Abstract
Background/Objectives:
The SNF1/AMPK protein kinase has a central role in energy homeostasis in eukaryotic cells. It is activated by energy depletion and stimulates processes leading to the production of ATP while it downregulates ATP-consuming processes. The yeast SNF1 complex is best known for its role in glucose derepression.
Methods:
We performed a network reconstruction of the Snf1 pathway based on a comprehensive literature review. The network was formalised in the rxncon language, and we used the rxncon toolbox for model validation and gap filling.
Results:
We present a machine-readable network definition that summarises the mechanistic knowledge of the Snf1 pathway. Furthermore, we used the known input/output relationships in the network to identify and fill gaps in the information transfer through the pathway, to produce a functional network model. Finally, we convert the functional network model into a rule-based model as a proof-of-principle.
Conclusions:
The workflow presented here enables large scale reconstruction, validation and gap filling of signal transduction networks. It is analogous to but distinct from that established for metabolic networks. We demonstrate the workflow capabilities, and the direct link between the reconstruction and dynamic modelling, with the Snf1 network. This network is a distillation of the knowledge from all previous publications on the Snf1/AMPK pathway. The network is a knowledge resource for modellers and experimentalists alike, and a template for similar efforts in higher eukaryotes. Finally, we envisage the workflow as an instrumental tool for reconstruction of large signalling networks across Eukaryota.
Similar content being viewed by others
Introduction
The adenosine monophosphate-activated protein kinase (AMPK) is the key regulator of energy homeostasis in eukaryotic cells. It orchestrates cellular adaptation to nutrient availability and ensures cell survival under stressful conditions.1 Its central role in energy regulation makes it a potential drug target for treating diseases like obesity, heart diseases or type 2 diabetes. Thus, the mechanisms of activation of and signalling through AMPK have received great attention.
The Sucrose Non-Fermenting kinase Snf1 is the AMPK orthologue in baker’s yeast, Saccharomyces cerevisiae.2 Yeast cells adjust their metabolism in response to nutrient availability to ensure cell survival.3 Glucose and fructose cause the repression of numerous genes encoding nutrient uptake systems and metabolic enzymes, and cells shifted to less preferred carbon sources change their transcriptome and proteome to utilise new carbon sources.4 This reprogramming is mediated by the Snf1 kinase, which is a key regulator of energy metabolism also in yeast.5
The architecture of the Snf1 pathway is well-known. At the core of the Snf1 pathway is the heterotrimeric SNF1 complex, consisting of a catalytic α-subunit (Snf1), a regulatory γ-subunit (Snf4) and one of three β-subunits (Sip1, Sip2 and Gal83) that serve as scaffold and targeting subunits. These three forms of the SNF1 complex have overlapping but distinct roles and localisation patterns, and are regulated by upstream kinases, phosphatases and other modifiers. SNF1 in turn regulates a range of cytoplasmic and nuclear targets, in particular the transcription factors responsible for the reprogramming of energy metabolism. Hence, also the physiological role of the pathway is well-defined.
These features make the Snf1 system a suitable target for network reconstruction. This process is well-established for metabolic networks, where a high degree of conservation enables sequence-based reconstruction.6 However, the specificities of signalling components are encoded in short or ill-defined sequences, and may be completely disrupted by point mutations.7 Therefore, reconstruction of signalling networks relies on experimentally validated reactions as extracted from literature, and the validation on physiological input/output relationships. Various approaches have been used to generate large, primarily graphical, signalling network reconstructions.8–11 However, most of these cannot be simulated and validated, as routinely done for metabolic networks.12 Thus, we need an integrated workflow for reconstruction, validation and refinement of signalling networks.
Here, we present such a workflow and apply it to create a reconstruction of the Snf1 network. We performed an exhaustive literature review with the explicit aim to collect all mechanistic information on the signal transfer, and to compile that information in a stringent, machine-readable format. We chose the rxncon language for its scalability and fidelity to the empirical data structure,11,13 and used the rxncon toolbox for model creation, validation and gap filling.14 We extended the literature-curated network to enable information transfer between each input and every output that it triggers. The resulting network fully reproduces the expected qualitative behaviour in Boolean simulations, and each of the gap filling additions constitutes a clearly phrased hypothesis awaiting experimental validation. Finally, we generated a rule-based model corresponding to the final reconstruction as a proof-of-principle. We provide both the initial literature curation and the gap-filled network model as community resources that summarise the complete Snf1 literature to date. We envisage them as useful tools in yeast Snf1/AMPK research and as templates for similar efforts in higher organisms, and the workflow as a key tool for large scale reconstruction of signalling networks.
Materials and Methods
As described in detail in the Supplementary Material section (supplementary information 1), the reconstruction was performed using the rxncon language and tool.11 During the reconstruction process, we collected two kinds of data from literature: mechanistic and physiological/functional data. The mechanistic data were further divided into elemental reactions and contingencies. The elemental reactions define possible state transition events that produce or consume elemental states. Importantly, the elemental states define only a single property of a component, such as a specific modification or binding. Hence, they correspond to the full set of specific states for which that modification and binding is true, and, correspondingly, an elemental reaction corresponds to a set of reactions (reviewed in ref. 13). These decontextualised reactions are equivalent to the protein–protein interactions in e.g., the BioGRID database.15 The contingency information defines how elemental reactions depend on elemental states, and hence defines the causality in the network. The distinction between reactions and contingencies is the same as in the SBGN entity relationship diagrams,16 and together the reactions and contingencies fully define the network and can be used for automatic model generation (Supplementary file 1; ref. 14).
