Abstract
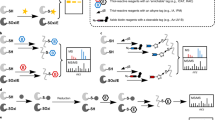
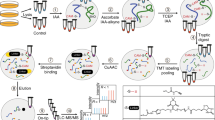
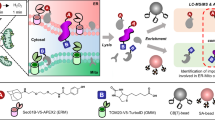
Reversible thiol oxidation of cysteine residues occurs in many intracellular catalytic and signaling processes. Here we describe an optimized protocol, which can be completed in ∼5 d, to unambiguously identify specific cysteine residues that are transiently and reversibly oxidized by comparing two complex biological samples obtained from yeast cell cultures at the proteome level. After 'freezing' the in vivo thiol stage of cysteine residues by medium acidification, we first block reduced thiols in extracts with iodoacetamide (IAM), and then we sequentially reduce and label reversible oxidized thiols with the biotin-based heavy or light IAM derivatives, which are known as isotope-coded affinity tag (ICAT) reagents, so that the two samples can be compared at once after combination of the labeled extracts, trypsin digestion, streptavidin-affinity purification of peptides containing oxidized cysteines, and liquid chromatography–tandem mass spectrometry (LC-MS/MS) analysis. For the same protein extracts, before cysteine-containing peptide enrichment, individual relative protein concentrations are obtained by stable-isotope dimethyl labeling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Gilbert, H.F. Molecular and cellular aspects of thiol-disulfide exchange. Adv. Enzymol. Relat. Areas Mol. Biol. 63, 69–172 (1990).
Ghezzi, P. Protein glutathionylation in health and disease. Biochim. Biophys. Acta. 1830, 3165–3172 (2013).
Auclair, J.R. et al. Structural consequences of cysteinylation of Cu/Zn-superoxide dismutase. Biochemistry 52, 6145–6150 (2013).
Hochgrafe, F. et al. S-cysteinylation is a general mechanism for thiol protection of Bacillus subtilis proteins after oxidative stress. J. Biol. Chem. 282, 25981–2595 (2007).
Salmeen, A. et al. Redox regulation of protein tyrosine phosphatase 1B involves a sulphenyl-amide intermediate. Nature 423, 769–773 (2003).
van Montfort, R.L., Congreve, M., Tisi, D., Carr, R. & Jhoti, H. Oxidation state of the active-site cysteine in protein tyrosine phosphatase 1B. Nature 423, 773–777 (2003).
Winterbourn, C.C. & Hampton, M.B. Thiol chemistry and specificity in redox signaling. Free Radic. Biol. Med. 45, 549–561 (2008).
D'Autreaux, B. & Toledano, M.B. ROS as signalling molecules: mechanisms that generate specificity in ROS homeostasis. Nat. Rev. Mol. Cell Biol. 8, 813–824 (2007).
Hess, D.T., Matsumoto, A., Kim, S.O., Marshall, H.E. & Stamler, J.S. Protein S-nitrosylation: purview and parameters. Nat. Rev. Mol. Cell Biol. 6, 150–166 (2005).
Meyer, Y., Buchanan, B.B., Vignols, F. & Reichheld, J.P. Thioredoxins and glutaredoxins: unifying elements in redox biology. Annu. Rev. Genet. 43, 335–367 (2009).
Vlamis-Gardikas, A. The multiple functions of the thiol-based electron flow pathways of Escherichia coli: Eternal concepts revisited. Biochim. Biophys. Acta. 1780, 1170–1200 (2008).
Sengupta, R. & Holmgren, A. Thioredoxin and thioredoxin reductase in relation to reversible S-nitrosylation. Antioxid. Redox Signal. 18, 259–269 (2013).
Benhar, M., Forrester, M.T. & Stamler, J.S. Protein denitrosylation: enzymatic mechanisms and cellular functions. Nat. Rev. Mol. Cell Biol. 10, 721–732 (2009).
