Abstract
Optical clearing methods facilitate deep biological imaging by mitigating light scattering in situ. Multi-scale high-resolution imaging requires preservation of tissue integrity for accurate signal reconstruction. However, existing clearing reagents contain chemical components that could compromise tissue structure, preventing reproducible anatomical and fluorescence signal stability. We developed ScaleS, a sorbitol-based optical clearing method that provides stable tissue preservation for immunochemical labeling and three-dimensional (3D) signal rendering. ScaleS permitted optical reconstructions of aged and diseased brain in Alzheimer's disease models, including mapping of 3D networks of amyloid plaques, neurons and microglia, and multi-scale tracking of single plaques by successive fluorescence and electron microscopy. Human clinical samples from Alzheimer's disease patients analyzed via reversible optical re-sectioning illuminated plaque pathogenesis in the z axis. Comparative benchmarking of contemporary clearing agents showed superior signal and structure preservation by ScaleS. These findings suggest that ScaleS is a simple and reproducible method for accurate visualization of biological tissue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dodt, H.-U. et al. Ultramicroscopy: 3D visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Ertürk, A. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 7, 1983–1995 (2012).
Ke, M.-T., Fujimoto, S. & Imai, T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 16, 1154–1161 (2013).
Kuwajima, T. et al. ClearT: a detergent- and solvent-free clearing method for neuronal and non-neuronal tissue. Development 140, 1364–1368 (2013).
Susaki, E.A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Tomer, R., Ye, L., Hsueh, B. & Deisseroth, K. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat. Protoc. 9, 1682–1697 (2014).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Saito, T. et al. Single App knock-in mouse models of Alzheimer′s disease. Nat. Neurosci. 17, 661–663 (2014).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Wang, H. et al. High-speed mapping of synaptic connectivity usingphotostimulation in Channelrhodopsin-2 transgenic mice. Proc. Natl. Acad. Sci. USA 104, 8143–8148 (2007).
Wilcock, D.M. et al. Intracranially administered anti-A beta antibodies reduce beta-amyloid deposition by mechanisms both independent of and associated with microglial activation. J. Neurosci. 23, 3745–3751 (2003).
Aho, L. et al. Immunohistochemical visualization of amyloid-β protein precursor and amyloid-β in extra- and intracellular compartments in the human brain. J. Alzheimers Dis. 20, 1015–1028 (2010).
Ono, M. et al. Push–pull benzothiazole derivatives as probes for detecting β-amyloid plaques in Alzheimer's brains. Bioorg. Med. Chem. 17, 7002–7007 (2009).
Kumar-Singh, S. et al. Dense-core plaques in Tg2576 and PSAPP mouse models of Alzheimer's disease are centered on vessel walls. Am. J. Pathol. 167, 527–543 (2005).
Ozaki, H.S. & Walsten, D. Prental formation of the normal mouse corpus callosum: a quantitative study with carbocyanine dye. J. Comp. Neurol. 323, 81–90 (1992).
Thal, D.R. & Capetillo-Zarate, E. Tracing of temporo-entorhinal connections in the human brain: cognitively impaired argyrophilic grain disease cases show of temporo-entorhinal association neurons. Acta Neuropathol. 115, 175–183 (2008).
Murak, R.E. Microglia in Alzheimer brain: a neuropathological perspective. Int. J. Alzheimers Dis. published online, doi:10.1155/2012/165021 (13 May 2012).
Torres-Platas, S.G. et al. Morphometric characterization of microglial phenotypes in human cerebral cortex. J. Neuroinflammation 11, 1–13 (2014).
Kozlowski, C. & Weimer, R.M. An automated method to quantify microglia morphology and application to monitor activation state. PLoS ONE 7, e31814 (2012).
Karperien, A., Ahammer, H. & Jelinek, H.F. Quantitating the subtleties of microglial morphology with fractal analysis. Front. Cell. Neurosci. 7, 3–18 (2013).
Fiala, J.C. Mechanisms of amyloid plaque pathogenesis. Acta Neuropathol. 114, 551–571 (2007).
Thal, D.R., Capetillo-Zarate, E., Tredici, K.D. & Braak, H. The development of amyloid β protein deposits in the aged brain. Sci. Aging Knowledge Environ. 2006, re1 (2006).
