Abstract
Jute (Corchorus sp.) is one of the most important sources of natural fibre, covering ∼80% of global bast fibre production1. Only Corchorus olitorius and Corchorus capsularis are commercially cultivated, though there are more than 100 Corchorus species2 in the Malvaceae family. Here we describe high-quality draft genomes of these two species and their comparisons at the functional genomics level to support tailor-designed breeding. The assemblies cover 91.6% and 82.2% of the estimated genome sizes for C. olitorius and C. capsularis, respectively. In total, 37,031 C. olitorius and 30,096 C. capsularis genes are identified, and most of the genes are validated by cDNA and RNA-seq data. Analyses of clustered gene families and gene collinearity show that jute underwent shared whole-genome duplication ∼18.66 million years (Myr) ago prior to speciation. RNA expression analysis from isolated fibre cells reveals the key regulatory and structural genes involved in fibre formation. This work expands our understanding of the molecular basis of fibre formation laying the foundation for the genetic improvement of jute.
Similar content being viewed by others
Bast (phloem) fibres are obtained from the stem of the plants such as jute, flax, hemp, ramie and kenaf. The annual global production of jute generates a farm value of ∼US$2.3 billion1. The cultivated species of jute, C. olitorius and C. capsularis, are morphologically and physiologically distinct (Supplementary Fig. 1), and a combination of useful traits from these species into a single genotype is highly desirable3. However, interspecific hybridization is limited because of their cross-incompatibility4,5. To facilitate comparative functional genomics and to understand the molecular basis of bast fibre biogenesis, genomes of two popular jute cultivars C. olitorius var. O-4 and C. capsularis var. CVL-1 are sequenced and analysed.
We performed whole-genome shotgun (WGS) sequencing with the Roche/454 platform (Supplementary Table 1) and assembled the genomes using CABOG6. The resulting assemblies were 445 Mb (scaffold N50 length, 3.3 Mb; longest scaffold, 45.5 Mb) for C. olitorius and 338 Mb (scaffold N50 length, 4.1 Mb; longest scaffold, 28.5 Mb) for C. capsularis (Table 1 and Supplementary Table 2). Eighty per cent of the C. olitorius and C. capsularis assemblies were covered with 415 scaffolds (minimum length 76 kb) and 231 scaffolds (minimum length 120 kb), respectively. We estimated the genome sizes for C. olitorius and C. capsularis to be ∼448 and ∼404 Mb (Supplementary Information and Supplementary Fig. 2), respectively, which were consistent with reported estimates7. Whole-genome optical mapping was used to improve the assemblies, resulting increase in N50 of the scaffolds to 4.0 Mb for C. olitorius and 8.5 Mb for C. capsularis (Supplementary Information and Supplementary Tables 3–6). We anchored ∼60% of each assembly to seven genetic linkage groups using a set of 1,389 molecular markers from a consensus genetic linkage map8–12 (Supplementary Fig. 3 and Supplementary Table 7). More than 99% (C. olitorius) and 97% (C. capsularis) of the isotigs generated from transcriptome sequencing of jute seedlings aligned to the respective genomes indicate comprehensive coverage of the gene-rich regions (Supplementary Tables 8 and 9). In addition, more than 97% of the conserved core eukaryotic genes13 were present in each of the draft genomes (Supplementary Table 10). Moreover, the single-base accuracy of the de novo assembled genomes was evaluated by mapping all reads onto the scaffolds using a CLC mapper (CLC bio, Aarhus, Denmark). It was observed that 82.29% and 78.28% of the reads are uniquely mapped to C. olitorius and C. capsularis, respectively (Supplementary Table 11). We predicted 37,031 C. olitorius and 30,096 C. capsularis protein-coding genes by using a combination of de novo, homology and transcriptome-based approaches (Table 1; Supplementary Fig. 4 and Supplementary Table 12). The averages of gene length, number of exons and coding sequence length of C. olitorius are 2,359 bp, 4.00 and 927 bp respectively and for C. capsularis are 2,759 bp, 4.58 and 1,036 bp respectively (Supplementary Table 13). These values are similar to other malvaceous crops such as cotton14 and cacao15. We also identified 1,010 and 666 microRNAs (miRNAs), 488 and 203 transfer RNAs (tRNAs) and 80 and 110 rRNAs in the C. olitorius and C. capsularis genomes, respectively (Table 1; Supplementary Tables 14–16). More than 50% of the C. olitorius and C. capsularis genomes were composed of repetitive elements, which is similar to cotton (∼57%) and double that of cacao (∼24%) (Supplementary Tables 17 and 18). The proportions of various types of repeats were similar in both genomes, with long terminal repeats being the most abundant (Supplementary Table 19) which is similar to that observed in Bamboo16 and Banana17.
