Abstract
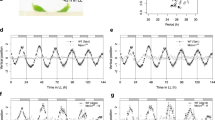
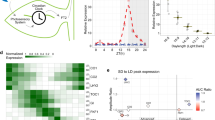
The circadian clock increases organisms' fitness by regulating physiological responses1. In mammals, the circadian clock in the suprachiasmatic nucleus (SCN) governs daily behavioural rhythms2. Similarly, in Arabidopsis, tissue-specific circadian clock functions have emerged, and the importance of the vasculature clock for photoperiodic flowering has been demonstrated3–5. However, it remains unclear if the vasculature clock regulates the majority of physiological responses, like the SCN in mammals, and if other environmental signals are also processed by the vasculature clock. Here, we studied the involvement of tissue-specific circadian clock regulation of flowering and cell elongation under different photoperiods and temperatures. We found that the circadian clock in vascular phloem companion cells is essential for photoperiodic flowering regulation; by contrast, the epidermis has a crucial impact on ambient temperature-dependent cell elongation. Thus, there are clear assignments of roles among circadian clocks in each tissue. Our results reveal that, unlike the more centralized circadian clock in mammals, the plant circadian clock is decentralized, where each tissue specifically processes individual environmental cues and regulates individual physiological responses. Our new conceptual framework will be a starting point for deciphering circadian clock functions in each tissue, which will lead to a better understanding of how circadian clock processing of environmental signals may be affected by ongoing climate change6.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Doherty, C. J. & Kay, S. A. Circadian control of global gene expression patterns. Annu. Rev. Genet. 44, 419–444 (2010).
Barclay, J. L., Tsang, A. H. & Oster, H. Interaction of central and peripheral clocks in physiological regulation. Prog. Brain Res. 199, 163–181 (2012).
James, A. B. et al. The circadian clock in Arabidopsis roots is a simplified slave version of the clock in shoots. Science 322, 1832–1835 (2008).
Yakir, E. et al. Cell autonomous and cell-type specific circadian rhythms in Arabidopsis. Plant J. 68, 520–531 (2011).
Endo, M. et al. Tissue-specific clocks in Arabidopsis show asymmetric coupling. Nature 515, 419–422 (2014).
Wolkovich, E. M. et al. Warming experiments underpredict plant phenological responses to climate change. Nature 485, 494–497 (2012).
Tahara, Y. & Shibata, S. Chronobiology and nutrition. Neuroscience 253, 78–88 (2013).
Eckel-Mahan, K. & Sassone-Corsi, P. Metabolism and the circadian clock converge. Physiol. Rev. 93, 107–135 (2013).
Baran, P. On Distributed Communications: I. Introduction to Distributed Communications Networks, RM-3420-PR (1964); http://www.rand.org/content/dam/rand/pubs/research_memoranda/2006/RM3420.pdf
Wigge, P. A. Ambient temperature signalling in plants. Curr. Opin. Plant Biol. 16, 661–666 (2013).
Wang, Z. Y. & Tobin, E. M. Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 93, 1207–1217 (1998).
Makino, S., Matsushika, A., Kojima, M., Yamashino, T. & Mizuno, T. The APRR1/TOC1 quintet implicated in circadian rhythms of Arabidopsis thaliana: I. Characterization with APRR1-overexpressing plants. Plant Cell Physiol. 43, 58–69 (2002).
Fukuda, H. Signals that control plant vascular cell differentiation. Nature Rev. Mol. Cell Biol. 5, 379–391 (2004).
Endo, M., Mochizuki, N., Suzuki, T. & Nagatani, A. CRYPTOCHROME2 in vascular bundles regulates flowering in Arabidopsis. Plant Cell 19, 84–93 (2007).
Takada, S. & Goto, K. TERMINAL FLOWER2, an Arabidopsis homolog of HETEROCHROMATIN PROTEIN1, counteracts the activation of FLOWERING LOCUS T by CONSTANS in the vascular tissues of leaves to regulate flowering time. Plant Cell 15, 2856–2865 (2003).
Abe, M. et al. FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309, 1052–1056 (2005).
Wigge, P. A. et al. Integration of spatial and temporal information during floral induction in Arabidopsis. Science 309, 1056–1059 (2005).
Huang, W. et al. Mapping the core of the Arabidopsis circadian clock defines the network structure of the oscillator. Science 336, 75–79 (2012).