D'Andrea, M.R., Cole, G.M. & Ard, M.D. The microglial phagocytic role with specific plaque types in the Alzheimer disease brain. Neurobiol. Aging 25, 675–683 (2004).
Hou, B. et al. Scalable and DiI-compatible optical clearance of the mammalian brain. Front. Neuroanat. 9, 19 (2015).
Aoyagi, Y., Kawakami, R., Osanai, H., Hibi, T. & Nemoto, T. A rapid optical clearing protocol using 2,2′-thiodiethanol for microscopic observation of fixed mouse brain. PLoS ONE 10, e0116280 (2015).
Richardson, D.S. & Lichtman, J.W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Davanlou, M. & Smith, D.F. Unbiased stereological estimation of different cell types in rat cerebral cortex. Image Anal. Stereol. 23, 1–11 (2004).
Serrano-Pozo, A. et al. Differential relationships of reactive astrocytes and microglia to fibrillar amyloid deposits in Alzheimer disease. J. Neuropathol. Exp. Neurol. 72, 462–471 (2013).
Chiang, A.-S. et al. Insect NMDA receptors mediate juvenile hormone biosynthesis. Proc. Natl. Acad. Sci. USA 99, 37–42 (2002).
Vidal, R. et al. The Psen1–L166P-knock-in mutant leads to amyloid deposition in human wild-type amyloid precursor protein YAC transgenic mice. FASEB J. 26, 2899–2910 (2012).
Zhu, D.-Y. et al. Expression of inducible nitric oxide synthetase after focal cerebral ischemia stimulates neurogenesis in adult rodent dentate gyrus. J. Neurosci. 23, 223–229 (2003).
Yoshikawa, F. et al. Phospholipase D family member 4, a transmembrane glycoprotein with no phospholipase D activity, expression in spleen and early postnatal microglia. PLoS ONE 5, e13932 (2010).
Condello, C., Schain, A. & Grutsedler, J. Multicolor time-stamp reveals the dynamics and toxicity of amyloid deposition. Sci. Rep. 1, 19 (2011).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. SMC-9, 62–66 (1979).
Oh, K.-J. et al. Staging of Alzheimer's pathology in triple transgenic mice: a light and electron microscopic analysis. Int. J. Alzheimers Dis. 2010, 1–24 (2010).
Acknowledgements
We thank H. Sakurai for general assistance, RIKEN BSI-Olympus Collaboration Center for technical support, K. Higuchi, K. Ishihara, D. Nishiwaki, Y. Ue, K. Okazaki, A. Yaguchi, Y. Sato, H. Sakuma, K. Koga and B. Zimmermann for help with acquiring and analyzing LM images, K. Okamoto-Furuta and H. Kohda (Division of Electron Microscopic Study, Center for Anatomical Studies, Graduate School of Medicine, Kyoto University) for their technical assistance in EM, G. Augustine (Duke University) for the ChR2-YFP Tg mice, J.R. Sanes (Harvard University) for the YFP-H line, R. Takahashi, Dr. K. Sohya and T. Tsumoto for advice on Tg mice, and C. Yokoyama and A. Terashima for critical reading and editing of the manuscript. This work was supported in part by grants from the Japan Ministry of Education, Culture, Sports, Science and Technology Grant-in-Aid for Scientific Research on Priority Areas, the Human Frontier Science Program, and the Brain Mapping by Integrated Neurotechnologies for Disease Studies (Brain/MINDS) from Japan Agency for Medical Research and development, AMED.
Author information
Authors and Affiliations
Contributions
A.M., H. Hama, H. Hioki and T.H. conceived the study. A.M., H. Hama, K.N., H. Hioki, T.H. and H.K. planned and executed the LM experiments and analyzed the data. H. Hioki, T.K. and T.A. planned and executed the EM experiments. H.K. devised the algorithms for data analysis. F.I. contributed to image data acquisition. T. Saido and T. Saito supervised the experiments that used AD mouse model and patient brains. A.M. wrote the manuscript and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Experimental protocols for ScaleA2, ScaleS, AbScale, ChemScale, and ScaleSQ
Tissue clearing protocols that use ScaleA2, ScaleS, and ScaleSQ solutions, and AbScale and ChemScale clearing/labeling protocols with relevant theoretical background.
In each protocol, a brain sample is incubated in a large volume (> 25 ml/g tissue) of the solution.
