Abstract
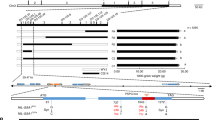
Grain size is a major determinant of grain yield in cereal crops. qSW5/GW5, which exerts the greatest effect on rice grain width and weight, was fine-mapped to a 2,263-bp/21-kb genomic region containing a 1,212-bp deletion, respectively. Here, we show that a gene encoding a calmodulin binding protein, located ∼5 kb downstream of the 1,212-bp deletion, corresponds to qSW5/GW5. GW5 is expressed in various rice organs, with highest expression level detected in young panicles. We provide evidence that the 1,212-bp deletion affects grain width most likely through influencing the expression levels of GW5. GW5 protein is localized to the plasma membrane and can physically interact with and repress the kinase activity of rice GSK2 (glycogen synthase kinase 2), a homologue of Arabidopsis BIN2 (BRASSINOSTEROID INSENSITIVE2) kinase, resulting in accumulation of unphosphorylated OsBZR1 (Oryza sativa BRASSINAZOLE RESISTANT1) and DLT (DWARF AND LOW-TILLERING) proteins in the nucleus to mediate brassinosteroid (BR)-responsive gene expression and growth responses (including grain width and weight). Our results suggest that GW5 is a novel positive regulator of BR signalling and a viable target for genetic manipulation to improve grain yield in rice and perhaps in other cereal crops as well.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sweeney, M. & McCouch, S. The complex history of the domestication of rice. Ann. Bot. 100, 951–957 (2007).
Zuo, J. & Li, J. Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu. Rev. Genet. 48, 99–118 (2014).
Zhang, C., Bai, M. Y. & Chong, K. Brassinosteroid-mediated regulation of agronomic traits in rice. Plant Cell Rep. 33, 683–696 (2014).
Hu, X. et al. The U-box E3 ubiquitin ligase TUD1 functions with a heterotrimeric G α subunit to regulate brassinosteroid-mediated growth in rice. PLoS Genet. 9, e1003391 (2013).
Weng, J. et al. Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res. 18, 1199–1209 (2008).
Shomura, A. et al. Deletion in a gene associated with grain size increased yields during rice domestication. Nat. Genet. 40, 1023–1028 (2008).
Wang, X. et al. Trans-golgi network-located AP1 gamma adaptins mediate dileucine motif-directed vacuolar targeting in Arabidopsis. Plant Cell 26, 4102–4118 (2014).
Chen, X., Huang, H. & Xu, L. The CaMV 35S enhancer has a function to change the histone modification state at insertion loci in Arabidopsis thaliana. J. Plant Res. 126, 841–846 (2013).
Benfey, P. N., Ren, L. & Chua, N. H. The CaMV 35S enhancer contains at least two domains which can confer different developmental and tissue-specific expression patterns. EMBO J. 8, 2195–2202 (1989).
Hudson, R. R., Kreitman, M. & Aguadé, M. A test of neutral molecular evolution based on nucleotide data. Genetics 116, 153–159 (1987).
Zhu, Q., Zheng, X., Luo, J., Gaut, B. S. & Ge, S. Multilocus analysis of nucleotide variation of Oryza sativa and its wild relatives: severe bottleneck during domestication of rice. Mol. Biol. Evol. 24, 875–888 (2007).
Liu, L. et al. KRN4 controls quantitative variation in maize kernel row number. PLoS Genet. 11, e1005670 (2015).
Ishii, T. et al. OsLG1 regulates a closed panicle trait in domesticated rice. Nat. Genet. 45, 462–465 (2013).
Tong, H. et al. DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell 24, 2562–2577 (2012).
Sun, S. et al. Brassinosteroid signaling regulates leaf erectness in Oryza sativa via the control of a specific U-type cyclin and cell proliferation. Dev. Cell 34, 220–228 (2015).
