Abstract
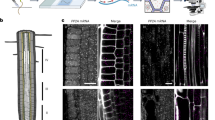
High throughput microarray transcription analyses provide us with the expression profiles for large amounts of plant genes. However, their tissue and cellular resolution is limited. Thus, for detailed functional analysis, it is still necessary to examine the expression pattern of selected candidate genes at a cellular level. Here, we present an in situ mRNA hybridization method that is routinely used for the analysis of plant gene expression patterns. The protocol is optimized for whole mount mRNA localizations in Arabidopsis seedling tissues including embryos, roots, hypocotyls and young primary leaves. It can also be used for comparable tissues in other species. Part of the protocol can also be automated and performed by a liquid handling robot. Here we present a detailed protocol, recommended controls and troubleshooting, along with examples of several applications. The total time to carry out the entire procedure is ∼7 d, depending on the tissue used.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Tautz, D. & Pfeifle, C. A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback. Chromosoma 98, 81–85 (1989).
de Almeid Engler, J., Van Montagu, M. & Engler, G. Whole-mount in situ hybridization in plants. Methods Mol. Biol. 82, 373–384 (1998).
Bennett, M.J. et al. Arabidopsis AUX1 gene: a permease-like regulator of root gravitropism. Science 273, 948–950 (1996).
Ludevid, D., Hofte, H., Himelblau, E. & Chrispeels, M.J. The expression pattern of the tonoplast intrinsic protein gamma-TIP in Arabidopsis thaliana is correlated with cell enlargement. Plant Physiol. 100, 1633–1639 (1992).
Plickert, G., Gajewski, M., Gehrke, G., Gausepohl, H., Schlossherr, J. & Ibrahim, H. Automated in situ detection (AISD) of biomolecules. Dev. Genes Evol. 207, 362–367 (1997).
Nonradioactive In Situ hybridization Application Manual, 2nd edition, 51–56 (Boehringer Manheim 1996).
Friml, J., Benková, E., Mayer, U., Palme, K. & Muster, G. Automated whole-mount localization techniques for plant seedlings. Plant J. 34, 115–124 (2003).
García-Aguilar, M., Dorantes-Acosta, A., Pérez-España, V. & Vielle-Calzada, J.P. Whole-mount in situ mRNA localization in developing ovules and seeds of Arabidopsis. Plant Mol. Biol. Reporter 23, 279–289 (2005).
Zachgo, S., Perbal, M.C., Saedler, H. & Schwarz-Sommer, Z. In situ analysis of RNA and protein expression in whole mounts facilitates detection of floral gene expression dynamics. Plant J. 23, 697–702 (2000).
Fisher, H.W. & Williams, R.C. Electron microscopic visualization of nucleic acids and of their complexes with proteins. Annu. Rev. Biochem. 48, 649–679 (1979).
Malamy, J.E. & Benfey, P.N. Organization and cell differentiation in lateral roots of Arabidopsis thaliana. Development, 124, 33–44 (1997).
Hamann, T., Benková, E., Bäurle, I., Kientz, M. & Jürgens, G. The Arabidopsis BODENLOS gene encodes an auxin response protein inhibiting MONOPTEROS-mediated embryo patterning. Genes & Dev. 16, 1610–1615 (2002).
Friml, J. et al. AtPIN4 mediates sink driven auxin gradients and patterning in Arabidopsis roots. Cell, 108, 661–673 (2002).
Blilou, I. et al. The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature, 433, 39–44 (2005).
Acknowledgements
J.H. was supported by the Ministry of Education of the Czech Republic (LC06034, MSM0021622415); P.B. and J.F. by the VolkswagenStiftung and the EMBO Young Investigator Program; E.B. by the Margarete von Wrangell-Habilitations program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Hejátko, J., Blilou, I., Brewer, P. et al. In situ hybridization technique for mRNA detection in whole mount Arabidopsis samples. Nat Protoc 1, 1939–1946 (2006). https://doi.org/10.1038/nprot.2006.333
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.333
This article is cited by
-
A rapid and sensitive, multiplex, whole mount RNA fluorescence in situ hybridization and immunohistochemistry protocol
Plant Methods (2023)
-
Whole-mount RNA in situ hybridization technique in Torenia ovules
Plant Reproduction (2023)
-
Specification of female germline by microRNA orchestrated auxin signaling in Arabidopsis
Nature Communications (2022)
-
Whole mount in situ localization of miRNAs and target mRNA transcripts in plants
3 Biotech (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.