Abstract
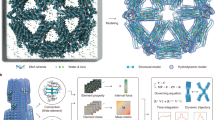
Hierarchical assemblies of biomolecular subunits can carry out versatile tasks at the cellular level with remarkable spatial and temporal precision1,2. As an example, the collective motion and mutual cooperation between complex protein machines mediate essential functions for life, such as replication3, synthesis4, degradation5, repair6 and transport7. Nucleic acid molecules are far less dynamic than proteins and need to bind to specific proteins to form hierarchical structures. The simplest example of these nucleic acid-based structures is provided by a rod-shaped tobacco mosaic virus, which consists of genetic material surrounded by coat proteins8. Inspired by the complexity and hierarchical assembly of viruses, a great deal of effort has been devoted to design similarly constructed artificial viruses9,10. However, such a wrapping approach makes nucleic acid dynamics insensitive to environmental changes. This limitation generally restricts, for example, the amplification of the conformational dynamics between the right-handed B form to the left-handed Z form of double-stranded deoxyribonucleic acid (DNA)11,12. Here we report a virus-like hierarchical assembly in which the native DNA and a synthetic coat undergo repeated collective helicity switching triggered by pH change under physiological conditions. We also show that this collective helicity inversion occurs during translocation of the DNA–coat assembly into intracellular compartments. Translating DNA conformational dynamics into a higher level of hierarchical dynamics may provide an approach to create DNA-based nanomachines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
11 April 2017
In the version of this Letter originally published, Fig. 1 contained chemical structures that were not drawn in our house stlye. These structures have been replaced in all versions of the Letter.
References
Kinbara, K. & Aida, T. Toward intelligent molecular machines: directed motions of biological and artificial molecules and assemblies. Chem. Rev. 105, 1377–1400 (2005).
Ishii, D. et al. Chaperonin-mediated stabilization and ATP-triggered release of semiconductor nanoparticles. Nature 423, 628–632 (2003).
Sun, J. et al. Cdc6-induced conformational changes in ORC bound to origin DNA revealed by cryo-electron microscopy. Structure 20, 534–544 (2012).
Alberts, B. et al. The cell as a collection of protein machines: preparing the next generation of molecular biologists. Cell 92, 291–294 (1998).
Olivares, A. O. et al. Mechanochemical basis of protein degradation by a double-ring A+A+A+ machine. Nat. Struct. Mol. Biol. 21, 871–875 (2014).
Saibil, H. et al. Chaperone machines for protein folding, unfolding and disaggregation. Nat. Rev. Mol. Cell Biol. 14, 630–642 (2013).
Ryan, K. J. et al. The nuclear pore complex: a protein machine bridging the nucleus and cytoplasm. Curr. Opin. Cell Biol. 12, 361–371 (2000).
Butler, P. J. G. et al. Self-assembly of tobacco mosaic virus: the role of an intermediate aggregate in generating both specificity and speed. Phil. Trans. R. Soc. Lond. 354, 537–550 (1999).
Lim, Y.-B. et al. Filamentous artificial virus from a self-assembled discrete nanoribbon. Angew. Chem. Int. Ed. 47, 4525–4528 (2008).
Hernandez-Garcia, A. et al. Design and self-assembly of simple coat proteins for artificial viruses. Nat. Nanotech. 9, 698–702 (2014).
Fuertes, M. A. et al. Molecular mechanisms for the B-Z transition in the example of poly[d(G-C).d((G-C)) polymers. A critical review. Chem. Rev. 106, 2045–2064 (2006).
Choi, J. et al. Conformational changes of non-B DNA. Chem. Soc. Rev. 40, 5893–5909 (2011).
Kim, H.-J. et al. Self-dissociating tubules from helical stacking of noncovalent macrocycles. Angew. Chem. Int. Ed. 139, 8471–8475 (2010).
Huang, Z . et al. Pulsating tubules from noncovalent macrocycles. Science 337, 1521–1526, (2012).