Poole, L.B. & Nelson, K.J. Discovering mechanisms of signaling-mediated cysteine oxidation. Curr. Opin. Chem. Biol. 12, 18–24 (2008).
Anand, P. & Stamler, J.S. Enzymatic mechanisms regulating protein S-nitrosylation: implications in health and disease. J. Mol. Med. 90, 233–44 (2012).
Sha, Y. & Marshall, H.E. S-nitrosylation in the regulation of gene transcription. Biochim. Biophys. Acta. 1820, 701–711 (2012).
Marozkina, N.V. & Gaston, B. S-Nitrosylation signaling regulates cellular protein interactions. Biochim. Biophys. Acta. 1820, 722–729 (2012).
Cumming, R.C. et al. Protein disulfide bond formation in the cytoplasm during oxidative stress. J. Biol. Chem. 279, 21749–21758 (2004).
Jaffrey, S.R., Erdjument-Bromage, H., Ferris, C.D., Tempst, P. & Snyder, S.H. Protein S-nitrosylation: a physiological signal for neuronal nitric oxide. Nat. Cell Biol. 3, 193–197 (2001).
Wang, X., Kettenhofen, N.J., Shiva, S., Hogg, N. & Gladwin, M.T. Copper dependence of the biotin switch assay: modified assay for measuring cellular and blood nitrosated proteins. Free Radic. Biol. Med. 44, 1362–1372 (2008).
Saurin, A.T., Neubert, H., Brennan, J.P. & Eaton, P. Widespread sulfenic acid formation in tissues in response to hydrogen peroxide. Proc. Natl. Acad. Sci. USA 101, 17982–17987 (2004).
Lind, C. et al. Identification of S-glutathionylated cellular proteins during oxidative stress and constitutive metabolism by affinity purification and proteomic analysis. Arch. Biochem. Biophys. 406, 229–240 (2002).
Leichert, L.I. et al. Quantifying changes in the thiol redox proteome upon oxidative stress in vivo. Proc. Natl. Acad. Sci. USA 105, 8197–8202 (2008).
Le Moan, N., Clement, G., Le Maout, S., Tacnet, F. & Toledano, M.B. The Saccharomyces cerevisiae proteome of oxidized protein thiols: contrasted functions for the thioredoxin and glutathione pathways. J. Biol. Chem. 281, 10420–10430 (2006).
Baty, J.W., Hampton, M.B. & Winterbourn, C.C. Proteomic detection of hydrogen peroxide-sensitive thiol proteins in Jurkat cells. Biochem. J. 389, 785–795 (2005).
Delahunty, C. & Yates, J.R. III Protein identification using 2D-LC-MS/MS. Methods 35, 248–255 (2005).
Hao, G., Derakhshan, B., Shi, L., Campagne, F. & Gross, S.S. SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures. Proc. Natl. Acad. Sci. USA 103, 1012–1017 (2006).
Sinha, V. et al. Proteomic and mass spectroscopic quantitation of protein S-nitrosation differentiates NO-donors. ACS Chem. Biol. 5, 667–680 (2010).
Foster, M.W. Methodologies for the characterization, identification and quantification of S-nitrosylated proteins. Biochim. Biophys. Acta 1820, 675–683 (2012).
Bechtold, E. & King, S.B. Chemical methods for the direct detection and labeling of S-nitrosothiols. Antioxid. Redox Signal. 17, 981–991 (2012).
Martinez-Acedo, P. et al. A novel strategy for global analysis of the dynamic thiol redox proteome. Mol. Cell Proteomics 11, 800–813 (2012).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Sethuraman, M. et al. Isotope-coded affinity tag (ICAT) approach to redox proteomics: identification and quantitation of oxidant-sensitive cysteine thiols in complex protein mixtures. J. Proteome Res. 3, 1228–1233 (2004).
Sethuraman, M. et al. Quantification of oxidative posttranslational modifications of cysteine thiols of p21ras associated with redox modulation of activity using isotope-coded affinity tags and mass spectrometry. Free Radic. Biol. Med. 42, 823–829 (2007).
