Abstract
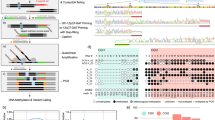
This protocol details a method for measuring the DNA methylation state of multiple target sites in single cells, otherwise known as single-cell restriction analysis of methylation (SCRAM). The basic steps include isolating and lysing single cells, digesting genomic DNA with a methylation-sensitive restriction endonuclease (MSRE) and amplification of multiple targets by two rounds of PCR to determine the methylation status of target sites. The method can reliably and accurately detect the methylation status of multiple target sites in each single cell, and it can be completed in a relatively short time (<2 d) at low cost. Consequently, the method may be preferable over whole-genome methods in applications requiring highly reliable and cost-effective coverage of specific target sites in all cells from a sample and in cases when the DNA methylation states of single CpG sites are representative of the methylation status of corresponding regions of interest.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Jaenisch, R. & Bird, A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 33, 245–254 (2003).
Reik, W. & Walter, J. Genomic imprinting: parental influence on the genome. Nat. Rev. Genet. 2, 21–32 (2001).
Goll, M.G. & Bestor, T.H. Eukaryotic cytosine methyltransferases. Annu. Rev. Biochem. 74, 481–514 (2005).
Messerschmidt, D.M., Knowles, B.B. & Solter, D. DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos. Genes Dev. 28, 812–828 (2014).
Smallwood, S.A. et al. Dynamic CpG island methylation landscape in oocytes and preimplantation embryos. Nat. Genet. 43, 811–814 (2011).
Santos, F., Hendrich, B., Reik, W. & Dean, W. Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev. Biol. 241, 172–182 (2002).
Smith, Z.D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Bock, C. et al. Quantitative comparison of genome-wide DNA methylation mapping technologies. Nat. Biotechnol. 28, 1106–1114 (2010).
Frommer, M. et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl. Acad. Sci. USA 89, 1827–1831 (1992).
Eckhardt, F. et al. DNA methylation profiling of human chromosomes 6, 20 and 22. Nat. Genet. 38, 1378–1385 (2006).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Colella, S., Shen, L., Baggerly, K.A., Issa, J.P.J. & Krahe, R. Sensitive and quantitative universal Pyrosequencing methylation analysis of CpG sites. Biotechniques 35, 146–151 (2003).
Herman, J.G., Graff, J.R., Myöhänen, S., Nelkin, B.D. & Baylin, S.B. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc. Natl. Acad. Sci. USA 93, 9821–9826 (1996).
Xiong, Z. & Laird, P.W. COBRA: a sensitive and quantitative DNA methylation assay. Nucleic Acids Res. 25, 2532–2534 (1997).
Eads, C.A. et al. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res. 28, e32 (2000).
Ehrich, M. et al. Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc. Natl. Acad. Sci. USA 102, 15785–15790 (2005).
Bibikova, M. et al. High-throughput DNA methylation profiling using universal bead arrays. Genome Res. 16, 383–393 (2006).
Weber, M. et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat. Genet. 37, 853–862 (2005).
Down, T.A. et al. A Bayesian deconvolution strategy for immunoprecipitation-based DNA methylome analysis. Nat. Biotechnol. 26, 779–785 (2008).
Khulan, B. et al. Comparative isoschizomer profiling of cytosine methylation: the HELP assay. Genome Res. 16, 1046–1055 (2006).
Melnikov, A.A., Gartenhaus, R.B., Levenson, A.S., Motchoulskaia, N.A. & Levenson, V.V. MSRE-PCR for analysis of gene-specific DNA methylation. Nucleic Acids Res. 33, e93 (2005).
Oakes, C.C., La Salle, S., Robaire, B. & Trasler, J.M. Evaluation of a quantitative DNA methylation analysis technique using methylation-sensitive/dependent restriction enzymes and real-time PCR. Epigenetics 1, 146–152 (2006).
Guo, H. et al. Single-cell methylome landscapes of mouse embryonic stem cells and early embryos analyzed using reduced representation bisulfite sequencing. Genome Res. 23, 2126–2135 (2013).
Smallwood, S.A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
El Hajj, N. et al. Limiting dilution bisulfite (pyro) sequencing reveals parent-specific methylation patterns in single early mouse embryos and bovine oocytes. Epigenetics 6, 1176–1188 (2011).
Gomez, D., Shankman, L.S., Nguyen, A.T. & Owens, G.K. Detection of histone modifications at specific gene loci in single cells in histological sections. Nat. Methods 10, 171–177 (2013).
Lorthongpanich, C. et al. Single-cell DNA-methylation analysis reveals epigenetic chimerism in preimplantation embryos. Science 341, 1110–1112 (2013).