(a) ScaleA2 protocol. See the original paper (Ref. 2) for details.
(b) ScaleS protocol. The formulas of ScaleS solutions are shown in Supplementary Table 1. This procedure assumes that an adult mouse cerebral hemisphere is processed as a sample to be cleared. When mouse brain slices are used, the incubation time for each step can be substantially shortened.
First, permeability of a sample is enhanced by incubation in ScaleS0 solution containing 20% sorbitol, 5% glycerol, 1 mM methyl-β-cyclodextrin, 1 mM γ-cyclodextrin, 1% N-acetyl-L-hydroxyproline, and 3% DMSO for 12 hrs. Methyl-β-cyclodextrin and γ-cyclodextrin extract cholesterol from biological membranes, whereas N-acetyl-L-hydroxyproline loosens collagen structures. It was noted that incubation in ScaleS0 can greatly render a fixed sample adaptable to tissue clearing solutions. This adaptation process was achieved in the original ScaleA2 protocol by a freeze/thaw procedure that involved cryoprotection, OCT embedding, and re-fixation. These time-consuming and laborious steps in the original protocol can be replaced by a simple incubation in ScaleS0. Second, the permeable (adapted) sample is incubated sequentially in ScaleS1, ScaleS2, and ScaleS3. These urea-containing and salt-free ScaleS solutions gradually clear the sample. Finally, the sample is restored by simple washing with PBS(–) (deScaling) for ≥ 6 hrs, and then incubated in ScaleS4 for 12 hrs prior to observation. Fresh ScaleS4 is used as the mounting medium.
Although our original protocol using ScaleA2 specified the incubation temperature of 4 °C (Ref. 2), we found that the clearing processes were enhanced at higher temperatures. This temperature effect applies to ScaleS as well. In the new protocol, the incubation temperature is 37 °C for all ScaleS solutions but 4 °C for PBS(–).
The incubation occurs in a temperature-controlled orbital shaker (70–80 rpm/min). Occasionally, micelles may form in the mounting (ScaleS4) solution, and can be removed by decreasing the concentration of DMSO from 25% to 15%.
(c) AbScale protocol. The formulas of the AbScale solution and AbScale rinse solution are as follows.
[AbScale solution: a PBS(–) solution containing 0.33 M urea, 0.1–0.5% Triton X-100]
[AbScale rinse solution: a 0.1× PBS(–) solution containing 2.5% BSA, 0.05% (w/v) Tween-20].
The formulas of all other solutions are shown in Supplementary Table 1. The procedure assumes that an adult mouse cerebral hemisphere or a 1–2-mm-thick brain slice is processed as a sample to be immunolabeled/cleared. The method starts with the sequential incubation of a fixed brain sample in multiple solutions: ScaleS0, ScaleA2, ScaleB4(0), and ScaleA2 for permeabilization/clearing. Then, after deScaling by PBS(–) wash, the sample is incubated with a fluorescently labeled primary Ab (Direct IHC) or a primary Ab and then a fluorescently labeled secondary Ab (Indirect IHC) in an AbScale solution. Before re-fixation with 4% PFA, the sample is rinsed in an AbScale rinse solution. Finally, the immunostained sample is optically cleared by incubation in ScaleS4.
(d) ChemScale protocol. The formulas of all the solutions are shown in Supplementary Table 1. This procedure assumes that a mouse cerebral hemisphere or a 1–2-mm-thick brain slice is processed as a sample to be labeled/cleared. When combined with AbScale, fluorescent chemicals can be put in an AbScale solution containing a fluorescently labeled primary antibody.
(e) ScaleSQ(0) protocol. (f) ScaleSQ(5) protocol. These procedures assume that a 1–2-mm-thick brain slice is processed as a sample to be cleared. The only problem is that ScaleSQ solutions must be kept constantly at temperatures above 30 °C; otherwise, urea would precipitate due to its high concentration. The formulas of ScaleSQ(0) and ScaleSQ(5) solutions are shown in Supplementary Table 1.