The physiological/functional data were used for validation of the network reconstruction. We searched for inputs known to activate Snf1 and for the downstream Snf1-dependent responses to these inputs, which we collected as a set of input/output relationships. For validation, we generated and simulated the corresponding bipartite Boolean model (bBM) with the rxncon toolbox, and visualised the attractor states on the regulatory graph in Cytoscape.14,17 We analysed only the attractor states, which are the end results of the simulations, due to the very crude time concept in Boolean models. The attractor states correspond to a qualitative steady state, which can be used to determine if the signal is transduced through the network or not. We scored functionality for each input–output relationship by determining if that output responded appropriately when the input was changed between on and off (Table 1). When necessary, we adapted the network definition to resolve blocks and/or constitutive activities as detailed in the Results section. All such adaptations have been clearly labelled as hypotheses in the updated network definition (Supplementary file 2). Finally, we translated the gap-filled network into a rule-based model in the BioNetGen language.18 All methods are described in more detail in the Supplementary Methods section.
Results
The Snf1 network reconstruction is based on a comprehensive literature review
We present a network reconstruction of the Snf1 pathway based on a comprehensive literature review. We used the Textpresso tool at Saccharomyces Genome Database (SGD) to search the literature with the ‘Snf1’ string, and extended this list with further papers manually. In total, we could find and download 444 publications dating from February 1977 to January 2015 from online research literature repositories (Supplementary file 3). We read and evaluated each of these papers, searching for and re-evaluating experimental evidence of interactions of pathway components. We extracted mechanistic information on the Snf1 pathway from 77 papers.19–95 The literature-derived network reconstruction (NR1) encompasses 71 reactions and 105 contingencies, each of which is associated with ⩾1 references (Supplementary file 1). Hence, this initial network reconstruction is fully referenced and based on careful manual curation of the entire Snf1 literature.
The network reconstruction encompasses 52 components taking part in 71 elemental reactions
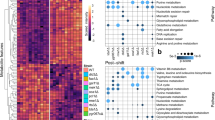
At the topological level, NR1 encompasses 52 components and 71 elemental reactions (Figure 1). The components are proteins (44), small molecules (1) and transcription factor-binding sites (7). The signalling pathway displays a clear bow-tie structure centred on the SNF1 kinase complex, which participates in 40 of the 71 reactions. The activity of the SNF1 kinase complex is controlled by posttranslational modification by a battery of upstream regulators. The key modification is phosphorylation of Thr210 in Snf1, which is indispensable for kinase activity. This residue is phosphorylated by any of the three upstream kinases Sak1, Elm1 or Tos3,75 and dephosphorylated by phosphoprotein phosphatase type 1 (PP1); consisting of the catalytic subunit Glc7 and either of the regulatory subunits Reg1 and Reg2.75 In addition, the SNF1 complex is regulated by ubiquitylation and sumoylation of the catalytic Snf1 subunit. The active SNF1 complex in turn regulates a wide range of targets that are primarily involved in energy metabolism and transcription.
The Snf1 pathway and its components. The reaction graph depicts the pathway components of the NR1 network, and the different reactions they take part in (excluding the nuclear import and export reactions, and the nuclear pore complex mediating them). The SNF1 complex is regulated by kinases, phosphatases and other modifiers shown at the top. The transcriptional regulation is shown at the bottom. The coloured circles represent components and coloured edges represent different classes of reactions. The information in the reaction graph is only topological, and the edges cannot be interpreted in terms of information flow. However, most of them do carry information, as shown in the regulatory graph (Figure 2).
The signal transmission in the Snf1 pathway is well-understood
At the regulatory level, NR1 encompasses 71 elemental reactions that produce or consume 64 elemental states, which in turn regulate the reactions via 108 contingencies (Figure 2; ref. 11). The network retains the bow-tie structure also at the regulatory level, where it is centred on the active forms of the SNF1 complex. The graph is well-connected, meaning that the causal relationships between reactions and states are well-known. Importantly, there are directed paths from the inputs to the outputs, indicating that the mechanism of information transfer is understood at the molecular, mechanistic level. However, there is one disconnected subgraph, reflecting our lack of knowledge on how—at a mechanistic level—glucose and energy regulate the Snf1 pathway.
The regulatory structure of the Snf1 pathway. The regulatory graph depicts the information flow through the NR1 network as a bipartite-directed graph. Elemental reactions (red nodes) produce or consume elemental states (blue nodes) via reaction edges, which correspond to the edges in the reaction graph (Figure 1). The elemental states in turn influence the elemental reactions via contingency edges, and they define contextual constraints on reactions. The inputs in form of external stimuli can be found at the top of the picture, as grey nodes with node names in square brackets. The cytoplasmic reactions and SNF1 complex formation are found in the middle part, and the nuclear reactions at the bottom with the transcriptional output again as grey nodes with names in square brackets. Information passes through the network along the unidirectional edges, either by production (blue edges) or consumption (purple edges) of elemental states by reactions, or by the regulatory effect of elemental states on elemental reactions. Positive contingencies from activating states to activated reactions are denoted in green, while inhibitory contingencies appear in red. More complex requirements, such as formation of the active forms of the SNF1 complex, are defined by Boolean states that are indicated as white triangles (AND) or diamonds (OR). The compartment borders in grey represent the plasma membrane and the nuclear membrane, and have been included as visual guides but carry no information. At the center, we have the three active forms of the SNF1 complex; SNF1*CYT, SNF1*VAC and SNF1*NUC, that are localised to the cytoplasm, the vacuole and the nucleus, respectively. Information can only pass along the direction of the edges, and we can follow them from inputs to outputs. In this network we have four inputs: The three grey stress inputs to the upper right, and extracellular glucose to the upper left. The last is part of an unconnected subgraph, highlighting the fact that we do not know how the glucose signal is sensed by the pathway. The original NR1 Cytoscape file is attached as Supplementary file 5.