We examined the evolutionary relationship between jute and 13 other sequenced plant genomes with representatives from the Malvids (cacao, cotton and Arabidopsis), Fabids (castor bean, flax, Medicago, soybean, Fragaria and Populus), Asterids (tomato and potato), Vitales (grape) and Monocots (rice). Phylogenetic analysis, based on a concatenated alignment of 13 single-copy gene families from 15 sequenced plant genomes, supported the placement of jute with cacao and cotton in the Malvaceae family (Fig. 1a). This inclusion is consistent with the analysis done with chloroplast DNA sequences18. All protein-coding genes from 15 genomes (Supplementary Table 20) were clustered into 47,186 gene families (two or more members), of which 8,177 were common to all five groups and 8,816 were confined to the Malvids (Supplementary Fig. 5). Among the Malvids-specific gene families only 613 and 152 are unique to C. olitorius and C. capsularis, respectively (Fig. 1b). These jute-specific gene families were significantly enriched with genes related to the oxidation–reduction process, transcription factors, transposases and defence-related proteins (Supplementary Tables 21 and 22). To investigate the expression of jute-specific genes in fibre cells, the RNA-seq data from isolated fibre cells and seedlings of C. olitorius and C. capsularis were compared. Among the jute-specific genes, Myb/SANT-like domain, Zinc finger, PHD-type, F-box domain cyclin-like proteins are highly expressed in fibre cells than seedlings (Supplementary Tables 23 and 24) indicating their involvement in bast fibre formation.
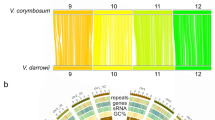
a, Phylogenetic analysis showing positions of C. olitorius and C. capsularis within the Malvids. b, Venn diagram of unique and shared gene families among five Malvids genomes. The analysis was performed with only the Malvids-specific gene families (8,816), as determined in Supplementary Fig. 5. c, Ks distributions of all paralogous gene pairs in the C. olitorius and C. capsularis genomes. The y-axis shows the percentage of the two-member gene clusters. Myr, million years. d, Syntenic blocks shared between the C. olitorius and C. capsularis linkage groups (LG).
The natural genetic diversity within the jute species is very narrow19,20 and it is one of the impediments for the breeder to develop high-yield and quality varieties. The extent of gene duplications in the C. olitorius and C. capsularis genomes were examined. By calculating the synonymous substitution rates (Ks) for paralogous gene pairs, two peaks at 0.24–0.32 and 0.72–0.92 for both of the genomes were found (Fig. 1c). The first peak reveals the whole-genome duplication (WGD) event occurred ∼18.6 (16.0–21.3) Myr ago prior to their separation at ∼6 Myr ago (Supplementary Fig. 6). The second peak is indicating an ancient WGD event occurred in jute ∼129.2 (110.7–141.5) Myr ago (Fig. 1c). The ancient duplication event corresponds to the ancient hexaploidization that is shared among the eudicots21. Comparison of the two genomes revealed that they share 160 syntenic blocks (five or more genes per block) with the linkage groups covering 58% and 65% of the assembled genome of C. olitorius and C. capsularis, respectively (Fig. 1d; Supplementary Fig. 7 and Supplementary Tables 25 and 26). It indicates that extensive synteny and conserved gene order exists between the genomes. A one-to-one relationship of the predominantly aligned syntenic regions denotes no WGD after speciation (Supplementary Fig. 8). The occurrence of tandem duplications, which tend to be biased towards genes involved in responses to environmental stimuli22, was relatively low for C. olitorius (7.2% of total genes) and C. capsularis (5.9% of total genes) (Supplementary Table 27) compared with other plant genomes23.