Supplementary Figure 2 Characterization of macroscopic structures of brain samples during and after ScaleS-treatment
(a) Three and two fixed whole mouse brains (9 weeks old) were treated with ScaleS (Supplementary Fig. 1b) (red lines) and PBS(–) (black lines), respectively. Sample volume was measured by liquid displacement (Ref. 2) throughout the protocol. ScaleS0 reduced the volume of a fixed whole mouse brain to approximately 55% of the original in approximately 6 hrs. Then, the urea-containing and salt-free ScaleS (ScaleS1, ScaleS2, and ScaleS3) solutions gradually cleared the sample with moderate expansion. After treatment with ScaleS3, the sample expanded to approximately 150% of the original volume. The deScaling before ScaleS4 incubation shrank the sample to approximately 90% of the original. After equilibration in ScaleS4, the volume returned to nearly 100% of the original (102.1 ± 3.5%, mean ± s.d., n = 3).
(b) Comparison of macroscopic structures in 2-mm-thick brain slices before and after ScaleS treatment.
A horizontal slice (left) and a coronal slice (right) both containing the hippocampus were prepared from YFP-H mice (21 weeks old). Slices were imaged using a fluorescence stereomicroscope for transmission (T) and fluorescence from YFP. The transmission images before and after the treatment were traced and colored blue and red, respectively (T trace). The blue image was superimposed on the red one to create a merged image.
In both merged images (horizontal and coronal), blue and red signals coexist almost everywhere, indicating that the ScaleS method is free from a tissue deformity problem. Scale bars: 5 mm.
(c) A whole mouse brain of a YFP-H mouse (13 weeks old) was fixed and cleared by ScaleS. After incubation in ScaleS4 solution for 15 months at 4 °C, a transmission image (T) and a fluorescence image (YFP) were taken using a fluorescence stereomicroscope. Scale bars: 5 mm.
Supplementary Figure 3 Comparative observation of optical transparency and fluorescence preservation between cleared and PBS(–)-treated hemispheres and 2-mm-thick brain slices
(a) ScaleS, CUBIC, 3DISCO, and SeeDB were evaluated compared with PBS(–). Images (except for #1) comparing the clarification (T: transmission) and preservation of YFP fluorescence (FL) between cleared (left) and PBS(–)-treated (right) hemispheres from YFP-H (solid bar) and ChR2-YFP (open bar) mice are shown. The age in week (W) and the gender of each mouse is shown in parentheses. Transmission images were taken using a single-lens reflex camera (NIKON D7000) with a patterned background. Fluorescence images were taken using a fluorescence stereomicroscope (NIKON SMZ18) with a color CCD camera (ZEISS AxioCam 506 color).
(b) ScaleS and PACT were evaluated compared with PBS(–). Images (except for #1) comparing T and FL between cleared (left) and PBS(–)-treated (right) brain slices (2 mm thick) from YFP-H mice are shown. Image acquisition was performed in the same way as described in (a).
Scale bars: 5 mm. See Fig. 2 for the quantitative measurements and the representative images (#1).
Supplementary Figure 4 Clearing young mouse samples by ScaleS and SeeDB
Two mouse embryos (E13.5) and the left and right halves of the brain of a neonatal mouse (P3) were used. After fixation with 4% PFA, they were cleared according to the protocols of ScaleS (Supplementary Fig. 1b) and SeeDB (Ref. 5). At time points indicated, clarification of the samples was examined by taking photos. The immersion solutions at picture acquisition are shown (right). Scale bar: 5 mm.
Supplementary Figure 5 Protein epitope preservation after ScaleS and CUBIC treatment
Immunohistochemistry on sections restored from ScaleS (left) and CUBIC (middle) compared with control sections stored in PBS(–) (right). DG, dentate gyrus; GCL, granule cell layer; MF, mossy fiber; SO, stratum oriens; SR, stratum radiatum. Scale bars: 100 μm. See Online Methods for details of 2D-IHC. Cytoskeleton proteins, such as MAP2 and GFAP, were well immunolocalized in the sections prepared from ScaleS- and CUBIC-treated samples. However, substantial differences in synaptic proteins were noted between the ScaleS- and CUBIC-treated samples. ScaleS treatment preserved the immunostaining of presynaptic and postsynaptic proteins enriched in the mossy fiber (MF) and CA3 regions, respectively. As a result, the signal contrast between the MF and CA3 regions was clearly distinguished. By contrast, CUBIC treatment attenuated the intensity and/or specificity of the immunostaining. In addition, the immunostaining of a cell adhesion molecule (PSA-NCAM) in granule cells in the dentate gyrus and MF regions was preserved in the ScaleS-treated sample but not the CUBIC-treated one. The experiment was performed in triplicate for each using different mouse brains.