Input/output validation reveals specific knowledge gaps
To validate NR1, we made use of the well-understood input/output relationship of the Snf1 pathway. We examined whether the network reconstruction sufficed to enable information transfer through the network as expected. For this purpose, we used a bBM to determine if the attractor states for every input combination correspond to that expected based on our empirical understanding of the pathway. Examining the input/output relationship of the Snf1 network reconstruction, we found that none of the documented input/output relationships could be reproduced by the bBM of NR1 (Table 1), highlighting important mechanistic gaps in the combined literature knowledge.
The gap filling process required three steps to generate a functional network model
We used iterative network improvement, bBM generation and validation as described above to identify and fill the gaps in the network. We identified blocks in the information transfer, i.e., reactions or states that do not vary as expected in response to varying inputs, and eliminated these blocks by minimal modifications of the network (Figure 3). The gap filling workflow on the Snf1 network reconstruction identified a total of 1 missing component, 1 missing reaction and 13 missing contingencies (Table 2), and resulted in a final network (NR2) with 53 components, 72 reactions and 118 contingencies (Supplementary file 2). The corresponding bBM reproduced the expected input/output relationships in all tested cases, as shown for glucose deprivation (Figure 4), basic conditions (Supplementary file 9), salt stress (Supplementary file 10), alkaline conditions (Supplementary file 11) and nitrogen starvation (Supplementary file 12). NR2 is a merge of the explicitly referenced original curation and clearly labelled hypothetical reactions and contingencies, each of which correspond to a testable hypothesis.
Three gap filling steps suffice to fully connect inputs with their respective outputs. The figure displays the difference between the initial (NR1) and final (NR2) networks. (a) Connection of the glucose subgraph to the main pathway. This connection required one abstract state, which is active in the presence of glucose as long as none of the stresses are active, and which activates sumoylation and inhibits phosphorylation of Snf1. It also activates the phosphorylation of Msn2. (b) Adaptation of transport reactions. The nuclear localisation pattern could only be reproduced if both import and export are regulated, otherwise the localisation oscillates. In addition, we added a direct edge from salt stress to inhibit Snf1 nuclear localisation, to account for the observation that Snf1 is phosphorylated but not nuclear on salt stress. (c) Deubiquitylation of Rod1. An unknown deubiqitylating enzyme was added, acting on Rod1. Faded nodes and edges were part of the initial network reconstruction (NR1), while nodes and edges with full colours were added to NR2 during the network refinement. The gap filled NR2 Cytoscape file is attached as Supplementary file 6.
The steady state of the Snf1 bBM under glucose deprivation is a point attractor. The end state of the bBM simulation under glucose-deprived conditions visualised on the regulatory graph of the updated network. The bBM was generated automatically based on the final network after validation, and the simulation initiated from the starting states defined in the Methods section. The pale nodes are inactive and the filled nodes are active. The Boolean model is attached in the BooleanNet and BoolNet formats as Supplementary files 7 and 8, respectively.
The rxncon network definition corresponds to a dynamic model
As proof-of-principle, we used the rxncon toolbox and NR2 to generate a rule-based model in the BioNetGen language (Supplementary file 4). The resulting model has 176 distinct parameters, precluding reliable parameter estimation based on current data and hence meaningful analysis. However, the model can be simulated using NFsim,96 demonstrating that the rxncon network reconstruction can be used as basis for dynamic simulations.
Discussion
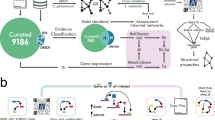
Here, we present a network reconstruction of the Snf1 signalling pathway. Network reconstruction is well-established for metabolic networks, and they are divided into four stages: (i) draft reconstruction, (ii) refinement, (iii) conversion into a computational model and (iv) network evaluation (including gap filling).12 However, these methods cannot directly be applied to signal transduction networks, and hence we developed an analogous, but distinct, workflow (Figure 5).
The network reconstruction workflow. The first phase of the process was the network reconstruction itself, where we extracted reaction and contingency information from published data. The curation process produced the initial network reconstruction (NR1). The second phase of the process was the validation and iterative improvement. It starts with the creation of the bBM, followed by simulation of the bBM and comparison to the known input/output relationships. If the model fails to transmit information as expected, the gap is identified and filled, and the updated network re-enters the workflow at model creation. If the model passes all tests, it is accepted and considered a validated network (NR2). The validated network constitutes a knowledge resource, but can also be further processed with the rxncon tool to generate graphical or mathematical models in a range of formats, as illustrated by the rule-based model (Supplementary file 4).
The first and arguably most important phase is the translation of diverse experimental findings into a single, machine-readable reconstruction of the system under study. For this purpose, we carefully curated and distilled the complete literature on the Snf1/AMPK pathway architecture into a network reconstruction. We searched for evidence of reactions between network components and of causal relationships between reactions (i.e. contingencies), and formalised the reactions and contingencies in the rxncon language. The resulting network is fully annotated, machine readable and can be used for automatic model generation with the rxncon software tool. Importantly, we found text mining insufficient for high quality network reconstruction and based the curation on manual re-evaluation of the data presented in the cited papers. Hence, this first phase corresponds to the first and second stages of metabolic network reconstruction.12 The result is a high-quality curation summarising the entire Snf1 literature in a fully annotated, machine-readable format.
The second phase is the network validation. This is again well-established for metabolic networks, based on the assumption that mass transfer paths exist such that all metabolites can be reached from the input nutrients.97 Again, this does not hold here, as there is no mass transfer through a signalling network. Instead, we used the known input/output relationships to define information paths through the network. The first step of phase II is the generation of a computational model, corresponding to stage 3 of metabolic network reconstruction.12 Here, we used the rxncon tool to generate and simulate the bBM corresponding to the network reconstruction, and used the bBM to determine whether information paths are functional.14 This analysis revealed that none of the input/output paths could be reproduced based on the literature curation itself (Table 1). Thus, critical information was missing from our understanding of the Snf1 pathway.