The genomics information of jute fibre biogenesis is merely available for the improvement of its yield and quality. RNA-seq data obtained from isolated fibre cells (elongated cells undergoing secondary cell wall (SCW) deposition) and seedlings of C. olitorius and C. capsularis were analysed to investigate the molecular events of jute fibre development (Supplementary Fig. 9 and Supplementary Tables 28 and 29). We identified 6,077 upregulated and 6,809 downregulated genes for C. olitorius and 7,695 upregulated and 7,809 downregulated genes for C. capsularis (Supplementary Fig. 10 and Supplementary Tables 30 and 31). Among them, 329 C. olitorius and 344 C. capsularis genes were identified based on the analysis of homologous genes reportedly involved in plant fibre formation24 which facilitated us to propose a model for bast fibre biogenesis in jute (Fig. 2a). It was found that 174 (53%) C. olitorius and 216 (63%) C. capsularis genes were expressed significantly within the fibre cells and seedlings (Supplementary Table 32). Genes encoding the WOX4, APL and HAT22 transcription factors and the TDIF signalling peptide, which are involved in vascular cambium initiation and proliferation25–28, were highly expressed in fibre cells, suggesting their importance in fibre differentiation. Moreover, several of the transcription factor genes involved in regulating SCW formation exhibited higher expression in the fibre cells (Supplementary Fig. 11). In particular, a substantial increase in expression was observed for the MYB83 homologue of Arabidopsis, a master regulator capable of activating the biosynthesis of all major SCW components (cellulose, lignin and xylan)29. The homologue of AtMYB46, which is co-expressed and functionally redundant with AtMYB83 in Arabidopsis29, showed little or no expression in jute fibres indicating that MYB83 is primarily accountable for the SCW regulatory network of jute.
a, Schematic representation of the fibre formation process in jute. Fibre formation-related genes are listed in Supplementary Table 32. b,c, Heat map showing relative expression of genes involved in the biosynthesis of lignin (b) and cellulose (c) between 4-day-old seedlings and fibre cells. A similar expression pattern was also observed between very young seedlings before bolting and fibre cells (Supplementary Fig. 12a1,a2). The heat maps of normalized RNA-seq data was prepared from three biological replicates from fibre cells and whole seedlings of C. olitorius and C. capsularis. Gene expression was measured by quantified transcription levels (fragments per kilobase of exon model per million mapped reads, FPKM) derived from RNA-seq analysis. Heat scale, log2(FPKM). To calculate the log2(FPKM) values of individual genes, all of the original FPKM values were added by a pseudo-count of 1. CH4, cinnamate-4-hydroxylase; PAL, L-phenylalanine ammonia-lyase; HCT, hydroxycinnamoyl transferase; C3H, 4-coumarate 3-hydroxylase; F5H, ferulate-5-hydroxylase; CAD, cinnamyl alcohol dehydrogenase.
The relatively high lignin content (∼15%) in jute fibre makes it coarser than other bast fibres such as flax and ramie (<5% lignin)30. Among the lignin biosynthetic genes detected in the C. olitorius and C. capsularis genomes, there were expansions of the 4-coumarate:CoA ligase (4CL), cinnamoyl-CoA reductase (CCR), trans-caffeoyl-CoA 3-O-methyltransferase (CCoAOMT) and caffeic acid O-methyltransferase (COMT) gene families compared with flax (Supplementary Table 33). The expression profiles of the lignification genes reveal that only a few homologues appeared to be preferentially expressed at high levels in the fibre cells for most of the gene families (Fig. 2b; Supplementary Table 32 and Supplementary Fig. 12a1), highlighting possible targets for engineering low-lignin jute fibres. Cellulose, synthesized by the cellulose synthase (CesA) complex, makes up the majority of the SCW in jute fibres (∼60%). We identified 10 CesA and 32 cellulose synthase-like (Csl) genes, similar to several other plants (Supplementary Table 34). SCW synthesis specific genes CesA4 and CesA7 were distinctly upregulated in fibre cells (Fig. 2c; Supplementary Table 32 and Supplementary Fig. 12a2) indicating their association with SCW cellulose deposition, whereas significant expression of CesA1, CesA3 and CesA6 in seedlings suggest their involvement in primary cell wall cellulose deposition (Fig. 2c; Supplementary Table 32 and Supplementary Fig. 12a2).
