Abstract
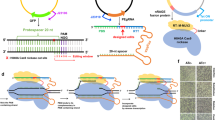
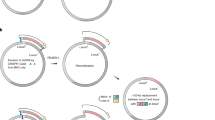
Multiplex automated genome engineering (MAGE) is a powerful technology for in vivo genome editing that uses synthetic single-stranded DNA (ssDNA) to introduce targeted modifications directly into the Escherichia coli chromosome. MAGE is a cyclical process that involves transformation of ssDNA (by electroporation) followed by outgrowth, during which bacteriophage homologous recombination proteins mediate annealing of ssDNAs to their genomic targets. By iteratively introducing libraries of mutagenic ssDNAs targeting multiple sites, MAGE can generate combinatorial genetic diversity in a cell population. Alternatively, MAGE can introduce precise mutant alleles at many loci for genome-wide editing or for recoding projects that are not possible with other methods. In recent technological advances, MAGE has been improved by strain modifications and selection techniques that enhance allelic replacement. This protocol describes the manual execution of MAGE wherein each cycle takes ∼2.5 h, which, if carried out by two people, allows ∼10 continuous cycles of MAGE-based mutagenesis per day.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
Accession codes
References
Metzker, M.L. Sequencing technologies—the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Ashworth, J. et al. Computational redesign of endonuclease DNA binding and cleavage specificity. Nature 441, 656–659 (2006).
Paddon, C.J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Carroll, D. Genome engineering with targetable nucleases. Annu. Rev. Biochem. 83, 409–439 (2014).
Jackel, C., Kast, P. & Hilvert, D. Protein design by directed evolution. Annu. Rev. Biophys. 37, 153–173 (2008).
Zhang, Y., Buchholz, F., Muyrers, J.P. & Stewart, A.F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 97, 5978–5983 (2000).
Ellis, H.M., Yu, D., DiTizio, T. & Court, D.L. High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc. Natl. Acad. Sci. USA 98, 6742–6746 (2001).
Isaacs, F.J. et al. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 333, 348–353 (2011).
Lajoie, M.J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Wang, H.H. et al. Genome-scale promoter engineering by coselection MAGE. Nat. Methods 9, 591–593 (2012).
Caruthers, M.H. Gene synthesis machines: DNA chemistry and its uses. Science 230, 281–285 (1985).
Carr, P.A. & Church, G.M. Genome engineering. Nat. Biotechnol. 27, 1151–1162 (2009).
Kosuri, S. et al. Scalable gene synthesis by selective amplification of DNA pools from high-fidelity microchips. Nat. Biotechnol. 28, 1295–1299 (2010).
Orr-Weaver, T.L., Szostak, J.W. & Rothstein, R.J. Yeast transformation: a model system for the study of recombination. Proc. Natl. Acad. Sci. USA 78, 6354–6358 (1981).
Kowalczykowski, S.C., Dixon, D.A., Eggleston, A.K., Lauder, S.D. & Rehrauer, W.M. Biochemistry of homologous recombination in Escherichia coli. Microbiol. Rev. 58, 401–465 (1994).
Muniyappa, K. & Radding, C.M. The homologous recombination system of phage λ. Pairing activities of β-protein. J. Biol. Chem. 261, 7472–7478 (1986).
Haseltine, E.L. & Arnold, F.H. Synthetic gene circuits: design with directed evolution. Annu. Rev. Biophys. Biomol. Struct. 36, 1–19 (2007).
Stemmer, W.P. DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution. Proc. Natl. Acad. Sci. USA 91, 10747–10751 (1994).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage-T4 DNA polymerase. Science 249, 505–510 (1990).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Soukup, G.A. & Breaker, R.R. Engineering precision RNA molecular switches. Proc. Natl. Acad. Sci. USA 96, 3584–3589 (1999).
Orth, J.D., Thiele, I. & Palsson, B.O. What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Burgard, A.P., Pharkya, P. & Maranas, C.D. Optknock: a bilevel programming framework for identifying gene knockout strategies for microbial strain optimization. Biotechnol. Bioeng. 84, 647–657 (2003).
Salis, H.M., Mirsky, E.A. & Voigt, C.A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Binder, S., Siedler, S., Marienhagen, J., Bott, M. & Eggeling, L. Recombineering in Corynebacterium glutamicum combined with optical nanosensors: a general strategy for fast producer strain generation. Nucleic Acids Res. 41, 6360–6369 (2013).
Carr, P.A. et al. Enhanced multiplex genome engineering through co-operative oligonucleotide co-selection. Nucleic Acids Res. 40, e132 (2012).
Datta, S., Costantino, N., Zhou, X. & Court, D.L. Identification and analysis of recombineering functions from Gram-negative and Gram-positive bacteria and their phages. Proc. Natl. Acad. Sci. USA 105, 1626–1631 (2008).
