Abstract
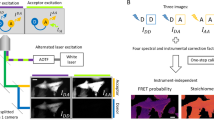
Genetically encoded fluorescent protein (FP)-based biosensor probes are useful tools for monitoring cellular events in living cells and tissues. Because these probes were developed for one-photon excitation approaches, their broad two-photon excitation (2PE) and poorly understood photobleaching characteristics have made their implementation in studies using two-photon laser-scanning microscopy (TPLSM) challenging. Here we describe a protocol that simplifies the use of Förster resonance energy transfer (FRET)-based biosensors in TPLSM. First, the TPLSM system is evaluated and optimized using FRET standards expressed in living cells, which enables the determination of spectral bleed-through (SBT) and the confirmation of FRET measurements from the known standards. Next, we describe how to apply the approach experimentally using a modified version of the A kinase activity reporter (AKAR) protein kinase A (PKA) biosensor as an example—first in cells in culture and then in hepatocytes in the liver of living mice. The microscopic imaging can be accomplished in a day in laboratories that routinely use TPLSM.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Day, R.N. & Davidson, M.W. The Fluorescent Protein Revolution (ed. Periasamy, A.) (Taylor & Francis, Boca Raton, FL, 2014).
Zhang, J., Campbell, R.E., Ting, A.Y. & Tsien, R.Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 3, 906–918 (2002).
VanEngelenburg, S.B. & Palmer, A.E. Fluorescent biosensors of protein function. Curr. Opin. Chem. Biol. 12, 60–65 (2008).
DiPilato, L.M. & Zhang, J. Fluorescent protein-based biosensors: resolving spatiotemporal dynamics of signaling. Curr. Opin. Chem. Biol. 14, 37–42 (2010).
Miyawaki, A. Development of probes for cellular functions using fluorescent proteins and fluorescence resonance energy transfer. Annu. Rev. Biochem. 80, 357–373 (2011).
Zhang, J., Ma, Y., Taylor, S.S. & Tsien, R.Y. Genetically encoded reporters of protein kinase A activity reveal impact of substrate tethering. Proc. Natl. Acad. Sci. USA 98, 14997–15002 (2001).
Patterson, G.H. & Piston, D.W. Photobleaching in two-photon excitation microscopy. Biophys. J. 78, 2159–2162 (2000).
Chen, T.S., Zeng, S.Q., Luo, Q.M., Zhang, Z.H. & Zhou, W. High-order photobleaching of green fluorescent protein inside live cells in two-photon excitation microscopy. Biochem. Biophys. Res. Commun. 291, 1272–1275 (2002).
Liu, L., Yermolaieva, O., Johnson, W.A., Abboud, F.M. & Welsh, M.J. Identification and function of thermosensory neurons in Drosophila larvae. Nat. Neurosci. 6, 267–273 (2003).
Ben Arous, J., Tanizawa, Y., Rabinowitch, I., Chatenay, D. & Schafer, W.R. Automated imaging of neuronal activity in freely behaving Caenorhabditis elegans. J. Neurosci. Methods 187, 229–234 (2010).
Kardash, E., Bandemer, J. & Raz, E. Imaging protein activity in live embryos using fluorescence resonance energy transfer biosensors. Nat. Protoc. 6, 1835–1846 (2011).
Hara, M. et al. Imaging endoplasmic reticulum calcium with a fluorescent biosensor in transgenic mice. Am. J. Physiol. Cell Physiol. 287, C932–C938 (2004).
Kamioka, Y. et al. Live imaging of protein kinase activities in transgenic mice expressing FRET biosensors. Cell Struct. Funct. 37, 65–73 (2012).
Oldach, L. & Zhang, J. Genetically encoded fluorescent biosensors for live-cell visualization of protein phosphorylation. Chem. Biol. 21, 186–197 (2014).
Aoki, K., Kamioka, Y. & Matsuda, M. Fluorescence resonance energy transfer imaging of cell signaling from in vitro to in vivo: basis of biosensor construction, live imaging, and image processing. Dev. Growth Differ. 55, 515–522 (2013).