He, J. X., Gendron, J. M., Yang, Y., Li, J. & Wang, Z. Y. The GSK3-like kinase BIN2 phosphorylates and destabilizes BZR1, a positive regulator of the brassinosteroid signaling pathway in Arabidopsis. Proc. Natl Acad. Sci. USA 99, 10185–10190 (2002).
Duan, P. et al. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat. Plants 2, 15203 (2015).
Hu, J. et al. A rare allele of GS2 enhances grain size and grain yield in rice. Mol. Plant 8, 1455–1465 (2015).
Che, R. et al. Control of grain size and rice yield by GL2-mediated brassinosteroid responses. Nat. Plants 2, 15195 (2015).
Lei, Y. et al. CRISPR-P: a web tool for synthetic single-guide RNA design of CRISPR-system in plants. Mol. Plant 7, 1494–1496 (2014).
Jefferson, R. The GUS reporter gene system. Nature 342, 837–838 (1989).
Bradbury, P. J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Rozas, J., Sanchez-DelBarrio, J. C., Messeguer, X. & Rozas, R. DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19, 2496–2497 (2003).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Waadt, R. & Kudla, J. In planta visualization of protein interactions using bimolecular fluorescence complementation (BiFC). CSH Protoc. http://dx.doi.org/10.1101/pdb.prot4995 (2008).
Lin, Q. et al. The SnRK2-APC/CTE regulatory module mediates the antagonistic action of gibberellic acid and abscisic acid pathways. Nat. Commun. 6, 7981 (2015).
Miernyk, J. A. & Thelen, J. J. Biochemical approaches for discovering protein–protein interactions. Plant J. 53, 597–609 (2008).
Wada, K., Marumo, S., Ikekawa, N., Morisaki, M. & Mori, K. Brassinolide and homobrassinolide promotion of lamina inclination of rice seedlings. Plant Cell Physiol. 22, 323–325 (1981).
Acknowledgements
We thank C. Chu (Chinese Academy of Sciences) for kindly providing the Go and Gi transgenic seeds; we thank X.W. Deng (Peking University), C. Wu (Chinese Academy of Agricultural Sciences) and J. Zhou (Frontier Laboratories of Systems Crop Design) for technical support. This work was supported by the National Natural Science Foundation of China (No. 91535302 and No. 31430008), National Key Research and Development Plan (2016YFD0100400) and National Transformation Science and Technology Program (2014ZX08001006).
Author information
Authors and Affiliations
Contributions
J.L. and J.C. performed most of the experiments; X.Zheng performed the analysis of molecular evolution; F.W., Q.L, Y.H., P.T., Z.C. and X.Y. provided technical assistance; K.Z. performed some of the sub-cellular localization assay; X.Zhang and X.G. generated the transgenic plants; J.Wang cultivated the transgenic plants in the field; J.Wan and H.W. supervised the project; J.Wan, H.W. and J.L. designed the research and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–22, Supplementary Tables 1 and 3, Legend for Supplementary Table 2. (PDF 5383 kb)
Supplementary Table 2
List of SNP's in 6.3-kb upstream region of GW5. (XLSX 132 kb)
Rights and permissions
About this article
Cite this article
Liu, J., Chen, J., Zheng, X. et al. GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nature Plants 3, 17043 (2017). https://doi.org/10.1038/nplants.2017.43
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2017.43
This article is cited by
-
MADS1-regulated lemma and awn development benefits barley yield
Nature Communications (2024)
-
Are cereal grasses a single genetic system?
Nature Plants (2024)
-
Identification of qGL4.1 and qGL4.2, two closely linked QTL controlling grain length in rice
Molecular Breeding (2024)
-
Dissection and validation of quantitative trait loci (QTLs) conferring grain size and grain weight in rice
Euphytica (2024)
-
Exploratory biological coordinate analysis on 2,4-epi-brassinolide-induced the size change of leaf angle in tobacco seedlings
Plant Growth Regulation (2024)