To identify these missing links, we went through an iterative gap filling and validation process. The bBM was modelled with alternating inputs to determine which paths worked as expected. For those that did not, we identified the missing steps manually and modified the network to solve the problem. After modification, a new bBM was generated and evaluated, and the process repeated until all paths were functional (Table 1). The modifications fall into one of three classes: missing causal links (i.e. contingencies), states that could not be consumed (missing reverse reactions) or artefacts from the binary assumptions in the Boolean model (missing/modified contingencies). Each modification corresponds to a testable hypothesis, and is clearly labelled as such to distinguish it from the reactions and contingencies that are based on existing literature evidence (Supplementary file 2). The result is a validated, functional network that qualitatively reproduces the input/output paths known form empirical observations.
The gap filling modifications fall into three groups (Figure 3; Table 2). First, the reconstruction lacks a connection between intracellular glucose and phosphorylation of Snf1, consistent with the current (lack of) understanding of how the pathway is activated. To enable the response to glucose, we included an unknown regulatory step connecting glucose and stress signals to Snf1, as proposed in ref. 98. It stimulates Snf1 sumoylation and inhibits phosphorylation such that Snf1 is activated on glucose deprivation. Second, we adapted the trans-organelle transport reactions. Both import and export need to be regulated to prevent components from cycling between compartments. While this is an artefact from the binary assumptions in the bBM, it is also likely that the nucleo-cytoplasmic shuttling is regulated in both directions to avoid futile cycling in vivo. Third, we added consumption reactions for states lacking them. Here, Rod1 is ubiquitylated by Rsp5 but the reverse reaction is unknown. To avoid permanent ubiquitylation of Rod1, we introduced an unknown deubiquitylating peptidase. Twenty such enzymes exist in yeast, although activity against Rod1 has not been reported.99 Hence, the gaps identified in the validation process could be filled with reactions and contingencies that are consistent with our knowledge, and which constitute explicit testable hypotheses.
The mechanism of activation remains an open question. Snf1 responds to a number of stress factors in addition to glucose limitation, namely high salt concentrations, alkaline pH and nitrogen limitation.44,100 The proposal that energy is the main cue is consistent with the wide but specific range of stresses that induces the pathway, and the unknown regulatory step we included may turn out to be a direct measure of the energy state of the cell. The strongest candidate is nucleotide binding, consistent with how AMPK works in higher organisms. There is some evidence in this direction, as ADP protects Snf1 from dephosphorylation101 and AMP appears to interact with Snf4.102 However, there are also reports that the SAGA acetyl transferase complex deubiquitylates Snf1, thereby affecting Snf1 kinase activity, Snf1 phosphorylation and SNF1 complex stability.103 Ubiquitylation is stimulated by SUMOylation, and Snf1 is SUMOylated by the SUMO-E3 ligase Mms21 under glucose conditions.93 As a consequence, Snf1 is ubiquitylated by Slx5-Slx8 and degraded.104 Thus, the mechanism remains elusive, and we summarise this gap in a single, glucose-regulated step in the final network.
To create this network reconstruction, we introduced a workflow for network reconstruction, validation and iterative refinement. Similar workflows have proven crucial for the success in large scale metabolic network reconstruction and modelling, but have hitherto not been available for signalling networks. Here, we demonstrate that this workflow works for a well-defined pathway. However, the challenges will aggravate as we extend the scope toward larger and eventually genome scale networks. First, the knowledge gaps will be much more severe in the grey areas between the traditional pathways as compared with their well-studied cores. This information bias is likely one of the reasons for the clear bow-tie structures in the SNF network (Figures 1 and 2). Second, the demands on the formats and methods increase drastically with larger network size. Several large mapping efforts have used the specific state based process description format, and these maps are highly valued community resources.8–10 Here, we chose a reaction-contingency based format, as the network definition scales more favourably with network size and has better congruence with experimental data (reviewed in ref. 13). This choice also enabled the use of the rxncon toolbox with the automatic model generation that was necessary for the iterative validation and gap filling processes. As we illustrate with the rule-based model generation, the rxncon network definition can also be used to create a dynamic model. However, meaningful parameterisation and efficient simulation of such large models are still outstanding challenges. Finally, the rxncon language enables clean bottom-up reconstructions, as each reaction and contingency can be defined independently of other reactions and contingencies, respectively. This stands in stark contrast to the specific state-based formats, where reactions and causalities are weaved together and must be adapted as new information appears. Taken together, the proposed workflow provides an approach to tackle large networks with partially very sparse knowledge. We are convinced that this or a similar approach will be instrumental in the reconstructions of genome scale signalling networks in eukaryotes.
References
Hardie DG, Ross FA, Hawley SA . AMP-activated protein kinase: a target for drugs both ancient and modern. Chem Biol 2012; 19: 1222–1236.
Carlson M, Osmond BC, Botstein D . Mutants of yeast defective in sucrose utilization. Genetics 1981; 98: 25–40.
Conrad M, Schothorst J, Kankipati HN, Van Zeebroeck G, Rubio-Texeira M, Thevelein JM . Nutrient sensing and signalling in the yeast Saccharomyces cerevisiae . FEMS Microbiol Rev 2014; 38: 254–299.
Zaman S, Lippman SI, Zhao X, Broach JR . How Saccharomyces responds to nutrients. Annu Rev Genet 2008; 42: 27–81.
Wilson WA, Hawley SA, Hardie DG . Glucose repression/derepression in budding yeast: SNF1 protein kinase is activated by phosphorylation under derepressing conditions, and this correlates with a high AMP:ATP ratio. Curr Biol 1996; 6: 1426–1434.