Senescence and cell death are the final phases of fibre biogenesis related to autophagy and proteolysis pathways. KEGG pathway enrichment analysis with fibre cell transcriptome data indicated an upregulation of autophagy and proteolysis pathways whereas most of the metabolic pathways were downregulated (Supplementary Tables 35 and 36). In flax phloem and poplar xylem fibres, a gradual degradation of the nucleus and cytoplasm is likely to be mediated by autophagy while deposition of the SCW continues31,32. In poplar xylem fibres32, all copies of ATG8 were upregulated in jute fibre cells and the expressions were the highest among the autophagy-related genes (Supplementary Table 32) suggesting a similar cell death programme.
RNA-seq results for fibre biogenesis pathway genes were validated with quantitative polymerase chain reaction with reverse transcription (RT–qPCR) on randomly selected genes (Supplementary Fig. 13). For most of the genes, similar upregulation and downregulation patterns were observed.
To explain the morphological and physiological differences between the species, we focused on comparing the transcripts and genes that are related to lignin deposition in SCW, fibre colour and response to abiotic stress. As C. olitorius fibre contains more lignin and less cellulose than that of C. capsularis (Supplementary Table 37)33, expression patterns of lignin and cellulose biosynthesis genes between them were different (Fig. 2b,c). Genes encoding most of the key enzymes for proanthocyanidin biosynthesis were highly expressed in the fibres of C. olitorius (golden colour) than in C. capsularis (whitish colour) (Supplementary Fig. 14), indicating their involvement for the differences in fibre pigmentation.
C. capsularis is comparatively tolerant to flood and drought but slightly more susceptible to diseases and pests than C. olitorius34. Moreover, C. capsularis is somewhat tolerant to salt stress compared with C. olitorius and can survive up to 60 mM concentrations of NaCl in nutrient media (Supplementary Fig. 15). We categorized gene ontology (GO) using the Blast2GO pipeline35 to identify gene copy number variation which is a major mechanism of phenotypic differentiation and evolutionary adaptation to the environment36,37 (Supplementary Tables 38 and 39). In the GO class ‘biological processes’, genes with GO terms associated with response to important environmental factors including salt and osmotic stress were over-represented in C. capsularis (Fig. 3a; Supplementary Table 40). Besides, in the GO class ‘molecular function’, we found that genes responding to abiotic stress were also significantly over-represented in C. capsularis (Fig. 3b; Supplementary Table 40). Among them transmembrane transport, vacuolar transport, homeostatic process, ATPase activity, transmembrane transporter activity, signal transducer activity, oxidoreductase activity were significantly higher in C. capsularis, indicating its adaptability in different habitats and environmental pressure. For example, transmembrane transport proteins mediate ion fluxes, including sodium ATPase, vacuolar H+-ATPase, chloride channel protein, and ABC transporters play important roles in ionic and osmotic homeostasis under salt environments38. To corroborate GO analysis results, correlated genes were identified (Supplementary Table 41) and most of them are associated with abiotic stress tolerance.
a, GO biological processes. b, GO molecular functions. The list of gene numbers in each of the GO categories and their associated P values are listed in Supplementary Table 40. The GO categories are as defined in Blast2GO. Categories with significant differences calculated using a chi-squared test, as described in the Supplementary Information. The complete list of GO categories for C. olitorius and C. capsularis genome are listed in Supplementary Tables 38 and 39. NS, non-significant. The remainder of the categories are significant at the 1% level. ORFs, open reading frames.