Berglund, K. et al. Imaging synaptic inhibition in transgenic mice expressing the chloride indicator, Clomeleon. Brain Cell Biol. 35, 207–228 (2006).
Haustein, M.D. et al. Conditions and constraints for astrocyte calcium signaling in the hippocampal mossy fiber pathway. Neuron 82, 413–429 (2014).
Madisen, L. et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 85, 942–958 (2015).
Johnsson, A.K. et al. The Rac-FRET mouse reveals tight spatiotemporal control of Rac activity in primary cells and tissues. Cell Rep. 6, 1153–1164 (2014).
Buchholz, C.J., Friedel, T. & Buning, H. Surface-engineered viral vectors for selective and cell type-specific gene delivery. Trends Biotechnol. 33, 777–790 (2015).
Babbey, C.M. et al. Quantitative intravital microscopy of hepatic transport. Intravital 1, 44–53 (2012).
Dunn, K.W. et al. Functional studies of the kidney of living animals using multicolor two-photon microscopy. Am. J. Physiol. Cell Physiol. 283, C905–C916 (2002).
Dunn, K.W., Sutton, T.A. & Sandoval, R.M. Live-animal imaging of renal function by multiphoton microscopy. Curr. Protoc. Cytom. 14, 12.9 (2007).
Ryan, J.C., Dunn, K.W. & Decker, B.S. Effects of chronic kidney disease on liver transport: quantitative intravital microscopy of fluorescein transport in the rat liver. Am. J. Physiol. Regul. Integr. Comp. Physiol. 307, R1488–R1492 (2014).
Presson, R.G. Jr. et al. Two-photon imaging within the murine thorax without respiratory and cardiac motion artifact. Am. J. Pathol. 179, 75–82 (2011).
Vinegoni, C. et al. Sequential average segmented microscopy for high signal-to-noise ratio motion-artifact-free in vivo heart imaging. Biomed. Opt. Express 4, 2095–2106 (2013).
Dombeck, D.A., Khabbaz, A.N., Collman, F., Adelman, T.L. & Tank, D.W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Dunn, K.W., Lorenz, K.S., Salama, P. & Delp, E.J. IMART software for correction of motion artifacts in images collected in intravital microscopy. Intravital 3, e28210 (2014).
Lee, S., Vinegoni, C., Sebas, M. & Weissleder, R. Automated motion artifact removal for intravital microscopy, without a priori information. Sci. Rep. 4, 4507 (2014).
Lorenz, K.S., Salama, P., Dunn, K.W. & Delp, E.J. Digital correction of motion artefacts in microscopy image sequences collected from living animals using rigid and nonrigid registration. J. Microsc. 245, 148–160 (2012).
Soulet, D., Pare, A., Coste, J. & Lacroix, S. Automated filtering of intrinsic movement artifacts during two-photon intravital microscopy. PLoS One 8, e53942 (2013).
Radbruch, H. et al. Intravital FRET: probing cellular and tissue function. Int. J. Mol. Sci. 16, 11713–11727 (2015).
Tao, W. et al. A practical method for monitoring FRET-based biosensors in living animals using two-photon microscopy. Am. J. Physiol. Cell Physiol. 309, C724–C735 (2015).
Shaner, N.C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat. Methods 5, 545–551 (2008).
Goedhart, J. et al. Bright cyan fluorescent protein variants identified by fluorescence lifetime screening. Nat. Methods 7, 137–139 (2010).
Goedhart, J. et al. Structure-guided evolution of cyan fluorescent proteins towards a quantum yield of 93%. Nat. Commun. 3, 751 (2012).
Markwardt, M.L. et al. An improved cerulean fluorescent protein with enhanced brightness and reduced reversible photoswitching. PLoS One 6, e17896 (2011).
Zipfel, W.R., Williams, R.M. & Webb, W.W. Nonlinear magic: multiphoton microscopy in the biosciences. Nat. Biotechnol. 21, 1369–1377 (2003).