Herrgard MJ, Swainston N, Dobson P, Dunn WB, Arga KY, Arvas M et al. A consensus yeast metabolic network reconstruction obtained from a community approach to systems biology. Nat Biotechnol 2008; 26: 1155–1160.
Zarrinpar A, Park SH, Lim WA . Optimization of specificity in a cellular protein interaction network by negative selection. Nature 2003; 426: 676–680.
Oda K, Kitano H . A comprehensive map of the toll-like receptor signalling network. Mol Syst Biol 2006; 2: 2006 0015.
Kaizu K, Ghosh S, Matsuoka Y, Moriya H, Shimizu-Yoshida Y, Kitano H . A comprehensive molecular interaction map of the budding yeast cell cycle. Mol Syst Biol 2010; 6: 415.
Croft D, O’Kelly G, Wu G, Haw R, Gillespie M, Matthews L et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res 2011; 39 (Database issue): D691–D697.
Tiger CF, Krause F, Cedersund G, Palmer R, Klipp E, Hohmann S et al. A framework for mapping, visualisation and automatic model creation of signal-transduction networks. Mol Syst Biol 2012; 8: 578.
Thiele I, Palsson BO . A protocol for generating a high-quality genome-scale metabolic reconstruction. Nat Protoc 2010; 5: 93–121.
Rother M, Munzner U, Thieme S, Krantz M . Information content and scalability in signal transduction network reconstruction formats. Mol biosyst 2013; 9: 1993–2004.
Flottmann M, Krause F, Klipp E, Krantz M . Reaction-contingency based bipartite Boolean modelling. BMC Syst Biol 2013; 7: 58.
Chatr-Aryamontri A, Breitkreutz BJ, Oughtred R, Boucher L, Heinicke S, Chen D et al. The BioGRID interaction database: 2015 update. Nucleic Acids Res 2015; 43 (Database issue): D470–D478.
Le Novere N, Hucka M, Mi H, Moodie S, Schreiber F, Sorokin A et al. The systems biology graphical notation. Nat Biotechnol 2009; 27: 735–741.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003; 13: 2498–2504.
Blinov ML, Faeder JR, Goldstein B, Hlavacek WS . BioNetGen: software for rule-based modelling of signal transduction based on the interactions of molecular domains. Bioinformatics (Oxford, England) 2004; 20: 3289–3291.
Ahuatzi D, Riera A, Pelaez R, Herrero P, Moreno F . Hxk2 regulates the phosphorylation state of Mig1 and therefore its nucleocytoplasmic distribution. J Biol Chem 2007; 282: 4485–4493.
Amodeo GA, Rudolph MJ, Tong L . Crystal structure of the heterotrimer core of Saccharomyces cerevisiae AMPK homologue SNF1. Nature 2007; 449: 492–495.
Becuwe M, Vieira N, Lara D, Gomes-Rezende J, Soares-Cunha C, Casal M et al. A molecular switch on an arrestin-like protein relays glucose signalling to transporter endocytosis. J Cell Biol 2012; 196: 247–259.
Benanti JA, Cheung SK, Brady MC, Toczyski DP . A proteomic screen reveals SCFGrr1 targets that regulate the glycolytic-gluconeogenic switch. Nat Cell Biol 2007; 9: 1184–1191.
Bertram PG, Choi JH, Carvalho J, Chan TF, Ai W, Zheng XF . Convergence of TOR-nitrogen and Snf1-glucose signalling pathways onto Gln3. Mol Cell Biol 2002; 22: 1246–1252.
Castermans D, Somers I, Kriel J, Louwet W, Wera S, Versele M et al. Glucose-induced posttranslational activation of protein phosphatases PP2A and PP1 in yeast. Cell Res 2012; 22: 1058–1077.
Celenza JL, Carlson M . Mutational analysis of the Saccharomyces cerevisiae SNF1 protein kinase and evidence for functional interaction with the SNF4 protein. Mol Cell Biol 1989; 9: 5034–5044.
Celenza JL, Eng FJ, Carlson M . Molecular analysis of the SNF4 gene of Saccharomyces cerevisiae: evidence for physical association of the SNF4 protein with the SNF1 protein kinase. Mol Cell Biol 1989; 9: 5045–5054.
Charbon G, Breunig KD, Wattiez R, Vandenhaute J, Noel-Georis I . Key role of Ser562/661 in Snf1-dependent regulation of Cat8p in Saccharomyces cerevisiae and Kluyveromyces lactis. Mol Cell Biol 2004; 24: 4083–4091.
De Wever V, Reiter W, Ballarini A, Ammerer G, Brocard C . A dual role for PP1 in shaping the Msn2-dependent transcriptional response to glucose starvation. EMBO J 2005; 24: 4115–4123.
DeVit MJ, Johnston M . The nuclear exportin Msn5 is required for nuclear export of the Mig1 glucose repressor of Saccharomyces cerevisiae . Curr Biol 1999; 9: 1231–1241.
Dombek KM, Voronkova V, Raney A, Young ET . Functional analysis of the yeast Glc7-binding protein Reg1 identifies a protein phosphatase type 1-binding motif as essential for repression of ADH2 expression. Mol Cell Biol 1999; 19: 6029–6040.
Dubacq C, Chevalier A, Mann C . The protein kinase Snf1 is required for tolerance to the ribonucleotide reductase inhibitor hydroxyurea. Mol Cell Biol 2004; 24: 2560–2572.
Elbing K, McCartney RR, Schmidt MC . Purification and characterization of the three Snf1-activating kinases of Saccharomyces cerevisiae . Biochem J 2006; 393 (Pt 3): 797–805.
Elbing K, Rubenstein EM, McCartney RR, Schmidt MC . Subunits of the Snf1 kinase heterotrimer show interdependence for association and activity. J Biol Chem 2006; 281: 26170–26180.