The comparative genome analysis opens up opportunities for the development of improved breeding strategies to meet the increasing demand of natural fibres in industry. The genome sequences provide a valuable resource to advance our understanding of bast fibre biogenesis in jute, thus serving as the foundation of genetic improvement for productivity and fibre quality.
Methods
Genome sequencing and assembly
The genomes of C. olitorius var. O-4 and C. capsularis var. CVL-1 were sequenced using the WGS approach on the 454 platform. A total of 13.04 Gb of sequence data was generated for the C. olitorius genome, consisting of 5.65 Gb of shotgun sequences, 2.56 Gb of 3 kb paired-end sequences, 2.47 Gb of 8 kb paired-end sequences and 2.36 Gb of 20 kb paired-end sequences. For the C. capsularis genome, 13.69 Gb of sequence data was generated, consisting of 7.87 Gb of shotgun sequences, 2.04 Gb of 3 kb paired-end sequences, 2.26 Gb of 8 kb paired-end sequences and 1.51 Gb of 20 kb paired-end sequences (Supplementary Table 1).
We used the CABOG tool sffToCA to identify mated reads and remove duplicate mate pairs. The remaining read sequence data were converted to the fastq format, trimmed to a length of 65 bases and used in the assembly. The CABOG v7.0 pipelines were then run with default parameters using a kmer parameter of 22, which was selected after testing a range of kmer settings.
We used whole-genome optical mapping technology to improve and validate the assemblies (Supplementary Table 3). A total of 360,906 and 260,615 single-molecule restriction maps longer than 250 kb each, with an average size of 356.37 and 356.99 kb, were generated using the KpnI restriction enzyme for C. olitorius and C. capsularis, respectively (Supplementary Table 4). Super-scaffolding with optical map data was performed using Genome-Builder software (OpGen). Super-scaffolds and scaffolds were anchored to seven linkage groups using a combination of traditional markers and whole-genome mapping data using ALLMAPS software.
The accuracy and completeness of the assemblies were assessed by aligning isotigs that were generated from transcriptome sequencing onto the WGS scaffolds using BLAT (Supplementary Table 9). We also checked the relative completeness of the assemblies by performing core gene annotation using the CEGMA v2.5 pipelines (Supplementary Table 10).
Gene annotation
Repetitive elements were identified and masked by RepeatModeler v1.0.7 and RepeatMasker Open-3.0 with default parameters. Gene prediction was performed using a combination of homology, de novo and transcript-based approaches (Supplementary Fig. 4). The predicted genes were analysed for functional domains and homologies by using InterProScan and Basic Local Alignment Search Tool (BLAST) against the NCBI non-redundant database, TrEMBL and SwissProt with Protein BLAST (BLASTP) (e < 1 × 10−5) and Blast2GO v3.3 with default parameters.
Genome comparison and evolution
Comparative analysis was performed to identify orthologous gene families among the genomes of C. olitorius, C. capsularis, Arabidopsis thaliana, Theobroma cacao, Gossypium raimondii, Glycine max, Populus trichocarpa, Ricinus communis, Fragaria vesca, Linum usitatissimum, Medicago truncatula, Vitis vinifera, Solanum lycopersicum, Solanum tuberosum and Oryza sativa. All predicted protein sequences of these plants (Supplementary Table 20) were searched against each other using BLASTP (e < 1 × 10−5) and clustered into orthologous groups using MCL-10-201 (inflation parameter, 1.5). Clusters containing single-copy orthologues were identified with exact one member per species. Phylogenetic relationships were determined from these single-copy orthologues using the maximum likelihood method.
Paralogous genes in C. olitorius and C. capsularis were detected by all-against-all protein sequence similarity searches using BLASTP. The synonymous substitution rate (Ks) was calculated for each gene pair. Paralogous genes were determined to be tandem duplicates if they were located within five genes from each other. Orthologous genes between C. olitorius and C. capsularis were identified using the reciprocal best hit method and the Ks values were calculated for each pair. Intra- and intergenomic regions of synteny were identified and visualized by SyMAP v4.0.
