Ruan, C., Fuller, C. W., Glazer, A. N. & Mathies, R. A. Fluorescence energy transfer dye-labeled primers for DNA sequencing and analysis. Proc. Natl Acad. Sci. USA 92, 4347–4351 (1995).
Ju, J., Glazer, A. N. & Mathies, R. A. Cassette labeling for facile construction of energy transfer fluorescent primers. Nucleic Acids Res. 24, 1144–1148 (1996).
Kheterpal, I. et al. DNA sequencing using a four-color confocal fluorescence capillary array scanner. Electrophoresis 17, 1852–1859 (1996).
Bowling, J. M., Bruner, K. L., Cmarik, J. L. & Tibbetts, C. Neighboring nucleotide interactions during DNA sequencing gel electrophoresis. Nucleic Acids Res. 19, 3089–3097 (1991).
Yamakawa, H. & Ohara, O. A DNA cycle sequencing reaction that minimizes compressions on automated fluorescent sequencers. Nucleic Acids Res. 25, 1311–1312 (1997).
Fitzgerald, M. C., Zhu, L. & Smith, L. M. The analysis of mock DNA sequencing reactions using matrix-assisted laser desorption/ionization mass spectrometry. Rapid Commun. Mass Spectrom. 7, 895–897 (1993).
Roskey, M. T. et al. DNA sequencing by delayed extraction-matrix-assisted laser desorption/ionization time of flight mass spectrometry. Proc. Natl Acad. Sci. USA. 93, 4724–4729 (1996).
Kirpekar, F. et al. DNA sequence analysis by MALDI mass spectrometry. Nucleic Acid Res. 26, 2554–2559 (1998).
Monforte, J. & Becker, C. High-throughput DNA analysis by time-of-flight mass spectrometry. Nature Med. 3, 360–362 (1997).
Fu, D. J. et al. Sequencing exons 5 to 8 of the p53 gene by MALDI-TOF mass spectrometry. Nature Biotechnol. 16, 381–384 (1998).
Sanger, F., Nicklen, S. & Coulson, A. R. DNA sequencing with chain-terminating inhibitors. Proc. Natl Acad. Sci. USA 74, 5463–5467 (1977).
Langer, P. R., Waldrop, A. A. & Ward, D. C. Enzymatic synthesis of biotin-labeled polynucleotides: novel nucleic acid affinity probes. Proc. Natl Acad. Sci. USA. 78, 6633–6637 (1981).
Hawkins, T. L., O'Connor-Morin, T., Roy, A. & Santillan, C. DNA purification and isolation using a solid-phase. Nucleic Acids Res. 22, 4543–4544 (1994).
Uhlen, M. Magnetic separation of DNA. Nature 340, 733–734 (1989).
Ju, J. DNA sequencing with solid phase capturable dideoxynucleotides and energy transfer primers. Anal. Biochem. 309, 35–39 (2002).
Edwards, J. R., Itagaki, Y. & Ju, J. DNA sequencing using biotinylated dideoxynucleotides and mass spectrometry. Nucleic Acids Res. 29, 1–5 (2001).
Prober, J. et al. A system for rapid DNA sequencing with fluorescent chain-terminating dideoxynucleotides. Science 238, 336–341 (1987).
Gebeyehu, G., Rao, P., SooChan, P., Simms, D. A. & Klevan, L. Novel biotinylated nucleotide analogs for labeling and calorimetric detection of DNA. Nucleic Acid Res. 15, 4513–4534 (1987).
Herman, T., Lefever, E. & Shimkus, M. Affinity chromatography of DNA labeled with chemically cleavable biotinylated nucleotide analogs. Anal. Biochem. 156, 48–55 (1986).
Ronaghi, M., Uhlén, M. & Nyrén, P. A sequencing method based on real-time pyrophosphate. Science 281, 363–365 (1998).
Fakhrai-Rad, H., Pourmand, N. & Ronaghi, M. Pyrosequencing™: an accurate detection platform for single nucleotide polymorphisms. Hum. Mutat. 19, 479–485 (2002).
Ruparel, H., Ulz, M. E., Kim, S. & Ju, J. Digital detection of genetic mutations using SPC–sequencing. Genome Res. (in the press).