Sethuraman, M., McComb, M.E., Heibeck, T., Costello, C.E. & Cohen, R.A. Isotope-coded affinity tag approach to identify and quantify oxidant-sensitive protein thiols. Mol. Cell Proteomics 3, 273–278 (2004).
Fu, C. et al. Quantitative analysis of redox-sensitive proteome with DIGE and ICAT. J. Proteome Res. 7, 3789–3802 (2008).
Garcia-Santamarina, S. et al. The oxidized thiol proteome in fission yeast–optimization of an ICAT-based method to identify H2O2-oxidized proteins. J. Proteomics 74, 2476–2486 (2011).
Garcia-Santamarina, S. et al. Is oxidized thioredoxin a major trigger for cysteine oxidation? Clues from a redox proteomics approach. Antioxid. Redox Signal. 18, 1549–1556 (2013).
Brandes, N., Reichmann, D., Tienson, H., Leichert, L.I. & Jakob, U. Using quantitative redox proteomics to dissect the yeast redoxome. J. Biol. Chem. 286, 41893–41903 (2011).
Kumsta, C., Thamsen, M. & Jakob, U. Effects of oxidative stress on behavior, physiology, and the redox thiol proteome of Caenorhabditis elegans. Antioxid. Redox Signal. 14, 1023–1037 (2011).
Knoefler, D. et al. Quantitative in vivo redox sensors uncover oxidative stress as an early event in life. Mol. Cell 47, 767–776 (2012).
Christoforou, A.L. & Lilley, K.S. Isobaric tagging approaches in quantitative proteomics: the ups and downs. Anal. Bioanal. Chem. 404, 1029–1037 (2012).
Boersema, P.J., Raijmakers, R., Lemeer, S., Mohammed, S. & Heck, A.J. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat. Protoc. 4, 484–494 (2009).
Wisniewski, J.R., Zougman, A. & Mann, M. Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J. Proteome Res. 8, 5674–5678 (2009).
Guo, J. et al. Resin-assisted enrichment of thiols as a general strategy for proteomic profiling of cysteine-based reversible modifications. Nat. Protoc. 9, 64–75 (2014).
Hansen, R.E. & Winther, J.R. An introduction to methods for analyzing thiols and disulfides: Reactions, reagents, and practical considerations. Anal. Biochem. 394, 147–158 (2009).
Gilbert, H.F. Thiol/disulfide exchange equilibria and disulfide bond stability. Methods Enzymol. 251, 8–28 (1995).
Rogers, L.K., Leinweber, B.L. & Smith, C.V. Detection of reversible protein thiol modifications in tissues. Anal. Biochem. 358, 171–184 (2006).
Held, J.M. & Gibson, B.W. Regulatory control or oxidative damage? Proteomic approaches to interrogate the role of cysteine oxidation status in biological processes. Mol. Cell Proteomics 11, R111 013037 (2012).
Sivaraman, T., Kumar, T.K., Jayaraman, G. & Yu, C. The mechanism of 2,2,2-trichloroacetic acid-induced protein precipitation. J. Protein Chem. 16, 291–297 (1997).
Bensadoun, A. & Weinstein, D. Assay of proteins in the presence of interfering materials. Anal. Biochem. 70, 241–250 (1976).
Rajalingam, D., Loftis, C., Xu, J.J. & Kumar, T.K. Trichloroacetic acid-induced protein precipitation involves the reversible association of a stable partially structured intermediate. Protein Sci. 18, 980–993 (2009).
Boja, E.S. & Fales, H.M. Overalkylation of a protein digest with iodoacetamide. Anal. Chem. 73, 3576–3582 (2001).
Gundlach, H.G., Moore, S. & Stein, W.H. The reaction of iodoacetate with methionine. J. Biol. Chem. 234, 1761–1764 (1959).