Pathway reconstruction
Metabolic and regulatory pathways were reconstructed with Pathway Studio software based on the Resnet-Plant 4.0 database. Predicted jute interologues and pathways were imported into a new Pathway Studio database for manual pathway reconstruction and genome analysis.
Fibre cell transcriptome sequencing and analysis
The transcriptomes of isolated fibre cells and whole seedlings were sequenced with an Illumina HiSeq 2,500 at HudsonAlpha Institute for Biotechnology, Huntsville, Alabama (Supplementary Tables 28 and 29). Expression patterns were compared by aligning the RNA-seq reads against the C. olitorius and C. capsularis genome sequences and quantifying the transcript abundances using the Cufflinks v2.2.1 package, which was visualized by R libraries. KEGG Orthology Based Annotation System (KOBAS) was used to identify the pathways in the C. olitorius and the C. capsularis genome using the model organism A. thaliana. KEGG (Release 74.0) and Biocyc v19.0 pathways were utilized to run R package piano v1.8.0 for Gene set analysis (GSA). Pathways in the distinct direction were selected for subsequent analysis based on adjusted P < 0.05. The differential gene expression from the in silico analysis were validated by RT–qPCR with randomly selected several fibre biosynthesis pathway genes. All primers used in this study are provided in Supplementary Table 42.
Statistical analyses
Two-tailed chi-squared tests were used to compare the distributions of GO subcategories between C. olitorius and C. capsularis (Fig. 3). For each GO subcategory, a 2 × 2 contingency table was constructed by recording the existence of the number of genes in a subcategory for each species and ranking the statistical significance of the differences.
Detailed methods and their associated references are in the Supplementary Information.
Data availability
The WGS projects have been deposited at NCBI GenBank under BioProject ID PRJNA215141 and accession no. AWUE00000000 for C. olitorius and BioProject ID PRJNA215142 and accession no. AWWV00000000 for C. capsularis. The genomic and transcriptomic raw data have been deposited in the NCBI Sequence Read Archive (SRA) under SRP049494 and SRP053213 for C. olitorius and C. capsularis, respectively.
Additional information
How to cite this article: Islam, M. S. et al. Comparative genomics of two jute species and insight into fibre biogenesis. Nat. Plants 3, 16223 (2017).
References
Statistical Bulletin-2014 (Food and Agriculture Organization of the United Nations, 2014).
Saunders, M. Recovery Plan for the Endangered Native Jute Species, Corchorus cunninghamii F. Muell in Queensland (2001–2006) (Natural Heritage Trust, 2006).
Mir, J. I. et al. SSR and RAPD profile based grouping of selected jute germplasm with respect to fibre fineness trait. J. Plant Biochem. Biotechnol. 17, 29–35 (2008).
Patel, G. I. & Datta, R. M. Interspecific hybridization between Corchorus capsularis L. and C. olitorius L. and the cytological basis of incompatibility between them. Euphytica 9, 89–110 (1960).
Swaminathan, M. S., Iyer, R. D. & Sulbha, K. Morphology, cytology and breeding behaviour of hybrids between Corchorus olitorius and C. capsularis. Curr. Sci. 30, 67–68 (1961).
Miller, J. R. et al. Aggressive assembly of pyrosequencing reads with mates. Bioinformatics 24, 2818–2824 (2008).
Benor, S., Fuschs, J. & Blattner, F. R. Genome size variation in Corchorus olitorius (Malvaceae s.l.) and its correlation with elevation and phenotypic traits. Genome 54, 575–585 (2011).
Joshi, A. et al. Chromosome-specific physical localisation of expressed sequence tag loci in Corchorus olitorius L. Plant Biol. 16, 1133–1139 (2014).
Topdar, N. et al. A complete genetic linkage map and QTL analyses for bast fibre quality traits, yield and yield components in jute (Corchorus olitorius L.). Cytol. Genet. 47, 129–137 (2013).