Supplementary Figure 6 Transmission electron microscopy of a restored sample after ScaleS and a control sample.
(a–f) ScaleS.
(a) Schematic diagrams of the experimental flow. The whole hemisphere of C57BL/6J mouse (10 weeks old) was fixed with 4% PFA, and cleared completely by ScaleS. The sample was incubated in the mounting solution (ScaleS4) at 37 °C for 14 hrs on the assumption that it was subjected to an overnight imaging experiment. The sample was restored by washing with PBS(–) and a 1-mm-cube of primary somatosensory cortex (S1) was excised for the preparation of ultrathin sections.
(b–f) Transmission electron microscopy (TEM) images. (b) A micrograph containing two somata of pyramidal neurons in cortical layer II/III. (c and d) Excitatory, asymmetric synapses in the two regions (boxes in (b)) of the neuropil. Solid arrowheads indicate thickened postsynaptic densities (PSDs). (e) A magnified micrograph corresponding to the box shown in (b). (f) A magnified micrograph corresponding to the box shown in (e). A symmetric synapse is indicated by open arrowheads. Den, dendrite; AH, axon hillock; AIS, axon initial segment; AT, axon terminal. Scale bars: 3 μm (b); 200 nm (c–f). Similar TEM images were obtained from 3 other mice of the same age.
(g–i) Control.
(g) TEM images were acquired in the same way as described for (a) except that the sample was not cleared. Briefly, the whole hemisphere of a C57BL/6J mouse (10 weeks old) was fixed with 4% PFA. Then a 1-mm-cube in the primary somatosensory cortex (S1) was taken out and used for the preparation of ultrathin sections. (h,i) Two representative TEM images representing excitatory synapses are shown. Solid arrowheads indicate thickened postsynaptic densities (PSDs). Den, dendrite; AT, axon terminal. Scale bars: 100 nm. Similar TEM images were obtained from 3 other mice of the same age.
Supplementary Figure 7 Multicolor AbScale method applied to brain tissues from AppNL-F/NL-F mice and AD patients for 3D reconstruction
a) Optical components used for triple-color AbScale for immunostaining neuronal nuclei and Aβ plaques by a fluorescence stereomicroscope (Fig. 4). The excitation (dotted line) and emission (solid line) spectra of Alexa488, Cy3, and Cy5 are normalized and shown with the same colors as used in the main figures in CMYK mode. The transmission properties of the excitation and emission filters are illustrated by boxes. The filter sets used for Alexa488, Cy3, and Cy5 were ET-GFP, DSR, and ET-Cy5, respectively.
(b) Colocalization of pre-cut staining signals (Cy5-A60) with post-cut staining signals (Cy3-αNeuN) in the same section as shown in Fig. 4. Fluorescence images of Cy3 and Cy5 were acquired using a confocal microscope (FV1000) equipped with the optical components (indicated by the spectral graph), and shown in green and red, respectively, in RGB mode. Their colocalization was quantified by correlation analysis. The Pearson correlation coefficient (r) was calculated to be 0.969. Scale bars: 1 mm.
(c) Alexa488 and PP-BTA-1 for confocal microscopy (see Fig. 5a,b).
(d) Alexa488 and Texas Red for SPIM (see Fig. 5d).
(e) Alexa488 and Alexa546 for SPIM (see Fig. 6a–h).
(f) Alexa488, Alexa546, and Cy5 for SPIM (see Fig. 6i-r).
The excitation (dotted line) and emission (solid line) spectra are normalized and shown with the same colors as used in the main figures. The transmission properties of the emission filters are illustrated by boxes. ex, excitation; em, emission. The used laser lines are indicated by arrows.
Supplementary Figure 8 Volume preservation after AbScale and ChemScale treatment
(a) A pair of fixed hemispheres was prepared from a C57BL/6J mouse (12 weeks old); the left hemisphere was AbScaled with Alexa488-A60 (anti-NeuN mAb) and the right one was incubated in PBS(–). Fluorescence (FL) and transmission (T) images were taken using a fluorescence stereomicroscope. Scale bars: 5 mm. The sample volume was measured by liquid displacement. The volume ratio of the stained sample to untreated sample was plotted on a bar graph. The experiment was duplicated.
