Abstract
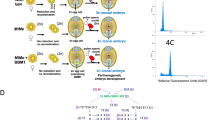
Hybrid crop varieties are traditionally produced by selecting and crossing parental lines to evaluate hybrid performance. Reverse breeding allows doing the opposite: selecting uncharacterized heterozygotes and generating parental lines from them. With these, the selected heterozygotes can be recreated as F1 hybrids, greatly increasing the number of hybrids that can be screened in breeding programs. Key to reverse breeding is the suppression of meiotic crossovers in a hybrid plant to ensure the transmission of nonrecombinant chromosomes to haploid gametes. These gametes are subsequently regenerated as doubled-haploid (DH) offspring. Each DH carries combinations of its parental chromosomes, and complementing pairs can be crossed to reconstitute the initial hybrid. Achiasmatic meiosis and haploid generation result in uncommon phenotypes among offspring owing to chromosome number variation. We describe how these features can be dealt with during a reverse-breeding experiment, which can be completed in six generations (∼1 year).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Link, W. & Melchinger, A.E. An approach to the genetic improvement of clonal cultivars via backcrosing. Crop Sci. 35, 931 (1995).
Dirks, R. et al. Reverse breeding: a novel breeding approach based on engineered meiosis. Plant Biotechnol. J. 7, 837–845 (2009).
Wijnker, E. et al. Reverse breeding in Arabidopsis thaliana generates homozygous parental lines from a heterozygous plant. Nat. Genet. 44, 467–470 (2012).
Bishop, D.K., Park, D., Xu, L. & Kleckner, N. DMC1: A meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69, 439–456 (1992).
Law, C.N., Snape, J.W. & Worland, A.J. in Wheat Breeding. Its Scientific Basis (ed. Lupton, F.G.H.) 71–108 (Chapman & Hall, 1987).
Koumproglou, R. et al. STAIRS: a new genetic resource for functional genomic studies of Arabidopsis. Plant J. 31, 355–364 (2002).
Singer, J.B. et al. Genetic dissection of complex traits with chromosome substitution strains of mice. Science 304, 445–448 (2004).
Kooke, R., Wijnker, E. & Keurentjes, J.B. in Quantitative Trait Loci (QTL) (ed. Rifkin, S.A.) 871, 3–16 (Humana Press, 2012).
Nadeau, J.H., Singer, J.B., Matin, A. & Lander, E.S. Analysing complex genetic traits with chromosome substitution strains. Nat. Genet. 24, 221–225 (2000).
Perera, S.A.C.N., Pooni, H.S. & Kearsey, M.J. Chromosome substitution lines for the analysis of heterosis in Arabidopsis thaliana. J. Natl. Sci. Found. Sri Lanka 36, 275–280 (2008).
Tuinstra, M.R., Ejeta, G. & Goldsbrough, P.B. Heterogeneous inbred family (HIF) analysis: a method for developing near-isogenic lines that differ at quantitative trait loci. Theor. Appl. Genet. 95, 1005–1011 (1997).
Keurentjes, J.J.B. et al. Development of a near-isogenic line population of Arabidopsis thaliana and comparison of mapping power with a recombinant inbred line population. Genetics 175, 891–905 (2007).
Ravi, M. & Chan, S.W.L. Haploid plants produced by centromere-mediated genome elimination. Nature 464, 615–618 (2010).
Bikard, D. et al. Divergent evolution of duplicate genes leads to genetic incompatibilities within A. thaliana. Science 323, 623–626 (2009).
Seymour, D.K. et al. Rapid creation of Arabidopsis doubled haploid lines for quantitative trait locus mapping. Proc. Natl. Acad. Sci. USA 109, 4227–4232 (2012).
Hartung, F. et al. The catalytically active tyrosine residues of both SPO11-1 and SPO11-2 are required for meiotic double-strand break induction in Arabidopsis. Plant Cell Online 19, 3090–3099 (2007).
Koornneef, M. & Veen, J.H. Trisomics in Arabidopsis thaliana and the location of linkage groups. Genetica 61, 41–46 (1983).
Henry, I.M., Dilkes, B.P., Miller, E.S., Burkart-Waco, D. & Comai, L. Phenotypic consequences of aneuploidy in Arabidopsis thaliana. Genetics 186, 1231–1245 (2010).
Martínez, P., López, C., Roldán, M., Sabater, B. & Martín, M. Plastid DNA of five ecotypes of Arabidopsis thaliana: sequence of ndhG gene and maternal inheritance. Plant Sci. 123, 113–122 (1997).
Zhang, X., Henriques, R., Lin, S.-S., Niu, Q.-W. & Chua, N.-H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 1, 641–646 (2006).
Edwards, K., Johnstone, C. & Thompson, C. A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res. 19, 1349–1349 (1991).
Berendzen, K. et al. A rapid and versatile combined DNA/RNA extraction protocol and its application to the analysis of a novel DNA marker set polymorphic between Arabidopsis thaliana ecotypes Col-0 and Landsberg erecta. Plant Methods 1, 4 (2005).
Kasajima, I. et al. A protocol for rapid DNA extraction from Arabidopsis thaliana for PCR analysis. Plant Mol. Biol. Rep. 22, 49–52 (2004).
Wang, H., Qi, M. & Cutler, A.J. A simple method of preparing plant samples for PCR. Nucleic Acids Res. 21, 4153–4154 (1993).
Singer, T. & Burke, E. in Plant Functional Genomics (ed. Grotewold, E.) 236, 241–271 (Humana Press, 2003).
O'Malley, R.C., Alonso, J.M., Kim, C.J., Leisse, T.J. & Ecker, J.R. An adapter ligation-mediated PCR method for high-throughput mapping of T-DNA inserts in the Arabidopsis genome. Nat. Protoc. 2, 2910–2917 (2007).
