Abstract
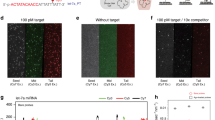
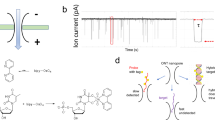
Small RNA molecules have an important role in gene regulation and RNA silencing therapy, but it is challenging to detect these molecules without the use of time-consuming radioactive labelling assays or error-prone amplification methods. Here, we present a platform for the rapid electronic detection of probe-hybridized microRNAs from cellular RNA. In this platform, a target microRNA is first hybridized to a probe. This probe:microRNA duplex is then enriched through binding to the viral protein p19. Finally, the abundance of the duplex is quantified using a nanopore. Reducing the thickness of the membrane containing the nanopore to 6 nm leads to increased signal amplitudes from biomolecules, and reducing the diameter of the nanopore to 3 nm allows the detection and discrimination of small nucleic acids based on differences in their physical dimensions. We demonstrate the potential of this approach by detecting picogram levels of a liver-specific miRNA from rat liver RNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene LIN-4 encodes small RNAs with antisense complementarity to LIN-14. Cell 75, 843–854 (1993).
Carthew, R. W. & Sontheimer, E. J. Origins and mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Winter, J., Jung, S., Keller, S., Gregory, R. I. & Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nature Cell Biol. 11, 228–234 (2009).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nature Rev. Drug Discov. 8, 129–138 (2009).
Oh, Y. K. & Park, T. G. siRNA delivery systems for cancer treatment. Adv. Drug Deliv. Rev. 61, 850–862 (2009).
Singh, S. K. & Hajeri, P. B. siRNAs: their potential as therapeutic agents—part II. Methods of delivery. Drug Discov. Today 14, 859–865 (2009).
Murphy, J. & Bustin, S. A. Reliability of real-time reverse-transcription PCR in clinical diagnostics: gold standard or substandard? Expert Rev. Mol. Diagnostics 9, 187–197 (2009).
Coulter, W. H. US patent 2,656,508 (1953).
Bezrukov, S. M., Vodyanoy, I. & Parsegian, V. A. Counting polymers moving through a single-ion channel. Nature 370, 279–281 (1994).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996).
Gu, L. Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686–690 (1999).
Butler, T. Z., Pavlenok, M., Derrington, I. M., Niederweis, M. & Gundlach, J. H. Single-molecule DNA detection with an engineered MspA protein nanopore. Proc. Natl Acad. Sci. USA 105, 20647–20652 (2008).
Wendell, D. et al. Translocation of double-stranded DNA through membrane-adapted phi29 motor protein nanopores. Nature Nanotech. 4, 765–772 (2009).
Howorka, S. & Siwy, Z. Nanopore analytics: sensing of single molecules. Chem. Soc. Rev. 38, 2360–2384 (2009).
Li, J. et al. Ion-beam sculpting at nanometre length scales. Nature 412, 166–169 (2001).
Storm, A. J., Chen, J. H., Ling, X. S., Zandbergen, H. W. & Dekker, C. Fabrication of solid-state nanopores with single-nanometre precision. Nature Mater. 2, 537–540 (2003).
Li, J. L., Gershow, M., Stein, D., Brandin, E. & Golovchenko, J. A. DNA molecules and configurations in a solid-state nanopore microscope. Nature Mater. 2, 611–615 (2003).
Heng, J. B. et al. The electromechanics of DNA in a synthetic nanopore. Biophys. J. 90, 1098–1106 (2006).
Iqbal, S. M., Akin, D. & Bashir, R. Solid-state nanopore channels with DNA selectivity. Nature Nanotech. 2, 243–248 (2007).
Fologea, D., Ledden, B., McNabb, D. S. & Li, J. L. Electrical characterization of protein molecules by a solid-state nanopore. Appl. Phys. Lett. 91, 053901 (2007).
Zhao, Q. et al. Stretching and unzipping nucleic acid hairpins using a synthetic nanopore. Nucleic Acids Res. 36, 1532–1541 (2008).