Das, M. et al. Development of SSR markers and construction of a linkage map in jute. J. Genet. 91, 21–31 (2012).
Biswas, C., Dey, P., Karmakar, P. G. & Satpathy, S. Discovery of large-scale SNP markers and construction of linkage map in a RIL population of jute (Corchorus capsularis). Mol. Breed. 35, 1–10 (2015).
Kundu, A. et al. A restriction-site-associated DNA (RAD) linkage map, comparative genomics and identification of QTL for histological fibre content coincident with those for retted bast fibre yield and its major components in jute (Corchorus olitorius L., Malvaceae s.l.). Mol. Breed. 35, 19 (2015).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Wang, K. et al. The draft genome of a diploid cotton Gossypium raimondii. Nat. Genet. 44, 1098–1103 (2012).
Argout, X. et al. The genome of Theobroma cacao. Nat. Genet. 43, 101–108 (2011).
Peng, Z. et al. The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla). Nat. Genet. 45, 456–461 (2013).
D'Hont, A. et al. The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488, 213–217 (2012).
Whitlock, B. A., Karol, K. G. & Alverson, W. S. Chloroplast DNA sequences confirm the placement of the enigmatic Oceanopapaver within Corchorus (Grewioideae: Malvaceae s.l., formerly Tiliaceae). Int. J. Plant Sci. 164, 35–41 (2003).
Palve, S. M. & Sinha, M. K. Genetic variation and interrelationships among fibre yield attributes in secondary gene pool of Corchorus spp. SABRAO J. Breed. Genet. 37, 55–64 (2005).
Basu, A. et al. Analysis of genetic diversity in cultivated jute determined by means of SSR markers and AFLP profiling. Crop Sci. 44, 678–685 (2004).
Tang, H. et al. Unraveling ancient hexaploidy through multiple-aligned angiosperm gene maps. Genome Res. 18, 1944–1954 (2008).
Hanada, K., Zou, C., Lehti-Shiu, M. D., Shinozaki, K. & Shiu, S. H. Importance of lineage-specific expansion of plant tandem duplicates in the adaptive response to environmental stimuli. Plant Physiol. 148, 993–1003 (2008).
Myburg, A. A. et al. The genome of Eucalyptus grandis. Nature 510, 356–362 (2014).
Gorshkova, T. et al. Plant fiber formation: state of the art, recent and expected progress, and open questions. Crit. Rev. Plant Sci. 31, 201–228 (2012).
Ji, J. et al. WOX4 promotes procambial development. Plant Physiol. 152, 1346–1356 (2010).
Bonke, M., Thitamadee, S., Mähönen, A. P., Hauser, M. T. & Helariutta, Y. APL regulates vascular tissue identity in Arabidopsis. Nature 426, 181–186 (2003).
Tornero, P., Conejero, V. & Vera, P. Phloem-specific expression of a plant homeobox gene during secondary phases of vascular development. Plant J. 9, 639–648 (1996).
Hirakawa, Y., Kondo, Y. & Fukuda, H. Establishment and maintenance of vascular cell communities through local signaling. Curr. Opin. Plant Biol. 14, 1–7 (2010).
McCarthy, R. L., Zhong, R. & Ye, Z. H. MYB83 is a direct target of SND1 and acts redundantly with MYB46 in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell Physiol. 50, 1950–1964 (2009).
Urquhart, A. R. & Howitt, F. O. The Structure of Textile Fibres (Textile Institute, 1953).
Ageeva, M. V., Chernova, T. E. & Gorshkova, T. A. Processes of protoplast senescence and death in flax fibers: an ultrastructural analysis. Russ. J. Dev. Biol. 43, 94–100 (2012).
Courtois-Moreau, C. L. et al. A unique program for cell death in xylem fibers of Populus stem. Plant J. 58, 260–274 (2009).
Sur, D. & Amin, M. N. in Jute Basics (ed. Sur, D. ) 35–55 (International Jute Study Group, 2010).