Friedman, L. S. et al. Confirmation of BRCA1 by analysis of germline mutations linked to breast and ovarian cancer in ten families. Nature Genet. 8, 399–404 (1994).
Steele, R. J. C., Thompson, A. M., Hall, P. A. & Lane, D. P. The p53 tumour suppressor gene. Br. J. Surgery 85, 1460–1467 (1998).
Kwok, P -Y. High-throughput genotyping assay approaches. Pharmacogenomics 1, 95–100 (2000).
Roses, A. Pharmacogenetics and the practice of medicine. Nature 405, 857–865 (2000).
The International SNP Map Working Group. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001).
Beavis, R. C. & Chait, B. T. Matrix-assisted laser-desorption mass spectrometry using 355 nm radiation. Rapid Commun. Mass Spectrom. 3, 436–439 (1989).
Stoerker, J., Mayo, J. D., Tetzlaff, C. N., Sarracino, D. A., Schwope, I. & Richert, C. Rapid genotyping by MALDI-monitored nuclease selection from probe libraries. Nature Biotechnol. 18, 1213–1216 (2000).
Ross, P. L., Lee, K. & Belgrader, P. Discrimination of single-nucleotide polymorphisms in human DNA using peptide nucleic acid probes detected by MALDI-TOF mass spectrometry. Anal. Chem. 69, 4197–4202 (1997).
Griffin, T. J., Hall, J. G., Prudent, J. R. & Smith, L. M. Direct genetic analysis by matrix-assisted laser desorption/ionization mass spectrometry. Proc. Natl Acad. Sci. USA, 96, 6301–6306 (1999).
Lyamichev, V. et al. Polymorphism identification and quantitative detection of genomic DNA by invasive cleavage of oligonucleotide probes. Nature Biotechnol. 17, 292–296 (1999).
Haff, L. A. & Smirnov, I. P. Multiplex genotyping of PCR products with mass tag-labeled primers. Nucleic Acids Res. 25, 3749–3750 (1997).
Tang, K. et al. Chip-based genotyping by mass spectrometry. Proc. Natl Acad. Sci. USA 96, 10016–10020 (1999).
Ross, P., Hall, L., Smirnov, I. P. & Haff, L. High level multiplex genotyping by MALDI-TOF mass spectrometry. Nature Biotechnol. 16, 1347–1351 (1998).
Fei, Z., Ono, T. & Smith, L. M. MALDI-TOF mass spectrometric typing of single nucleotide polymorphisms with mass-tagged ddNTPs. Nucleic Acids Res. 26, 2827–2828 (1998).
Griffin, T. J. & Smith, L. M. Single-nucleotide polymorphism analysis by MALDI-TOF mass spectrometry. Trends. Biotechnol. 18, 77–84 (2000).
Ding, C. & Cantor, C. R. A high-throughput gene expression analysis technique using competitive PCR and matrix-assisted laser desorption ionization time-of-flight MS. Proc. Natl Acad. Sci. USA, 100, 3059–3064 (2003).
Ding, C. & Cantor, C. R. Direct molecular haplotyping of long-range genomic DNA with M1-PCR. Proc. Natl Acad. Sci. USA 100, 7449–7453 (2003).
Kim, S., Edwards, J. R., Deng, L., Chung, W. & Ju, J. Solid phase capturable dideoxynucleotides for multiplex genotyping using mass spectrometry. Nucleic Acids Res. 30, 1–6 (2002).
Kim, S. et al. Multiplex genotyping of the human β2-adrenergic receptor gene using solid phase capturable dideoxynucleotides and mass spectrometry. Anal. Biochem. 316, 251–258 (2003).
Hanson, E. H., Imperatore, G. & Burke, W. HFE gene and hereditary hemochromatosis: A HuGE review. Am. J. Epidem. 154, 193–206 (2001).
Hollstein, M., Sidransky, D., Vogelstein, B. & Harris, C. C. p53 mutations in human cancers. Science 253, 49–53 (1991).
Bardelli, A. et al. Mutational analysis of the tyrosine kinome in colorectal cancers. Science 300, 949 (2003).
Kim, S. et al. Thirty fold multiplex genotyping of the p53 gene using solid phase capturable dideoxynucleotides and mass spectrometry Genomics (in the press).