Rizzo, M.A., Springer, G., Segawa, K., Zipfel, W.R. & Piston, D.W. Optimization of pairings and detection conditions for measurement of FRET between cyan and yellow fluorescent proteins. Microsc. Microanal. 12, 238–254 (2006).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Thaler, C., Koushik, S.V., Blank, P.S. & Vogel, S.S. Quantitative multiphoton spectral imaging and its use for measuring resonance energy transfer. Biophys. J. 89, 2736–2749.
Zhou, X., Herbst-Robinson, K.J. & Zhang, J. Visualizing dynamic activities of signaling enzymes using genetically encodable FRET-based biosensors from designs to applications. Methods Enzymol. 504, 317–340 (2012).
Rincon, M.Y., VandenDriessche, T. & Chuah, M.K. Gene therapy for cardiovascular disease: advances in vector development, targeting, and delivery for clinical translation. Cardiovasc. Res. 108, 4–20 (2015).
McGhee, E.J. et al. FLIM-FRET imaging in vivoreveals 3D-environment spatially regulates RhoGTPase activity during cancer cell invasion. Small GTPases 2, 239–244 (2011).
Nobis, M. et al. Intravital FLIM-FRET imaging reveals dasatinib-induced spatial control of src in pancreatic cancer. Cancer Res. 73, 4674–4686 (2013).
Janssen, A., Beerling, E., Medema, R. & van Rheenen, J. Intravital FRET imaging of tumor cell viability and mitosis during chemotherapy. PLoS One 8, e64029 (2013).
Thestrup, T. et al. Optimized ratiometric calcium sensors for functional in vivo imaging of neurons and T lymphocytes. Nat. Methods 11, 175–182 (2014).
Luo, J. et al. A protocol for rapid generation of recombinant adenoviruses using the AdEasy system. Nat. Protoc. 2, 1236–1247 (2007).
Miller, R.A. et al. Biguanides suppress hepatic glucagon signaling by decreasing production of cyclic AMP. Nature 494, 256–260 (2013).
Day, R.N. Measuring protein interactions using Forster resonance energy transfer and fluorescence lifetime imaging microscopy. Methods 66, 200–207 (2014).
Broussard, J.A., Rappaz, B., Webb, D.J. & Brown, C.M. Fluorescence resonance energy transfer microscopy as demonstrated by measuring the activation of the serine/threonine kinase Akt. Nat. Protoc. 8, 265–281 (2013).
Periasamy, A. & Day, R.N. Molecular Imaging: FRET Microscopy and Spectroscopy (Oxford University Press, New York, 2005).
Acknowledgements
This research was supported by the National Institutes of Health O'Brien Center for Advanced Renal Microscopic Analysis (NIH-NIDDK P30DK079312 to R.N.D. and K.W.D.). Microscopy studies were conducted at the Indiana Center for Biological Microscopy. The authors thank M. Kamocka and S. Winfree for their assistance in microscopy. This article is dedicated to the memory of M.W. Davidson.
Author information
Authors and Affiliations
Contributions
R.N.D. and K.W.D. wrote the manuscript. W.T. conducted the experiments, and R.N.D and K.W.D. supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data
Sequence archive: DNA sequence information for plasmids used in this protocol (PDF 123 kb)
Rights and permissions
About this article
Cite this article
Day, R., Tao, W. & Dunn, K. A simple approach for measuring FRET in fluorescent biosensors using two-photon microscopy. Nat Protoc 11, 2066–2080 (2016). https://doi.org/10.1038/nprot.2016.121
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.121
This article is cited by
-
Dopamine D1 receptor signalling in dyskinetic Parkinsonian rats revealed by fiber photometry using FRET-based biosensors
Scientific Reports (2020)
-
Fit-free analysis of fluorescence lifetime imaging data using the phasor approach
Nature Protocols (2018)
-
Biosensing Technologies for Medical Applications, Manufacturing, and Regenerative Medicine
Current Stem Cell Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.