(b) A pair of fixed hemispheres was prepared from a C57BL/6J mouse (12 weeks old), and the left hemisphere was ChemScaled with propidium iodide (PI) and the right one was incubated in PBS(–). Fluorescence (FL) and transmission (T) images were taken using a fluorescence stereomicroscope. Scale bars: 5 mm. The sample volume was measured by liquid displacement. The volume ratio of the stained sample to untreated sample was plotted on a bar graph. The experiment was duplicated.
Supplementary Figure 9 Compatibility of ScaleS with DiI labeling
(a) DiI-labeled neural tracts in the hippocampus were well preserved with clarification by ScaleS but not by CUBIC, 3DISCO, or PACT.
top, Bright-field images showing coronal slices (2 mm thick) with DiI spotting on the hippocampus.
bottom, DiI fluorescence (FL) images of left and right halves after clarification. The cleared samples are outlined by white dotted lines; significant shrinkage was noticed after 3DISCO treatment. DiI staining regions are circled by white broken lines. Scale bar: 5 mm.
(b) Dual-color ChemScale to visualize FSB-labeled Aβ plaques and DiI-labeled neurites in a clarified block sample from an AD patient brain (#1625, see Fig. 6).
left, The optical components for dual-color ChemScale. The excitation (dotted line) and emission (solid line) spectra are normalized and shown with the same colors as used in the images. The transmission properties of the emission filters are illustrated by boxes. ex, excitation; em, emission. The used laser lines are indicated by arrows.
right, A 5-mm-cube prepared from the postmortem brain sample was shot by a DiI crystal, incubated in a solution containing FSB and 2% PFA, and then clarified in ScaleS4. The sample was imaged using a confocal microscope (Olympus FV1200), and representative images with and without FSB-labeled plaques are shown. Scale bars: 0.1 mm. Similar results were obtained with another cube from #1625.
Supplementary Figure 10 Immunoelectron microscopy of Aβ plaques in association with neighboring synapses
(a–c) ImmunoEM images of brain regions from a cleared/restored hemisphere of an aged AppNL-F/NL-F mouse (24 months old). Excitatory synapses identified by high-magnification EM observation are encircled in yellow. A, axon; BV, blood vessel. (a) The same field as shown in Fig. 5e. (b) The plaque involved a neuron (delineated in magenta), which was invaded by a microglial cell (indicated by an arrow). Scale bars: 10 μm.
(d) The identified synapses were counted inside and outside plaques, and their densities per 1,000 μm2 extracellular area were calculated and plotted (solid circles, triangles, and squares for (a), (b), and (c), respectively). Likewise, intact excitatory synapses were counted in 7 other regions. Collective data are shown on a bar graph (mean ± s.e.m.) (n = 10). The synapse density was significantly higher outside plaques than inside. Statistical analysis was performed by Welch’s unpaired t-test (two-tailed) (p < 0.001).
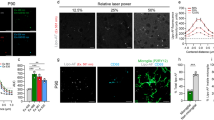
Supplementary Figure 11 Plaque-centered investigation of microglial association in the brain of aged AppNL-F/NL-F mice at different ages
Each histogram shows the distribution of the distances of activated and resting microglia to the plaque edge for individual plaques. Isolated plaques that were not associated with any activated microglia in their vicinity (≤ 21 μm) are labeled in blue. Other plaques showing microglial association in their vicinity (≤ 21 μm) are labeled in red. Vertical dotted lines are drawn at 21 μm to distinguish between closely neighboring and distant microglia.
(a) Thirty-nine plaques in the brain region of a 20-month-old AppNL-F/NL-F mouse (see Fig. 6a–h).
(b) Twenty-seven plaques in the brain region of a 10-month-old AppNL-F/NL-F mouse.
(c) Sixteen plaques in the brain region of a 12-month-old AppNL-F/NL-F mouse.
(d) Thirty-one plaques in the brain region of a 14-month-old AppNL-F/NL-F mouse.
(b–d) Two volume rendered images (xy) are shown on the right. top, a low-magnification image generated from the observed region; bottom, a high-magnification image containing analyzed plaques.
(e) A bar chart showing the percentage of isolated plaques analyzed at age 10, 12, 14, and 20 months.