Estruch F, Carlson M . Increased dosage of the MSN1 gene restores invertase expression in yeast mutants defective in the SNF1 protein kinase. Nucleic Acids Res 1990; 18: 6959–6964.
Fernandez-Garcia P, Pelaez R, Herrero P, Moreno F . Phosphorylation of yeast hexokinase 2 regulates its nucleocytoplasmic shuttling. J Biol Chem 2012; 287: 42151–42164.
Frederick DL, Tatchell K . The REG2 gene of Saccharomyces cerevisiae encodes a type 1 protein phosphatase-binding protein that functions with Reg1p and the Snf1 protein kinase to regulate growth. Mol Cell Biol 1996; 16: 2922–2931.
Garreau H, Hasan RN, Renault G, Estruch F, Boy-Marcotte E, Jacquet M . Hyperphosphorylation of Msn2p and Msn4p in response to heat shock and the diauxic shift is inhibited by cAMP in Saccharomyces cerevisiae . Microbiology 2000; 146 (Pt 9): 2113–2120.
Gimeno-Alcaniz JV, Sanz P . Glucose and type 2A protein phosphatase regulate the interaction between catalytic and regulatory subunits of AMP-activated protein kinase. J Mol Biol 2003; 333: 201–209.
Hahn JS, Thiele DJ . Activation of the Saccharomyces cerevisiae heat shock transcription factor under glucose starvation conditions by Snf1 protein kinase. J Biol Chem 2004; 279: 5169–5176.
Hardy TA, Huang D, Roach PJ . Interactions between cAMP-dependent and SNF1 protein kinases in the control of glycogen accumulation in Saccharomyces cerevisiae . J Biol Chem 1994; 269: 27907–27913.
Hedbacker K, Carlson M . Regulation of the nucleocytoplasmic distribution of Snf1-Gal83 protein kinase. Eukaryot Cell 2006; 5: 1950–1956.
Hedbacker K, Hong SP, Carlson M . Pak1 protein kinase regulates activation and nuclear localisation of Snf1-Gal83 protein kinase. Mol Cell Biol 2004; 24: 8255–8263.
Hedbacker K, Townley R, Carlson M . Cyclic AMP-dependent protein kinase regulates the subcellular localisation of Snf1-Sip1 protein kinase. Mol Cell Biol 2004; 24: 1836–1843.
Hong SP, Carlson M . Regulation of snf1 protein kinase in response to environmental stress. J Biol Chem 2007; 282: 16838–16845.
Hong SP, Leiper FC, Woods A, Carling D, Carlson M . Activation of yeast Snf1 and mammalian AMP-activated protein kinase by upstream kinases. Proc Natl Acad Sci USA 2003; 100: 8839–8843.
Jiang R, Carlson M . Glucose regulates protein interactions within the yeast SNF1 protein kinase complex. Genes Dev 1996; 10: 3105–3115.
Jiang R, Carlson M . The Snf1 protein kinase and its activating subunit, Snf4, interact with distinct domains of the Sip1/Sip2/Gal83 component in the kinase complex. Mol Cell Biol 1997; 17: 2099–2106.
Jin C, Barrientos A, Epstein CB, Butow RA, Tzagoloff A . SIT4 regulation of Mig1p-mediated catabolite repression in Saccharomyces cerevisiae . FEBS Lett 2007; 581: 5658–5663.
Kaniak A, Xue Z, Macool D, Kim JH, Johnston M . Regulatory network connecting two glucose signal transduction pathways in Saccharomyces cerevisiae . Eukaryot Cell 2004; 3: 221–231.
Kim JH, Brachet V, Moriya H, Johnston M . Integration of transcriptional and posttranslational regulation in a glucose signal transduction pathway in Saccharomyces cerevisiae . Eukaryot Cell 2006; 5: 167–173.
Kuchin S, Vyas VK, Carlson M . Snf1 protein kinase and the repressors Nrg1 and Nrg2 regulate FLO11, haploid invasive growth, and diploid pseudohyphal differentiation. Mol Cell Biol 2002; 22: 3994–4000.
Kuchin S, Vyas VK, Kanter E, Hong SP, Carlson M . Std1p (Msn3p) positively regulates the Snf1 kinase in Saccharomyces cerevisiae . Genetics 2003; 163: 507–514.
Kuchin SV, Kartasheva NN, Benevolensky SV . Genes required for derepression of an extracellular glucoamylase gene, STA2, in the yeast Saccharomyces. Yeast (Chichester, England) 1993; 9: 533–541.
Lesage P, Yang X, Carlson M . Analysis of the SIP3 protein identified in a two-hybrid screen for interaction with the SNF1 protein kinase. Nucleic Acids Res 1994; 22: 597–603.
Lesage P, Yang X, Carlson M . Yeast SNF1 protein kinase interacts with SIP4, a C6 zinc cluster transcriptional activator: a new role for SNF1 in the glucose response. Mol Cell Biol 1996; 16: 1921–1928.
Lin SS, Manchester JK, Gordon JI . Sip2 an N-myristoylated beta subunit of Snf1 kinase, regulates aging in Saccharomyces cerevisiae by affecting cellular histone kinase activity, recombination at rDNA loci, and silencing. J Biol Chem 2003; 278: 13390–13397.
Liu Y, Xu X, Carlson M . Interaction of SNF1 protein kinase with its activating kinase Sak1. Eukaryot Cell 2011; 10: 313–319.
Liu Y, Xu X, Singh-Rodriguez S, Zhao Y, Kuo MH . Histone H3 Ser10 phosphorylation-independent function of Snf1 and Reg1 proteins rescues a gcn5- mutant in HIS3 expression. Mol Cell Biol 2005; 25: 10566–10579.