Lapko, V.N., Smith, D.L. & Smith, J.B. Identification of an artifact in the mass spectrometry of proteins derivatized with iodoacetamide. J. Mass Spectrom. 35, 572–575 (2000).
Crestfield, A.M., Stein, W.H. & Moore, S. Alkylation and identification of the histidine residues at the active site of ribonuclease. J. Biol. Chem. 238, 2413–2419 (1963).
Park, S.S. et al. Effective correction of experimental errors in quantitative proteomics using stable isotope labeling by amino acids in cell culture (SILAC). J. Proteomics 75, 3720–3732 (2012).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Kruger, R., Hung, C.W., Edelson-Averbukh, M. & Lehmann, W.D. Iodoacetamide-alkylated methionine can mimic neutral loss of phosphoric acid from phosphopeptides as exemplified by nano-electrospray ionization quadrupole time-of-flight parent ion scanning. Rapid Commun. Mass Spectrom. 19, 1709–1716 (2005).
Hung, C.W., Schlosser, A., Wei, J. & Lehmann, W.D. Collision-induced reporter fragmentations for identification of covalently modified peptides. Anal. Bioanal. Chem. 389, 1003–1016 (2007).
Gouw, J.W. & Krijgsveld, J. MSQuant: a platform for stable isotope-based quantitative proteomics. Methods Mol. Biol. 893, 511–522 (2012).
Mortensen, P. et al. MSQuant, an open source platform for mass spectrometry-based quantitative proteomics. J. Proteome Res. 9, 393–403 (2010).
Acknowledgements
We acknowledge the Centre for Genomic Regulation-Universitat Pompeu Fabra (CRG-UPF) proteomic facility where the LC-MS/MS experiments were performed. This work was supported by the Spanish Ministry of Science and Innovation (nos. BFU2009-06933 and BFU2012-32045), by PLAN E and Fondo Europeo de Desarrollo Regional (FEDER), by the Spanish program Consolider-Ingenio 2010 (grant no. CSD 2007-0020) and by grant no. SGR2009-195 from Generalitat de Catalunya (Spain) to E.H. E.H. and J.A. are recipients of Institució Catalana de Recerca i Estudis Avançats (ICREA) Academia Awards (Generalitat de Catalunya).
Author information
Authors and Affiliations
Contributions
S.G.-S. and S.B. developed the protocol and conducted the experiments. S.G.-S., H.M. and S.B. interpreted the data. H.M. performed the LC-MS/MS experiments and edited the manuscript. A.D. performed some control experiments. J.A. provided intellectual expertise. S.G.-S. drafted the manuscript. E.H. was responsible for project supervision, data interpretation, manuscript editing and providing grant support.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 TCA followed by IAM, but not only IAM, is required to display the transient H2O2-dependent in vivo oxidation of Pap1.
Wild-type strain 972 was treated or not with 0.2 mM H2O2 for 5 min. Cells were then lysed in the presence of TCA, proteins in the lysates were then alkylated with IAM, and Pap1 redox state was analyzed by non-reducing electrophoresis and Western blot analysis as described before (Vivancos et al. 2005, PNAS 102:8875) (A). Alternatively, 75 mM IAM was added to both cell cultures and cell pellets prior to lysis (B), or only to cell pellets prior to lysis (C), and samples were processed for Western blot analysis as described above.
Supplementary Figure 2 Effect of SDS and Tris-HCl in the labeling of oxidized cysteines and proteins in their N-terminal sites.