Scale bars in the perspective VR images (yellow) show the distance on the foreground plane: 50 μm.
Supplementary Figure 12 ScaleSQ(0) and ScaleSQ(5): two quick versions of ScaleS applicable to brain slices
(a–h) ScaleSQ(0).
(a–e) Fluorescence (a and e) and transmission (b–d) images of a 1-mm-thick brain slice prepared from an 8-week-old YFP-H mouse before (a) and after 0- (b), 1- (c), and 2- (d and e) hour incubation in ScaleSQ(0) at 37 °C. Similar observation was performed using 9 other slices.
(f–h) TEM observation of a brain sample restored from ScaleSQ(0). Ultrathin sections were prepared and imaged in the same way as for Supplementary Fig. 6. (f) A micrograph containing a neuronal soma. (g and h) Magnified micrographs corresponding to the boxes in (f), showing an excitatory, asymmetric synapse, and a myelinated axon, respectively. Similar ultrastructure was observed for other 25 excitatory synapses and 9 myelinated axons.
(i–p) ScaleSQ(5).
(i–m) Fluorescence (i and m) and transmission (j–l) images of a 1-mm-thick brain slice prepared from an 8-week-old YFP-H mouse before (i) and after 0- (j), 1- (k), and 2- (l and m) hour incubation in ScaleSQ(5) at 37 °C. Similar observation was performed using 4 other slices.
(n–p) TEM observation of a brain sample restored from ScaleSQ(5). Ultrathin sections were prepared and imaged in the same way as for Supplementary Fig. 6. (n) A micrograph containing a neuronal soma. (o and p) Magnified micrographs corresponding to the boxes in (n), possibly showing an excitatory, asymmetric synapse, and a myelinated axon, respectively. Similar ultrastructure was observed for other 9 excitatory synapses and 4 myelinated axons.
(g,o) Solid arrowheads indicate thickened postsynaptic densities. Den, dendrite; AT, axon terminal.
(h,p) Ax, axon; My, myelin.
(q) Change in size of slices after PBS(–)-, ScaleSQ(0)-, and ScaleSQ(5)-treatment. Four slices were used for each method. Three distances on each slice were measured, and the values after treatment were normalized to the initial values. Mean ± s.e.m. (n = 12).
Scale bars: a–e, i–m, 5 mm; f, n, 4 μm; g, h, o, p, 200 nm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Tables 1–3 (PDF 1638 kb)
Supplementary Methods Checklist
(PDF 450 kb)
3D visualization of the spatial association between plaques and blood vessels.
Animation (zooming in and out) of 3D image data (580 × 580 × 860 mm volume) in the cereberal cortex of a 20-month-old AppNL-F/NL-F mouse. The entire vasculature was labelled with Texas Red-labeled lectin. Aβ plaques in the left hemisphere were immunolabelled with AbScale using Alexa488-6E10. Red, blood vessels; Green, Aβ plaques. Images were acquired using SPIM. Backward and forward perspective images were created at different depths and from different angles. See Fig. 4d. (MPG 5010 kb)
Rapid clarification of brain slices by ScaleSQ.
Time-lapse imaging of coronal slices (1 mm thick) prepared from an 8-week-old YFP-H mice during the incubation in ScaleSQ(0) and ScaleSQ(5) at 37 °C. Transmitted-light bright-field images were acquired with a digital single lens reflex camera (Nikon). See Fig. 7. (MPG 1466 kb)
Source data
Rights and permissions
About this article
Cite this article
Hama, H., Hioki, H., Namiki, K. et al. ScaleS: an optical clearing palette for biological imaging. Nat Neurosci 18, 1518–1529 (2015). https://doi.org/10.1038/nn.4107
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4107
This article is cited by
-
Biomechanical Remodeling of Aortic Valve Interstitial Cells During Calcified Lesion Formation In Vitro
Annals of Biomedical Engineering (2024)
-
TSA-PACT: a method for tissue clearing and immunofluorescence staining on zebrafish brain with improved sensitivity, specificity and stability
Cell & Bioscience (2023)
-
Bat teeth illuminate the diversification of mammalian tooth classes
Nature Communications (2023)
-
Reconstituting human somitogenesis in vitro
Nature (2023)
-
Multimodal optical clearing to minimize light attenuation in biological tissues
Scientific Reports (2023)