Roy, A. et al. Evaluation of genetic diversity in jute (Corchorus species) using STMS, ISSR and RAPD markers. Plant Breed. 125, 292–297 (2006).
Conesa, A. & Götz, S. Blast2GO: a comprehensive suite for functional analysis in plant genomics. Intl. J. Plant Genomics 2008, 619832 (2008).
Dassanayake, M. et al. Transcription strength and halophytic lifestyle. Trends Plant Sci. 16, 1–3 (2011).
Hastings, P. J. et al. Mechanisms of change in gene copy number. Nat. Rev. Genet. 10, 551–564 (2009).
Wang, X. et al. Proteomic analysis of the response to high-salinity stress in Physcomitrella patens. Planta 228, 167–177 (2008).
Acknowledgements
This paper is dedicated to A. Shamsul Islam, in memory of his contributions to biotechnological research on jute. We thank C. Detter (US Defense Threat Reduction Agency), P. Chain (Los Alamos National Laboratory), M. Zafar Iqbal (Shahjalal University of Science and Technology) and M. Mutahar Hossain (Hossain & Khan Associates) for suggestions and comments on the project and manuscript. This research was funded by the Government of Bangladesh.
Author information
Authors and Affiliations
Contributions
M.A. conceived, designed and executed the project as principal investigator. M.M.A. and M.K.U. managed the project. M.S.I., M.S. Haque, Q.M.M.H., M.Z.H., R.H., N.A., U.H., R.I., and H.K. performed material preparation and multiplication, DNA and RNA extractions. S.H. and X.W. performed genome and transcriptome sequencing. M.S.I., E.M.E., M.M.I., M.S.R., M.M.R., M.S.A.K., S.M.T.K., M.M.A. and J.A.S. contributed to genome analysis including assembly, annotation, genome evolution and comparative genomics. M.S.I., R.A., B.A. and M.S. Hossain conducted fibre cell isolation for RNA extraction and conducted RT–qPCR of fibre biosynthesis genes. M.S.I., E.M.E., R.A., B.A., M.S. Hossain, S.M.T.K. and X.W. contributed to expression analysis of the fibre cell transcriptome. M.S.I., E.M.E., Q.M.M.H. and J.A.S. analysed the optical mapping data. B.A., A.H., M.M.K. and A.Y. conducted pathway and network analysis. M.S.I., M.S. Haque, M.M.I., A.H., M.Z.H., Q.M.M.H., J.A.S., T.H., M.S.A., M.M.M., S.M.E., A.M.M.H., N.M., M.S., S.S., N.S.S., S.J., S.R., A.C., A.I.A., G.M.N., K.S.U., T.R., S.M.E.H., A.R.S., S.M., S.A.M., M.K.I., M.Z.H.L., M.Z. and H.K. contributed to de novo genome assembly of Corchorus olitorius. M.S.I., J.A.S. and M.A. wrote and/or revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Note, Supplementary Figures 1–15. (PDF 2483 kb)
Supplementary Tables
Supplementary Tables 1–42. (XLSX 5106 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Islam, M., Saito, J., Emdad, E. et al. Comparative genomics of two jute species and insight into fibre biogenesis. Nature Plants 3, 16223 (2017). https://doi.org/10.1038/nplants.2016.223
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2016.223
This article is cited by
-
Impact of NaOH Treatment on the Chemical, Structural, Physico-mechanical, and Thermal Characteristics of Jute Species
Fibers and Polymers (2024)
-
Identification and Expression Profiles of Auxin-Related Gene Families Involved in Fiber Development in Jute (Corchorus capsularis)
Tropical Plant Biology (2024)
-
Genome-wide investigation of SnRK2 gene family in two jute species: Corchorus olitorius and Corchorus capsularis
Journal of Genetic Engineering and Biotechnology (2023)
-
Comparative metagenomics and microbial dynamics of jute retting environment
International Microbiology (2023)
-
Genome-wide in silico identification of phospholipase D (PLD) gene family from Corchorus capsularis and Corchorus olitorius: reveals their responses to plant stress
Journal of Genetic Engineering and Biotechnology (2022)