Left panel, oxidized cysteines from wild-type (WT) and Δtrr1 cells were labelled with a fluorescent derivative of iodoacetamide (FIAM) as described in García-Santamarina et al. 2011, J. Proteomics 74:2476, using either ICAT buffer (buffer 1; 200 mM Tris-HCl, 0.05% SDS, 6 M urea, 5 mM EDTA) or a buffer without Tris-HCl and SDS (buffer 2; 200 mM Triethyl-ammonium bicarbonate 1 M, pH 8.5 , 6 M urea). Right panel, 100 µg of protein extracts in 50 µl from wild-type (WT) and Δtrr1 cells were labelled at their N-terminal sites with 2.5 µl of 1 mg/ml fluorescein isothiocyanate (FITC), either in ICAT buffer (buffer 1) or in a buffer without Tris-HCl and SDS (buffer 2). In both cases, protein extracts were electrophoretically separated by reducing SDS-PAGE. After electrophoresis fluorescence was scanned in a Typhoon 8600 using λex. 532 nm and λem. 526 nm. Silver staining was used as a protein loading control.
Supplementary information
Supplementary Figure 1
TCA followed by IAM, but not only IAM, is required to display the transient H2O2-dependent in vivo oxidation of Pap1. (PDF 93 kb)
Supplementary Figure 2
Effect of SDS and Tris-HCl in the labeling of oxidized cysteines and proteins in their N-terminal sites. (PDF 319 kb)
Supplementary Table 1
Peptides with oxidized cysteines (ICAT ratio vs protein concentration >1.5 in two biological replicates) identified from wild-type H2O2-treated vs wild-type untreated fission yeast cells. (XLS 48 kb)
Supplementary Table 2
Peptides with oxidized cysteines (ICAT ratio vs protein concentration >1.5 in two biological replicates) identified from Δtrr1 vs wild-type untreated fission yeast cells. (XLS 48 kb)
Supplementary Table 3
Peptides with oxidized cysteines (ICAT ratio vs protein concentration >1.5 in two biological replicates) identified from Δtrx1 vs wild-type untreated fission yeast cells. (XLS 37 kb)
Supplementary Data 1
Description LC gradient with 84 min (marked '120 minutes') and 147 min (marked '180 minutes') resolving gradients. (PDF 40 kb)
Supplementary Data 2
MS and MS/MS parameters for a Q-Exactive mass spectrometer operated in data dependent (dd) mode. (PDF 100 kb)
Supplementary Data 3
Retention time comparison of identical cysteine-containing peptides modified by carbamidomethyl (ca) and/or ICAT measured in the same experiment. (PDF 109 kb)
Supplementary Data 4
MS/MS spectra of the triply charged peptide. (PDF 130 kb)
Supplementary Data 5
MS spectrum of the triply charged peptide. (PDF 143 kb)
Supplementary Data 6
Three dimensional plot of the ICAT-labelled peptide pair shown in Supplementary Data 5. (PDF 283 kb)
Supplementary Data 7
Modifications used to search tandem MS data (MS/MS). (PDF 70 kb)
Supplementary Data 8
Five important steps needed to launch a MaxQuant 1.3.0.5 analysis. (PDF 150 kb)
Supplementary Data 9
The 'peptide.txt' file contains the dimethyl-based quantification results as well as the ICAT-based quantification results. (PDF 52 kb)
Supplementary Data 10
Scatter plot of measured ICAT ratios. (PDF 54 kb)
Rights and permissions
About this article
Cite this article
García-Santamarina, S., Boronat, S., Domènech, A. et al. Monitoring in vivo reversible cysteine oxidation in proteins using ICAT and mass spectrometry. Nat Protoc 9, 1131–1145 (2014). https://doi.org/10.1038/nprot.2014.065
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.065
This article is cited by
-
Metabolism-based targeting of MYC via MPC-SOD2 axis-mediated oxidation promotes cellular differentiation in group 3 medulloblastoma
Nature Communications (2023)
-
A quantitative thiol reactivity profiling platform to analyze redox and electrophile reactive cysteine proteomes
Nature Protocols (2020)
-
Proteome-wide detection of S-nitrosylation targets and motifs using bioorthogonal cleavable-linker-based enrichment and switch technique
Nature Communications (2019)
-
Proteome-wide analysis of cysteine oxidation reveals metabolic sensitivity to redox stress
Nature Communications (2018)
-
T-REX on-demand redox targeting in live cells
Nature Protocols (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.