Lo WS, Duggan L, Emre NC, Belotserkovskya R, Lane WS, Shiekhattar R et al. Snf1--a histone kinase that works in concert with the histone acetyltransferase Gcn5 to regulate transcription. Science (New York, NY) 2001; 293: 1142–1146.
Lo WS, Gamache ER, Henry KW, Yang D, Pillus L, Berger SL . Histone H3 phosphorylation can promote TBP recruitment through distinct promoter-specific mechanisms. EMBO J 2005; 24: 997–1008.
Ludin K, Jiang R, Carlson M . Glucose-regulated interaction of a regulatory subunit of protein phosphatase 1 with the Snf1 protein kinase in Saccharomyces cerevisiae . Proc Natl Acad Sci USA 1998; 95: 6245–6250.
Lutfiyya LL, Iyer VR, DeRisi J, DeVit MJ, Brown PO, Johnston M . Characterization of three related glucose repressors and genes they regulate in Saccharomyces cerevisiae . Genetics 1998; 150: 1377–1391.
Martinez-Pastor MT, Marchler G, Schuller C, Marchler-Bauer A, Ruis H, Estruch F . The Saccharomyces cerevisiae zinc finger proteins Msn2p and Msn4p are required for transcriptional induction through the stress response element (STRE). EMBO J 1996; 15: 2227–2235.
Momcilovic M, Carlson M . Alterations at dispersed sites cause phosphorylation and activation of SNF1 protein kinase during growth on high glucose. J Biol Chem 2011; 286: 23544–23551.
Momcilovic M, Iram SH, Liu Y, Carlson M . Roles of the glycogen-binding domain and Snf4 in glucose inhibition of SNF1 protein kinase. J Biol Chem 2008; 283: 19521–19529.
Nath N, McCartney RR, Schmidt MC . Yeast Pak1 kinase associates with and activates Snf1. Mol Cell Biol 2003; 23: 3909–3917.
Olivari MT, Homans DC, Wilson RF, Kubo SH, Ring WS . Coronary artery disease in cardiac transplant patients receiving triple-drug immunosuppressive therapy. Circulation 1989; 80 (5 Pt 2): Iii111–Iii115.
Ostling J, Ronne H . Negative control of the Mig1p repressor by Snf1p-dependent phosphorylation in the absence of glucose. Eur J Biochem 1998; 252: 162–168.
Pelaez R, Herrero P, Moreno F . Functional domains of yeast hexokinase 2. Biochem J. 2010; 432: 181–190.
Randez-Gil F, Bojunga N, Proft M, Entian KD . Glucose derepression of gluconeogenic enzymes in Saccharomyces cerevisiae correlates with phosphorylation of the gene activator Cat8p. Mol Cell Biol 1997; 17: 2502–2510.
Randez-Gil F, Herrero P, Sanz P, Prieto JA, Moreno F . Hexokinase PII has a double cytosolic-nuclear localisation in Saccharomyces cerevisiae . FEBS Lett 1998; 425: 475–478.
Reifenberger E, Boles E, Ciriacy M . Kinetic characterization of individual hexose transporters of Saccharomyces cerevisiae and their relation to the triggering mechanisms of glucose repression. Eur J Biochem 1997; 245: 324–333.
Rodriguez-Colman MJ, Sorolla MA, Vall-Llaura N, Tamarit J, Ros J, Cabiscol E . The FOX transcription factor Hcm1 regulates oxidative metabolism in response to early nutrient limitation in yeast. Role of Snf1 and Tor1/Sch9 kinases. Biochim Biophys Acta 2013; 1833: 2004–2015.
Roth S, Kumme J, Schuller HJ . Transcriptional activators Cat8 and Sip4 discriminate between sequence variants of the carbon source-responsive promoter element in the yeast Saccharomyces cerevisiae . Curr Genet 2004; 45: 121–128.
Rubenstein EM, McCartney RR, Zhang C, Shokat KM, Shirra MK, Arndt KM et al. Access denied: Snf1 activation loop phosphorylation is controlled by availability of the phosphorylated threonine 210 to the PP1 phosphatase. J Biol Chem 2008; 283: 222–230.
Sanz P, Alms GR, Haystead TA, Carlson M . Regulatory interactions between the Reg1-Glc7 protein phosphatase and the Snf1 protein kinase. Mol Cell Biol 2000; 20: 1321–1328.
Simpson-Lavy KJ, Johnston M . SUMOylation regulates the SNF1 protein kinase. Proc Natl Acad Sci USA 2013; 110: 17432–17437.
Smith FC, Davies SP, Wilson WA, Carling D, Hardie DG . The SNF1 kinase complex from Saccharomyces cerevisiae phosphorylates the transcriptional repressor protein Mig1p in vitro at four sites within or near regulatory domain 1. FEBS Lett 1999; 453: 219–223.
Soontorngun N, Larochelle M, Drouin S, Robert F, Turcotte B . Regulation of gluconeogenesis in Saccharomyces cerevisiae is mediated by activator and repressor functions of Rds2. Mol Cell Biol 2007; 27: 7895–7905.
Tabba S, Mangat S, McCartney R, Schmidt MC . PP1 phosphatase-binding motif in Reg1 protein of Saccharomyces cerevisiae is required for interaction with both the PP1 phosphatase Glc7 and the Snf1 protein kinase. Cell Signal 2010; 22: 1013–1021.
Tamai KT, Liu X, Silar P, Sosinowski T, Thiele DJ . Heat shock transcription factor activates yeast metallothionein gene expression in response to heat and glucose starvation via distinct signalling pathways. Mol Cell Biol 1994; 14: 8155–8165.
Tillman TS, Ganster RW, Jiang R, Carlson M, Schmidt MC . STD1 (MSN3) interacts directly with the TATA-binding protein and modulates transcription of the SUC2 gene of Saccharomyces cerevisiae . Nucleic Acids Res 1995; 23: 3174–3180.
