Abstract
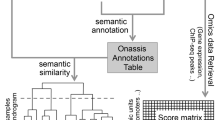
Our work concerns the elucidation of the cancer (epi)genome, transcriptome and proteome to better understand the complex interplay between a cancel cell's molecular state and its response to anti-cancer therapy. To study the problem, we have previously focused on data warehousing technologies and statistical data integration. In this paper, we present recent work on extending our analytical capabilities using Semantic Web technology. A key new component presented here is a SPARQL endpoint to our existing data warehouse. This endpoint allows the merging of observed quantitative data with existing data from semantic knowledge sources such as Gene Ontology (GO). We show how such variegated quantitative and functional data can be integrated and accessed in a universal manner using Semantic Web tools. We also demonstrate how Description Lobic (DL) reasoning can be used to infer previously unstated conclusions from existing knowledge bases. As proof of concept, we illustrate the ability of our setup to answer complex queries on resistance of cancer cells to Decitabine, a demethylating agent.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Holford, M., McCusker, J., Cheung, K. et al. Analysis of Cancer Omics Data In A Semantic Web Framework. Nat Prec (2010). https://doi.org/10.1038/npre.2010.5391.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2010.5391.1