Ravi, M. et al. Meiosis-specific loading of the centromere-specific histone CENH3 in Arabidopsis thaliana. PLoS Genet. 7, e1002121 (2011).
Acknowledgements
We thank C. Pillen, T. Stoker, G. Stunnenberg and B. Essenstam (Wageningen) for their help in the greenhouse. P. Flood (Wageningen) is kindly acknowledged for providing a picture of GFP-tailswap plants. This work was in part supported by a European Molecular Biology Organization (EMBO) long-term fellowship (E.W.).
Author information
Authors and Affiliations
Contributions
K.v.D., C.L.C.L., H.d.J. and R.D. conceived the research. E.W. performed crosses. C.B.d.S. constructed vectors and performed genotyping. L.D., J.v.d.B., H.B. and F.B. assisted in experimental work and plant phenotyping. E.W. analyzed the data with the help of C.B.d.S., J.J.B.K., M.R. and S.W.L.C. E.W. and J.J.B.K. wrote the manuscript. All authors were involved in planning and design of experiments, and all authors read and improved the manuscript.
Corresponding author
Ethics declarations
Competing interests
E.W. is a former employee of Rijk Zwaan. K.v.D., C.B.d.S., C.L.C.L. and R.D. are employees of Rijk Zwaan. H.d.J. and J.J.B.K. have received research funding from Rijk Zwaan in recent years.
Integrated supplementary information
Supplementary Figure 1 Comparison of WT (left) and RNAi:DMC1 transformed achiasmatic line (right) in a CVI background.
Note the long siliques on the WT plant in comparison to the semi-sterile line on the right. Semi-sterile lines typically show no or strongly reduced elongation of siliques as compared to WT plants, as is clearly visible on the main stem of the inflorescence.
Supplementary Figure 2 Typical offspring of a cross of a WT F1 to the GFP-tailswap haploid inducer showing GFP-tailswap selfs, haploids, aneuploids and diploid heterozygote offspring between GFP-tailswap and the WT F1.
Top row (left to right): GFP-tailswap self, heterozygote, aneuploid, aneuploid and a heterozygote. Second row: GFP-tailswap, haploid, heterozygote, haploid, heterozygote. Third row: GFP-tailswap, haploid, haploid, haploid, heterozygote. Bottom row: haploid, haploid, haploid, haploid, heterozygote. The WT F1 in this cross is a cross between two late-flowering accessions. Note the curly leaf and (uniform!) early flowering phenotype of the three GFP-tailswap plants; haploids predominantly shown narrow leaves in comparison to GFP-tailswap x WT F1 heterozygotes that grow more vigorously. Also note that due to segregation of alleles, heterozygotes and haploids do not show uniform phenotypes. The two aneuploids show aberrant phenotypes.
Supplementary Figure 3 Identification of aneuploids.
The top row shows six WT Col-0 plants. Note that these specific plants do not grow uniformly. Intra-line variation indicates that in this case plants may have experienced some stress. The bottom row shows RNAi:DMC1 transgenic offspring, originating from an achiasmatic parental Col-0. Comparisons to the upper row suggest that plants two, three, four and six (counting from the left) show similar phenotypes as WT Col-0 and can reliably be identified as eudiploid offspring. Plants one (lower left) and five are aneuploids showing aberrant phenotypes. Note that without comparison to WT plants, aneuploid selection can be very difficult.
Supplementary Figure 4 Recognition of GFP-tailswap seedlings.
Three seedlings are shown, 72 hours after germination on an agar plate. The two plants on the left are haploid or diploid seedlings, showing a long, well developed root. The seedling on the right is a GFP-tailswap seedling resulting from self-pollination. Note the short root and the presence of adventitious roots on the hypocotyl.
Supplementary Figure 5 Phenotype of GFP-tailswap plants.
The GFP-tailswap plants shown were grown in a growth chamber, conditions which result in a curly leaf phenotype. In contrast, when grown in greenhouse conditions GFP-tailswap will resemble more closely WT Col plants. Note that the plants shown were for several weeks grown in a cold growth chamber, leading to relatively large rosettes with many leaves.
Supplementary information
Supplementary Figure 1
Comparison of WT (left) and RNAi:DMC1 transformed achiasmatic line (right) in a CVI background. (PDF 3368 kb)
Supplementary Figure 2
Typical offspring of a cross of a WT F1 to the GFP-tailswap haploid inducer showing GFP-tailswap selfs, haploids, aneuploids and diploid heterozygote offspring between GFP-tailswap and the WT F1. (PDF 24103 kb)
Supplementary Figure 3
Identification of aneuploids. (PDF 21060 kb)
Supplementary Figure 4
Recognition of GFP-tailswap seedlings. (PDF 3030 kb)
Supplementary Figure 5
Phenotype of GFP-tailswap plants. (PDF 2026 kb)
Rights and permissions
About this article
Cite this article
Wijnker, E., Deurhof, L., van de Belt, J. et al. Hybrid recreation by reverse breeding in Arabidopsis thaliana. Nat Protoc 9, 761–772 (2014). https://doi.org/10.1038/nprot.2014.049
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.049
This article is cited by
-
Haploid induction and its application in maize breeding
Molecular Breeding (2021)
-
Reciprocal cybrids reveal how organellar genomes affect plant phenotypes
Nature Plants (2020)
-
State-of-the-art and novel developments of in vivo haploid technologies
Theoretical and Applied Genetics (2019)
-
Cichorium intybus L. × Cicerbita alpina Walbr.: doubled haploid chicory induction and CENH3 characterization
Euphytica (2019)
-
A haploid genetics toolbox for Arabidopsis thaliana
Nature Communications (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.