Malkin, D., Sexsmith, E., Yeger, H., Williams, B. R. G. & Coppes, M. J. Mutations of the p53 tumor suppressor gene occur infrequently in Wilms' tumor. Cancer Res. 54, 2077–2079 (1994).
Lahoti, C., Thorner, P., Malkin, D. & Yeger, H. Immunohistochemical detection of p53 in Wilms' tumors correlates with unfavorable outcome. Am. J. Pathol. 148, 1577–1589 (1996).
Min, B. M. et al. Inactivation of the p53 gene by either mutation or HPV infection is extremely frequent in human oral squamous cell carcinoma cell lines. Oral Oncol. Eur. J. Cancer Part B 30, 338–345 (1994).
Popanda, O. et al. Mutation analysis of replicative genes encoding the large subunits of DNA polymerase-α and replication factors A and C in human sporadic colorectal cancers. Int. J. Cancer 86, 318–324 (2000).
Lin-Lee, Y. C., Tatebe, S., Savaraj, N., Ishikawa, T. & Kuo, M. T. Differential sensitivities of the MRP gene family and γ-glutamylcysteine synthetase to prooxidants in human colorectal carcinoma cell lines with different p53 status. Biochem. Pharmacol. 61, 555–563 (2001).
Rodi, C. P., Darnhofer-Patel, B., Stanssens, P., Zabeau, M. and van den Boom, D. A strategy for the rapid discovery of disease markers using the massARRAY™ system. Biotechniques 32 (Suppl.), 62–69 (2002).
Chen, X., Levine, L. & Kwok, P -U. Fluorescence polarization in homogeneous nucleic acid analysis. Genome Res. 9, 492–498 (1999).
Hardenbol, P. et al. Multiplexed genotyping with sequence-tagged molecular inversion probes. Nature Biotechnol. 21, 673–678 (2003).
Dressman, D., Yan, H., Traverso, G., Kinzler, K. W. & Vogelstein, B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variation. Proc. Natl Acad. Sci. USA 100, 8817–8822 (2003).
Acknowledgements
This work was supported by a Packard Fellowship for Science and Engineering (J.J.) and the Columbia University Genomics Initiative.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Related links
Glossary
- ANALYTE
-
A molecule that is of interest for analysis in a particular study.
- COMPRESSION
-
A phenomenon in gel electrophoresis that occurs as a result of the thermodynamic stability of hairpin formations in DNA sequences that terminate in a string of G and C nucleotides.
- FLOW CYTOMETRY
-
The analysis of single cells or subcellular particles by the detection of their light absorption, scattering and/or fluorescence properties as they pass through a laser beam in a directed fluid stream.
- INVASIVE CLEAVAGE
-
The excision of redundant portions of DNA by DNA repair enzymes such as 5′ to 3′ exonuclease or 5′ nuclease. These redundancies are caused when two oligonucleotides, which are hybridized to the same DNA template, overlap along some of their terminal bases. The cleavage of the overlapped portion causes the formation of a nick at the position of redundancy that can later be repaired by ligation.
- LASER-INDUCED FLUORESCENCE DETECTION
-
The measurement of emitted fluorescence signals from molecules that are excited by laser radiation.
- PHARMACOGENOMICS
-
The study of the influence of genetic differences on the variability in the response of individuals to drugs.
- PHLEBOTOMY
-
The removal of blood from a vein for diagnostic therapeutic purposes.
- SANGER DNA SEQUENCING
-
A sequencing method that involves the enzymatic synthesis of DNA chains of different length using dideoxynucleotides, the separation of the DNA fragments by size and the identification of the fragments to reveal the sequences.
- TAG SNPS
-
A small subset of SNPs that is needed to uniquely identify a complete haplotype.
Rights and permissions
About this article
Cite this article
Kim, S., Ruparel, H., Gilliam, T. et al. Digital genotyping using molecular affinity and mass spectrometry. Nat Rev Genet 4, 1001–1008 (2003). https://doi.org/10.1038/nrg1230
Issue Date:
DOI: https://doi.org/10.1038/nrg1230
This article is cited by
-
Identification of two functional markers associated with drought resistance in maize
Molecular Breeding (2015)
-
A causal C–A mutation in the second exon of GS3 highly associated with rice grain length and validated as a functional marker
Theoretical and Applied Genetics (2009)