Gendron, J. M. et al. Arabidopsis circadian clock protein, TOC1, is a DNA-binding transcription factor. Proc. Natl Acad. Sci. USA 109, 3167–3172 (2012).
Imaizumi, T. Arabidopsis circadian clock and photoperiodism: time to think about location. Curr. Opin. Plant Biol. 13, 83–89 (2010).
Kumar, S. V. Transcription factor PIF4 controls the thermosensory activation of flowering. Nature 484, 242–245 (2012).
King, R. W., Hisamatsu, T., Goldschmidt, E. E. & Blundell, C. The nature of floral signals in Arabidopsis. I. Photosynthesis and a far-red photoresponse independently regulate flowering by increasing expression of FLOWERING LOCUS T (FT). J. Exp. Bot. 59, 3811–3820 (2008).
Kutschera, U. & Niklas, K. J. The epidermal-growth-control theory of stem elongation: an old and a new perspective. J. Plant Physiol. 164, 1395–1409 (2007).
Nusinow, D. A. et al. The ELF4-ELF3-LUX complex links the circadian clock to diurnal control of hypocotyl growth. Nature 475, 398–402 (2011).
Hornitschek, P. et al. Phytochrome interacting factors 4 and 5 control seedling growth in changing light conditions by directly controlling auxin signaling. Plant J. 71, 699–711 (2012).
Stokes, M. E., Chattopadhyay, A., Wilkins, O., Nambara, E. & Campbell, M. M. Interplay between sucrose and folate modulates auxin signaling in Arabidopsis. Plant Physiol. 162, 1552–1565 (2013).
Turchi, L. et al. Arabidopsis HD-Zip II transcription factors control apical embryo development and meristem function. Development 140, 2118–2129 (2013).
Nomoto, Y., Kubozono, S., Yamashino, T., Nakamichi, N. & Mizuno, T. Circadian clock- and PIF4-controlled plant growth: a coincidence mechanism directly integrates a hormone signaling network into the photoperiodic control of plant architectures in Arabidopsis thaliana. Plant Cell Physiol. 53, 1950–1964 (2012).
Blázquez, M. A., Ahn, J. H. & Weigel, D. A thermosensory pathway controlling flowering time in Arabidopsis thaliana. Nature Genet. 33, 168–171 (2003).
Michael, T. P., Salome, P. A. & McClung, C. R. Two Arabidopsis circadian oscillators can be distinguished by differential temperature sensitivity. Proc. Natl Acad. Sci. USA 100, 6878–6883 (2003).
Shimada, T. L., Shimada, T. & Hara-Nishimura, I. A rapid and non-destructive screenable marker, FAST, for identifying transformed seeds of Arabidopsis thaliana. Plant J. 61, 519–528 (2010).
Porra, R. J., Thompson, W. A. & Kriedeman, P. E. Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim. Biophys. Acta 975, 384–394 (1989).
Acknowledgements
We thank M. Niwa, K. Ifuku, and Y.C. Brenda for technical assistance; Y. Kondo, S.L. Harmer and T. Imaizumi for helpful advice; J.A. Hejna for English proofreading. This work was supported by a JST PRESTO 14529738 (to M.E.), JSPS KAKENHI grants 25650097 (to M.E.), a Nakatani Foundation (to M.E.), a Mitsubishi Foundation (to M.E.), Grant-in-Aid for Scientific Research on Innovative Areas 87006029 (to M.E.), 26113510 (to M.E.) and 25113005 (to T.A.).
Author information
Authors and Affiliations
Contributions
M.E. planned the experiments. H.S., T.K., K.T. and K.K. performed experiments. H.S. and M.E. wrote the manuscript. M.E. and T.A. supervised the project. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Shimizu, H., Katayama, K., Koto, T. et al. Decentralized circadian clocks process thermal and photoperiodic cues in specific tissues. Nature Plants 1, 15163 (2015). https://doi.org/10.1038/nplants.2015.163
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2015.163
This article is cited by
-
Diurnal oscillations of epigenetic modifications are associated with variation in rhythmic expression of homoeologous genes in Brassica napus
BMC Biology (2023)
-
Rhythms of Transcription in Field-Grown Sugarcane Are Highly Organ Specific
Scientific Reports (2020)
-
ELF in the root
Nature Plants (2020)
-
Coronatine is more potent than jasmonates in regulating Arabidopsis circadian clock
Scientific Reports (2020)
-
The epidermis coordinates thermoresponsive growth through the phyB-PIF4-auxin pathway
Nature Communications (2020)