Abstract
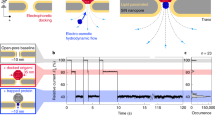
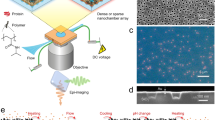
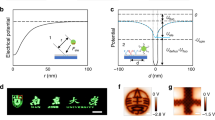
The development of systematic approaches to explore protein–protein interactions and dynamic protein networks is at the forefront of biological sciences. Nanopatterned protein arrays offer significant advantages for sensing applications, including short diffusion times, parallel detection of multiple targets and the requirement for only tiny amounts of sample1,2,3. Atomic force microscopy (AFM) based techniques have successfully demonstrated patterning of molecules, including stable proteins, with submicrometre resolution4,5,6,7,8,9,10,11,12,13,14,15. Here, we introduce native protein nanolithography for the nanostructured assembly of even fragile proteins or multiprotein complexes under native conditions. Immobilized proteins are detached by a novel vibrational AFM mode (contact oscillation mode) and replaced by other proteins, which are selectively self-assembled from the bulk. This nanolithography permits rapid writing, reading and erasing of protein arrays in a versatile manner. Functional protein complexes may be assembled with uniform orientation at dimensions down to 50 nm. Such fabrication of two-dimensionally arranged nano-objects with biological activity will prove powerful for proteome-wide interaction screens and single molecule/virus/cell analyses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
MacBeath, G. & Schreiber, S. L. Printing proteins as microarrays for high-throughput function determination. Science 289, 1760–1763 (2000).
Zhu, H. et al. Global analysis of protein activities using proteome chips. Science 293, 2101–2105 (2001).
Lynch, M. et al. Functional protein nanoarrays for biomarker profiling. Proteomics 4, 1695–1702 (2004).
Unal, K., Frommer, J. & Wickramasinghe, H. K. Ultrafast molecule sorting and delivery by atomic force microscopy. Appl. Phys. Lett. 88, 183105 (2006).
Jaschke, M. & Butt, H.-J. Deposition of organic material by the tip of a scanning force microscope. Langmuir 11, 1061–1064 (1995).
Ginger, D. S., Zhang, H. & Mirkin, C. A. The evolution of dip-pen nanolithography. Angew. Chem. Int. Edn 43, 30–45 (2004).
Lee, K. B., Park, S. J., Mirkin, C. A., Smith, J. C. & Mrksich, M. Protein nanoarrays generated by dip-pen nanolithography. Science 295, 1702–1705 (2002).
Piner, R. D., Zhu, J., Xu, F., Hong, S. & Mirkin, C. A. ‘Dip-Pen’ nanolithography. Science 283, 661–663 (1999).
Nam, J. M. et al. Bioactive protein nanoarrays on nickel oxide surfaces formed by dip-pen nanolithography. Angew. Chem. Int. Edn 43, 1246–1249 (2004).
Xu, S. & Liu, G.-Y. Nanometer-scale fabrication by simultanuous nanoshaving and molecular self-assembly. Langmuir 13, 127–129 (1997).
Wadu-Mesthrige, K., Xu, S., Amro, N. A. & Liu, G. Y. Fabrication and imaging of nanometer-sized protein patterns. Langmuir 15, 8580–8583 (1999).
Wadu-Mesthrige, K., Amro, N. A., Garno, J. C., Xu, S. & Liu, G. Fabrication of nanometer-sized protein patterns using atomic force microscopy and selective immobilization. Biophys. J. 80, 1891–1899 (2001).
Xu, S., Miller, S., Laibinis, P. E. & Liu, G. Y. Fabrication of nanometer scale patterns within self-assembled monolayers by nanografting. Langmuir 15, 7244–7251 (1999).
Wouters, D. & Schubert, U. S. Nanolithography and nanochemistry: probe-related patterning techniques and chemical modification for nanometer-sized devices. Angew. Chem. Int. Edn 43, 2480–2495 (2004).
Bruckbauer, A. et al. Multicomponent submicron features of biomolecules created by voltage controlled deposition from a nanopipet. J. Am. Chem. Soc. 125, 9834–9839 (2003).
Hu, Y., Das, A., Hecht, M. H. & Scoles, G. Nanografting de novo proteins onto gold surfaces. Langmuir 21, 9103–9109 (2005).