Treitel MA, Carlson M . Repression by SSN6-TUP1 is directed by MIG1, a repressor/activator protein. Proc Natl Acad Sci USA 1995; 92: 3132–3136.
Treitel MA, Kuchin S, Carlson M . Snf1 protein kinase regulates phosphorylation of the Mig1 repressor in Saccharomyces cerevisiae . Mol Cell Biol 1998; 18: 6273–6280.
Tu J, Carlson M . REG1 binds to protein phosphatase type 1 and regulates glucose repression in Saccharomyces cerevisiae . EMBO J 1995; 14: 5939–5946.
Van Driessche B, Coddens S, Van Mullem V, Vandenhaute J . Glucose deprivation mediates interaction between CTDK-I and Snf1 in Saccharomyces cerevisiae . FEBS Lett 2005; 579: 5318–5324.
Vincent O, Carlson M . Gal83 mediates the interaction of the Snf1 kinase complex with the transcription activator Sip4. EMBO J 1999; 18: 6672–6681.
Vincent O, Kuchin S, Hong SP, Townley R, Vyas VK, Carlson M . Interaction of the Srb10 kinase with Sip4, a transcriptional activator of gluconeogenic genes in Saccharomyces cerevisiae . Mol Cell Biol 2001; 21: 5790–5796.
Vincent O, Townley R, Kuchin S, Carlson M . Subcellular localisation of the Snf1 kinase is regulated by specific beta subunits and a novel glucose signalling mechanism. Genes Dev 2001; 15: 1104–1114.
Vyas VK, Kuchin S, Berkey CD, Carlson M . Snf1 kinases with different beta-subunit isoforms play distinct roles in regulating haploid invasive growth. Mol Cell Biol 2003; 23: 1341–1348.
Vyas VK, Kuchin S, Carlson M . Interaction of the repressors Nrg1 and Nrg2 with the Snf1 protein kinase in Saccharomyces cerevisiae . Genetics 2001; 158: 563–572.
Wilson MA, Koutelou E, Hirsch C, Akdemir K, Schibler A, Barton MC et al. Ubp8 and SAGA regulate Snf1 AMP kinase activity. Mol Cell Biol 2011; 31: 3126–3135.
Wohlschlegel JA, Johnson ES, Reed SI, Yates JR 3rd . Global analysis of protein sumoylation in Saccharomyces cerevisiae . J Biol Chem 2004; 279: 45662–45668.
Wu J, Trumbly RJ . Multiple regulatory proteins mediate repression and activation by interaction with the yeast Mig1 binding site. Yeast (Chichester, England) 1998; 14: 985–1000.
Yang X, Hubbard EJ, Carlson M . A protein kinase substrate identified by the two-hybrid system. Science (New York, NY) 1992; 257: 680–682.
Sneddon MW, Faeder JR, Emonet T . Efficient modelling, simulation and coarse-graining of biological complexity with NFsim. Nat Methods 2011; 8: 177–183.
Orth JD, Palsson BO . Systematizing the generation of missing metabolic knowledge. Biotechnol Bioeng 2010; 107: 403–412.
Garcia-Salcedo R, Lubitz T, Beltran G, Elbing K, Tian Y, Frey S et al. Glucose de-repression by yeast AMP-activated protein kinase SNF1 is controlled via at least two independent steps. FEBS J 2014; 281: 1901–1917.
Finley D, Ulrich HD, Sommer T, Kaiser P . The ubiquitin-proteasome system of Saccharomyces cerevisiae . Genetics 2012; 192: 319–360.
Hedbacker K, Carlson M . SNF1/AMPK pathways in yeast. Front Biosci 2008; 13: 2408–2420.
Chandrashekarappa DG, McCartney RR, Schmidt MC . Subunit and domain requirements for adenylate-mediated protection of Snf1 kinase activation loop from dephosphorylation. J Biol Chem 2011; 286: 44532–44541.
Mayer FV, Heath R, Underwood E, Sanders MJ, Carmena D, McCartney RR et al. ADP regulates SNF1, the Saccharomyces cerevisiae homolog of AMP-activated protein kinase. Cell Metab 2011; 14: 707–714.
Wilson MA, Koutelou E, Hirsch C, Akdemir K, Schibler A, Barton MC et al. Ubp8 and SAGA regulate Snf1 AMP kinase activity. Mol Cell Biol 2011; 31: 3126–3135.
Simpson-Lavy KJ, Johnston M . SUMOylation regulates the SNF1 protein kinase. Proc Natl Acad Sci USA 2013; 110: 17432–17437.
Acknowledgements
We thank Max Flöttmann for his helpful support with the Boolean simulations and their visualisations, and Falko Krause for the initial setup of the software tools we used for the literature curation. This work was supported by the European Commission via the UNICELLSYS project (contract 201142) to S.H. and E.K. and ISOLATE (contract 28995) to S.H., by the German Federal Ministry of Education and Research: OncoPath (FKZ0316184B) to E.K., e:Bio Cellemental (FKZ0316193) to M.K. and by the Swedish Research Council to S.H.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lubitz, T., Welkenhuysen, N., Shashkova, S. et al. Network reconstruction and validation of the Snf1/AMPK pathway in baker’s yeast based on a comprehensive literature review. npj Syst Biol Appl 1, 15007 (2015). https://doi.org/10.1038/npjsba.2015.7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/npjsba.2015.7
This article is cited by
-
A scalable method for parameter-free simulation and validation of mechanistic cellular signal transduction network models
npj Systems Biology and Applications (2020)
-
Transcriptional regulatory proteins in central carbon metabolism of Pichia pastoris and Saccharomyces cerevisiae
Applied Microbiology and Biotechnology (2020)