Kantlehner, M. et al. A high-throughput DNA methylation analysis of a single cell. Nucleic Acids Res. 39, e44 (2011).
Ferguson-Smith, A.C. Genomic imprinting: the emergence of an epigenetic paradigm. Nat. Rev. Genet. 12, 565–575 (2011).
Suter, C.M., Martin, D.I. & Ward, R.L. Germline epimutation of MLH1 in individuals with multiple cancers. Nat. Genet. 36, 497–501 (2004).
Sermon, K., Van Steirteghem, A. & Liebaers, I. Preimplantation genetic diagnosis. Lancet 363, 1633–1641 (2004).
Barrera, V. & Peinado, M.A. Evaluation of single CpG sites as proxies of CpG island methylation states at the genome scale. Nucleic Acids Res. 40, 11490–11498 (2012).
Hansen, K.D. et al. Increased methylation variation in epigenetic domains across cancer types. Nat. Genet. 43, 768–775 (2011).
Diercks, A., Kostner, H. & Ozinsky, A. Resolving cell population heterogeneity: real-time PCR for simultaneous multiplexed gene detection in multiple single-cell samples. PLoS ONE 4, e6326 (2009).
Roberts, R.J., Vincze, T., Posfai, J. & Macelis, D. REBASE—a database for DNA restriction and modification: enzymes, genes and genomes. Nucleic Acids Res. 38, D234–D236 (2010).
Peixoto, A., Monteiro, M., Rocha, B. & Veiga-Fernandes, H. Quantification of multiple gene expression in individual cells. Genome Res. 14, 1938–1947 (2004).
Sanchez-Freire, V., Ebert, A.D., Kalisky, T., Quake, S.R. & Wu, J.C. Microfluidic single-cell real-time PCR for comparative analysis of gene expression patterns. Nat. Protoc. 7, 829–838 (2012).
Shapiro, H.M. Practical Flow Cytometry (John Wiley & Sons, 2005).
Yin, H. & Marshall, D. Microfluidics for single cell analysis. Curr. Opin. Biotechnol. 23, 110–119 (2012).
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. Manipulating the Mouse Embryo: a Laboratory Manual (Cold Spring Harbor Laboratory Press, 2003).
Suarez-Quian, C.A. et al. Laser capture microdissection of single cells from complex tissues. Biotechniques 26, 328–335 (1999).
Acknowledgements
This work was supported in part by a Visiting Investigator Programme grant (project nos. 092 110 0080 and 1235e00049) from the Joint Council Office of the Singapore Agency for Science, Technology and Research, a Career Development Award (to L.F.C., project no. 14302FG095) from the Joint Council Office of the Singapore Agency for Science, Technology and Research and by the Singapore Institute of Molecular and Cell Biology.
Author information
Authors and Affiliations
Contributions
L.F.C., S.R.Q., W.F.B. and D.M.M. have contributed to the development of the SCRAM assay. L.F.C. wrote the manuscript with assistance from W.F.B. and D.M.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Primer organization layout.
The high-throughput nature of the assay requires systematic primer organization. We pre-aliquot 50 µM primer mixes as depicted: For each locus/target (1, 2,..., 24) the two primer combinations (forward-short/reverse [green wells] and forward-long/reverse [blue wells]) are arranged pairwise on a 96-well plate. Each row (A-D) is then systematically transferred to the Assay Inlet ports of the Fluidigm array. Photo courtesy of Fluidigm Corporation.
Supplementary Figure 2 Melting curve analysis.
An exemplary melting curve analysis showing specific amplification in most of the single cell samples (blue arrowhead) and nonspecific amplification in a single sample (red arrowhead).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 (PDF 252 kb)
Rights and permissions
About this article
Cite this article
Cheow, L., Quake, S., Burkholder, W. et al. Multiplexed locus-specific analysis of DNA methylation in single cells. Nat Protoc 10, 619–631 (2015). https://doi.org/10.1038/nprot.2015.041
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.041
This article is cited by
-
Bisulfite-free epigenomics and genomics of single cells through methylation-sensitive restriction
Communications Biology (2021)
-
Development of a multiplex methylation-sensitive restriction enzyme-based SNP typing system for deconvolution of semen-containing mixtures
International Journal of Legal Medicine (2021)
-
DNA methylation studies in cattle
Journal of Applied Genetics (2021)
-
A novel fluorescent biosensor based on dendritic DNA nanostructure in combination with ligase reaction for ultrasensitive detection of DNA methylation
Journal of Nanobiotechnology (2019)
-
Single-cell multimodal profiling reveals cellular epigenetic heterogeneity
Nature Methods (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.