Tinazli, A. et al. High-affinity chelator thiols for switchable and oriented immobilization of histidine-tagged proteins: a generic platform for protein chip technologies. Chem. Eur. J. 11, 5249–5259 (2005).
Seemüller, E. et al. Proteasome from Thermoplasma acidophilum—a threonine protease. Science 268, 579–582 (1995).
Voges, D., Zwickl, P. & Baumeister, W. The 26S proteasome: A molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 68, 1015–1068 (1999).
Zwickl, P., Voges, D. & Baumeister, W. The proteasome: a macromolecular assembly designed for controlled proteolysis. Phil. Trans. R. Soc. B 354, 1501–1511 (1999).
Bochtler, M., Ditzel, L., Groll, M., Hartmann, C. & Huber, R. The proteasome. Annu. Rev. Biophys. Biomol. Struct. 28, 295–317 (1999).
Löwe, J. et al. Crystal structure of the 20S proteasome from the archaeon T. acidophilum at 3.4 Å resolution. Science 268, 533–539 (1995).
Hutschenreiter, S., Tinazli, A., Model, K. & Tampé, R. Two-substrate association with the 20S proteasome at single-molecule level. EMBO J. 23, 2488–2497 (2004).
Spurlino, J. C., Lu, G. Y. & Quiocho, F. A. The 2.3-Å resolution structure of the maltose- or maltodextrin-binding protein, a primary receptor of bacterial active transport and chemotaxis. J. Biol. Chem. 266, 5202–5219 (1991).
Waldeck, D. H., Alivisatos, A. P. & Harris, C. B. Nonradiative damping of molecular electronic excited states by metal surfaces. Surf. Sci. 158, 103–125 (1985).
Lata, S. & Piehler, J. Stable and functional immobilization of histidine-tagged proteins via multivalent chelator headgroups on a molecular poly(ethylene glycol) brush. Anal. Chem. 77, 1096–1105 (2005).
Piehler, J. & Schreiber, G. Fast transient cytokine-receptor interactions monitored in real time by reflectometric interference spectroscopy. Anal. Biochem. 289, 173–186 (2001).
Butt, H.-J., Wang, D. N., Hansma, P. K. & Kühlbrandt, W. Effect of surface roughness of carbon support films on high-resolution electron diffraction of two-dimensional protein crystals. Ultramicroscopy 36, 307–318 (1991).
Butt, H.-J., Müller, T. & Gross, H. Immobilizing biomolecules for scanning force microscopy by embedding in carbon. J. Struct. Biol. 110, 127–132 (1993).
Hegner, M., Wagner, P. & Semenza, G. Ultralarge atomically flat template-stripped Au surfaces for scanning probe microscopy. Surf. Sci. 291, 39–46 (1993).
Acknowledgements
We thank Gerhard Spatz-Kümbel for technical assistance, Suman Lata and Annett Reichel for providing OG488MBP-His10, Eva Jaks for site-specifically biotinylated IFNα2, and Alart Mulder and Katrin Schulze for stimulating discussions. The work was supported by grants from the Federal Ministry of Education and Research (BMBF) (grant program: Nanobiotechnology) and the Deutsche Forschungsgemeinschaft (DFG).
Author information
Authors and Affiliations
Contributions
A.T. and R.T. conceived and designed the experiments, A.T. and M.B. performed the experiments, A.T., J.P., M.B., R.G., and R.T. analysed the data, and A.T. and R.T. co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary movie S1 (MOV 1884 kb)
Supplementary Information
Supplementary figures S1 and S2 (PDF 424 kb)
Rights and permissions
About this article
Cite this article
Tinazli, A., Piehler, J., Beuttler, M. et al. Native protein nanolithography that can write, read and erase. Nature Nanotech 2, 220–225 (2007). https://doi.org/10.1038/nnano.2007.63
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2007.63
This article is cited by
-
Probing fibronectin adsorption on chemically defined surfaces by means of single molecule force microscopy
Scientific Reports (2020)
-
Atomic force microscopy-based characterization and design of biointerfaces
Nature Reviews Materials (2017)
-
Arrays of Individual DNA Molecules on Nanopatterned Substrates
Scientific Reports (2017)
-
Multivalent chelators for spatially and temporally controlled protein functionalization
Analytical and Bioanalytical Chemistry (2014)
-
Investigating bioconjugation by atomic force microscopy
Journal of Nanobiotechnology (2013)