Zhang, Y., Muyrers, J.P., Rientjes, J. & Stewart, A.F. Phage annealing proteins promote oligonucleotide-directed mutagenesis in Escherichia coli and mouse ES cells. BMC Mol. Biol. 4, 1 (2003).
Li, X.T. et al. Identification of factors influencing strand bias in oligonucleotide-mediated recombination in Escherichia coli. Nucleic Acids Res. 31, 6674–6687 (2003).
Lajoie, M.J., Gregg, C.J., Mosberg, J.A., Washington, G.C. & Church, G.M. Manipulating replisome dynamics to enhance λ Red–mediated multiplex genome engineering. Nucleic Acids Res. 40, e170 (2012).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Babic, I., Andrew, S.E. & Jirik, F.R. MutS interaction with mismatch and alkylated base containing DNA molecules detected by optical biosensor. Mutat. Res. 372, 87–96 (1996).
Sawitzke, J.A. et al. Probing cellular processes with oligo-mediated recombination and using the knowledge gained to optimize recombineering. J. Mol. Biol. 407, 45–59 (2011).
Wang, H.H., Xu, G., Vonner, A.J. & Church, G. Modified bases enable high-efficiency oligonucleotide-mediated allelic replacement via mismatch repair evasion. Nucleic Acids Res. 39, 7336–7347 (2011).
Swingle, B. et al. Oligonucleotide recombination in Gram-negative bacteria. Mol. Microbiol. 75, 138–148 (2010).
Costantino, N. & Court, D.L. Enhanced levels of Red–mediated recombinants in mismatch repair mutants. Proc. Natl. Acad. Sci. USA 100, 15748–15753 (2003).
Nyerges, A. et al. Conditional DNA repair mutants enable highly precise genome engineering. Nucleic Acids Res. 42, e62 (2014).
Mosberg, J.A., Gregg, C.J., Lajoie, M.J., Wang, H.H. & Church, G.M. Improving Red genome engineering in Escherichia coli via rational removal of endogenous nucleases. PLoS ONE 7, e44638 (2012).
DeVito, J.A. Recombineering with tolC as a selectable/counter-selectable marker: remodeling the rRNA operons of Escherichia coli. Nucleic Acids Res. 36, e4 (2008).
Warming, S., Costantino, N., Court, D.L., Jenkins, N.A. & Copeland, N.G. Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res. 33, e36 (2005).
Gregg, C.J. et al. Rational optimization of tolC as a powerful dual-selectable marker for genome engineering. Nucleic Acids Res. 42, 4779–4790 (2014).
Herschma, H.R. & Helinski, D.R. Comparative study of events associated with colicin induction. J. Bacteriol. 94, 691 (1967).
Schwartz, S.A. & Helinski, D.R. Purification and characterization of colicin E1. J. Biol. Chem. 246, 6318–6327 (1971).
Wang, H.H. & Church, G.M. Multiplexed genome engineering and genotyping methods applications for synthetic biology and metabolic engineering. Methods Enzymol. 498, 409–426 (2011).
Acknowledgements
Funding was provided by the US Department of Energy (DE-FG02-02ER63445), Defense Advanced Research Projects Agency (N66001-12-C-4020, N66001-12-C-4211), and the Arnold and Mabel Beckman Foundation (F.J.I.).
Author information
Authors and Affiliations
Contributions
R.R.G., Z.L. and F.J.I. wrote the manuscript with contributions on mathematical modeling from A.O.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1: Safe insertion regions in E. coli MG1655
(Numbering according to NCBI's RefSeq accession number PRJNA57779). (PDF 227 kb)
Rights and permissions
About this article
Cite this article
Gallagher, R., Li, Z., Lewis, A. et al. Rapid editing and evolution of bacterial genomes using libraries of synthetic DNA. Nat Protoc 9, 2301–2316 (2014). https://doi.org/10.1038/nprot.2014.082
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.082
This article is cited by
-
A synthetic population-level oscillator in non-microfluidic environments
Communications Biology (2023)
-
Phosphite synthetic auxotrophy as an effective biocontainment strategy for the industrial chassis Pseudomonas putida
Microbial Cell Factories (2022)
-
Development and characterization of a glycine biosensor system for fine-tuned metabolic regulation in Escherichia coli
Microbial Cell Factories (2022)
-
Protein nanowires with tunable functionality and programmable self-assembly using sequence-controlled synthesis
Nature Communications (2022)
-
Lifestyle-specific S-nitrosylation of protein cysteine thiols regulates Escherichia coli biofilm formation and resistance to oxidative stress
npj Biofilms and Microbiomes (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.