Tabard-Cossa, V. et al. Single-molecule bonds characterized by solid-state nanopore force spectroscopy. ACS Nano 3, 3009–3014 (2009).
Skinner, G. M., van den Hout, M., Broekmans, O., Dekker, C. & Dekker, N. H. Distinguishing single- and double-stranded nucleic acid molecules using solid-state nanopores. Nano Lett. 9, 2953–2960 (2009).
Kowalczyk, S. W., Hall, A. R. & Dekker, C. Detection of local protein structures along DNA using solid-state nanopores. Nano Lett. 10, 324–328 (2010).
Smeets, R. M. M., Kowalczyk, S. W., Hall, A. R., Dekker, N. H. & Dekker, C. Translocation of RecA-coated double-stranded DNA through solid-state nanopores. Nano Lett. 9, 3089–3095 (2009).
Wanunu, M., Sutin, J. & Meller, A. DNA profiling using solid-state nanopores: detection of DNA-binding molecules. Nano Lett. 9, 3498–3502 (2009).
Singer, A. et al. Nanopore based sequence specific detection of duplex DNA for genomic profiling. Nano Lett. 10, 738–742 (2010).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nature Biotechnol. 26, 1146–1153 (2008).
Garaj, S. et al. Graphene as a subnanometre trans-electrode membrane. Nature 467, 190–193 (2010).
Schneider, G. F. et al. DNA translocation through graphene nanopores. Nano Lett. 10, 3163–3167 (2010).
Merchant, C. A. et al. DNA translocation through graphene nanopores. Nano Lett. 10, 2915–2921 (2010).
Krutzfeldt, J. Silencing of microRNAs in vivo with et al. ‘antagomirs’. Nature 438, 685–689 (2005).
Bai, S. M. et al. MicroRNA-122 inhibits tumorigenic properties of hepatocellular carcinoma cells and sensitizes these cells to sorafenib. J. Biol. Chem. 284, 32015–32027 (2009).
Lanford, R. E. et al. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science 327, 198–201 (2010).
Park, N. M., Choi, C. J., Seong, T. Y. & Park, S. J. Quantum confinement in amorphous silicon quantum dots embedded in silicon nitride. Phys. Rev. Lett. 86, 1355–1357 (2001).
Kim, M. J., Wanunu, M., Bell, D. C. & Meller, A. Rapid fabrication of uniformly sized nanopores and nanopore arrays for parallel DNA analysis. Adv. Mater. 18, 3149–3155 (2006).
Wu, M. Y. et al. Control of shape and material composition of solid-state nanopores. Nano Lett. 9, 479–484 (2009).
Smeets, R. M. M. et al. Salt dependence of ion transport and DNA translocation through solid-state nanopores. Nano Lett. 6, 89–95 (2006).
Hall, J. E. Access resistance of a small circular pore. J. Gen. Physiol. 66, 531–532 (1975).
Vodyanoy, I. & Bezrukov, S. M. Sizing of an ion pore by access resistance measurements. Biophys. J. 62, 10–11 (1992).
Akeson, M., Branton, D., Kasianowicz, J. J., Brandin, E. & Deamer, D. W. Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys. J. 77, 3227–3233 (1999).
Mathe, J., Aksimentiev, A., Nelson, D. R., Schulten, K. & Meller, A. Orientation discrimination of single-stranded DNA inside the alpha-hemolysin membrane channel. Proc. Natl Acad. Sci. USA 102, 12377–12382 (2005).
Wanunu, M., Sutin, J., McNally, B., Chow, A. & Meller, A. DNA translocation governed by interactions with solid-state nanopores. Biophys. J. 95, 4716–4725 (2008).
Aksimentiev, A. Deciphering ionic current signatures of DNA transport through a nanopore. Nanoscale 2, 468–483 (2010).
Lide, D. R. (ed.) Ionic conductivity and diffusion at infinite dilution, in CRC Handbook of Chemistry and Physics (CRC Press, 2005).
Wanunu, M., Morrison, W., Rabin, Y., Grosberg, A. Y. & Meller, A. Electrostatic focusing of unlabelled DNA into nanoscale pores using a salt gradient. Nature Nanotech. 5, 160–165 (2010).