Kypr, J . et al. Circular dichroism and conformational polymorphism of DNA. Nucl. Acids Res. 37, 1713–1725 (2009).
Benevides, J. M. et al. Characterization of DNA structures by Raman spectroscopy: high-salt and low-salt forms of double helical poly(dG-dC) in H2O and D2O solutions and application to B, Z and A-DNA. Nucl. Acids Res. 11, 5747–5761 (1983).
Misra, V. K. et al. The electrostatic contribution to the B-to-Z transition of DNA. Biochemistry 35, 1115–1124 (1996).
Sugiyama, H. et al. Synthesis, structure and thermodynamic properties of 8-methylguanine-containing oligonucleotides: Z-DNA under physiological salt conditions. Nucleic Acids Res. 24, 1272–1278 (1996).
Barone, G . et al. Confinement effects on the interaction of native DNA with Cu(II)-5-(triethylammoniunethyl) salicylidene ortho-pheylendiiminate in C12E4 liquid crystals. Dalton Trans. 4172–4178 (2008).
Callahan, L. et al. B- to Z-DNA transition probed by oligonucleotides containing methylphosphonates. Proc. Natl Acad. Sci. USA 83, 1617–1621 (1986).
Zacharias, W . et al. Conditions which cause the right-handed to left-handed DNA conformational transitions. J. Biol. Chem. 257, 2775–2782 (1982).
El-Sayed, A. & Harashima, H. Endocytosis of gene delivery vectors: from clathrin-dependent to lipid raft-mediated endocytosis. Mol. Ther. 21, 1118–1130 (2013).
Geisow, M. J. & Evans, W. H. pH in the endosome. Measurements during pinocytosis and receptor-mediated endocytosis. Exp. Cell Res. 150, 36–46 (1984).
Kunitz, M. Crystalline desoxyribonuclease; isolation and general properties; spectrophotometric method for the measurement of desoxyribonuclease activity. J. Gen. Physiol. 33, 349–362 (1950).
Herbert, A. & Rich, A. Left-handed Z-DNA: structure and function. Genetica 106, 37–47 (1999).
Kim, Y. et al. Open–closed switching of synthetic tubular pores. Nat. Commun. 6, 8650 (2015).
Gift, A. D., Stewart, S. M. & Bokashanga, P. K. Experimental determination of pKa values by use of NMR chemical shifts, revisited. J. Chem. Educ. 89, 1458–1460 (2012).
Beniash, E. et al. Self-assembling peptide amphiphile nanofiber matrices for cell entrapment. Acta Biomater. 1, 387–397 (2005).
Acknowledgements
This work was supported by the 1000 Program, and the National Natural Science Foundation China (no. 51473062, no. 21574055, no. 21634005 and no. 21550110493).
Author information
Authors and Affiliations
Contributions
Y.K. designed and performed most of the experiments, H.L. synthesized molecules and carried out AFM experiments, Y.H. performed TEM experiments, X.C. carried out ITC experiments, X.M. performed cell cultures and imaging experiments and M.L. developed the concept, supervised the research and wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 15825 kb)
Rights and permissions
About this article
Cite this article
Kim, Y., Li, H., He, Y. et al. Collective helicity switching of a DNA–coat assembly. Nature Nanotech 12, 551–556 (2017). https://doi.org/10.1038/nnano.2017.42
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2017.42
This article is cited by
-
Reversible chirality inversion of an AuAgx-cysteine coordination polymer by pH change
Nature Communications (2024)
-
Magnetic control of self-assembly and disassembly in organic materials
Nature Communications (2023)
-
Nanoparticles exhibiting virus-mimic surface topology for enhanced oral delivery
Nature Communications (2023)
-
Enantiocontrolled macrocyclization by encapsulation of substrates in chiral capsules
Nature Synthesis (2023)
-
A General Approach for Synthesis of Circularly Assembled Supramolecular Polymers by Means of Region-confined Amphiphilic Supramolecular Polymerization
Chemical Research in Chinese Universities (2023)