Luan, B. Q. & Aksimentiev, A. Electro-osmotic screening of the DNA charge in a nanopore. Phys. Rev. E 78, 4 (2008).
van Dorp, S., Keyser, U. F., Dekker, N. H., Dekker, C. & Lemay, S. G. Origin of the electrophoretic force on DNA in solid-state nanopores. Nature Phys. 5, 347–351 (2009).
Gu, L. Q., Cheley, S. & Bayley, H. Electroosmotic enhancement of the binding of a neutral molecule to a transmembrane pore. Proc. Natl Acad. Sci. USA 100, 15498–15503 (2003).
Kasianowicz, J. J., Nguyen, T. L. & Stanford, V. M. Enhancing molecular flux through nanopores by means of attractive interactions. Proc. Natl Acad. Sci. USA 103, 11431–11432 (2006).
Bissels, U. et al. Absolute quantification of microRNAs by using a universal reference. RNA 15, 2375–2384 (2009).
Foldes-Papp, Z. et al. Trafficking of mature miRNA-122 into the nucleus of live liver cells. Curr. Pharm. Biotechnol. 10, 569–578 (2009).
Vargason, J. M., Szittya, G., Burgyan, J. & Hall, T. M. T. Size selective recognition of siRNA by an RNA silencing suppressor. Cell 115, 799–811 (2003).
Jin, J., Cid, M., Poole, C. B. & McReynolds, L. A. Protein-mediated miRNA detection and siRNA enrichment using p19. Biotechniques 48, 17–23 (2010).
Wu, M. Y., Krapf, D., Zandbergen, M., Zandbergen, H. & Batson, P. E. Formation of nanopores in a SiN/SiO2 membrane with an electron beam. Appl. Phys. Lett. 87, 113106 (2005).
van den Hout, M. et al. Controlling nanopore size, shape and stability. Nanotechnology 21, 115304 (2010).
Acknowledgements
The authors would like to thank M. Fischbein for assistance with optimizing the plasma etch process, the laboratory of C. Johnson for the use of their plasma etcher, D. Yates for assistance with ADF-STEM, and G. Rosenbloom and B. Cooperman for the tRNA molecule. The authors also thank J. Bartel, J. Fairfield, M. Goulian, K. Healy, R. Johnson, C. Johnson, N. Peterman, G.V. Soni and L. Willis for helpful advice regarding the manuscript. This work was supported by the National Institutes of Health (grant no. R21HG004767) and the Penn Genomics Frontier Institute. T.D. acknowledges support from the Israel Science Foundation (Bikura postdoctoral fellowship).
Author information
Authors and Affiliations
Contributions
M.W. and L.M. conceived and designed the experiments. T.D., V.R. and M.W. designed, fabricated and characterized the nanopore devices. J.J. and L.M. developed and performed the miRNA enrichment protocols. M.W. performed the nanopore experiments and analysed the data. M.W., T.D., L.M. and M.D. wrote the manuscript, and all other authors commented on it.
Corresponding authors
Ethics declarations
Competing interests
J.J. and L.M. are employees of New England BioLabs, a company that sells p19 and other proteins for RNA and DNA research.
Supplementary information
Supplementary information
Supplementary information (PDF 1743 kb)
Rights and permissions
About this article
Cite this article
Wanunu, M., Dadosh, T., Ray, V. et al. Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nature Nanotech 5, 807–814 (2010). https://doi.org/10.1038/nnano.2010.202
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2010.202
This article is cited by
-
Nanopore sequencing of DNA-barcoded probes for highly multiplexed detection of microRNA, proteins and small biomarkers
Nature Nanotechnology (2023)
-
Breakdown of the Nernst–Einstein relation in carbon nanotube porins
Nature Nanotechnology (2023)
-
Simultaneous identification of viruses and viral variants with programmable DNA nanobait
Nature Nanotechnology (2023)
-
Optimizing the sensitivity and resolution of hyaluronan analysis with solid-state nanopores
Scientific Reports (2022)
-
Focus on using nanopore technology for societal health, environmental, and energy challenges
Nano Research (2022)