Abstract
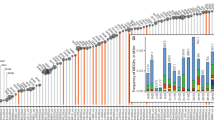
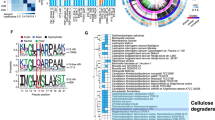
Cellulosomes are large, multiprotein complexes that tether plant biomass-degrading enzymes together for improved hydrolysis1. These complexes were first described in anaerobic bacteria, where species-specific dockerin domains mediate the assembly of enzymes onto cohesin motifs interspersed within protein scaffolds1. The versatile protein assembly mechanism conferred by the bacterial cohesin–dockerin interaction is now a standard design principle for synthetic biology2,3. For decades, analogous structures have been reported in anaerobic fungi, which are known to assemble by sequence-divergent non-catalytic dockerin domains (NCDDs)4. However, the components, modular assembly mechanism and functional role of fungal cellulosomes remain unknown5,6. Here, we describe a comprehensive set of proteins critical to fungal cellulosome assembly, including conserved scaffolding proteins unique to the Neocallimastigomycota. High-quality genomes of the anaerobic fungi Anaeromyces robustus, Neocallimastix californiae and Piromyces finnis were assembled with long-read, single-molecule technology. Genomic analysis coupled with proteomic validation revealed an average of 312 NCDD-containing proteins per fungal strain, which were overwhelmingly carbohydrate active enzymes (CAZymes), with 95 large fungal scaffoldins identified across four genera that bind to NCDDs. Fungal dockerin and scaffoldin domains have no similarity to their bacterial counterparts, yet several catalytic domains originated via horizontal gene transfer with gut bacteria. However, the biocatalytic activity of anaerobic fungal cellulosomes is expanded by the inclusion of GH3, GH6 and GH45 enzymes. These findings suggest that the fungal cellulosome is an evolutionarily chimaeric structure—an independently evolved fungal complex that co-opted useful activities from bacterial neighbours within the gut microbiome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fontes, C. M. G. A. & Gilbert, H. J. Cellulosomes: highly efficient nanomachines designed to deconstruct plant cell wall complex carbohydrates. Annu. Rev. Biochem. 79, 655–681 (2010).
You, C., Myung, S. & Zhang, Y.-H. P. Facilitated substrate channeling in a self-assembled trifunctional enzyme complex. Angew. Chem. Int. Ed. 51, 8787–8790 (2012).
Liu, F., Banta, S. & Chen, W. Functional assembly of a multi-enzyme methanol oxidation cascade on a surface-displayed trifunctional scaffold for enhanced NADH production. Chem. Commun. 49, 3766–3768 (2013).
Fanutti, C. C., Ponyi, T. T., Black, G. W. G., Hazlewood, G. P. G. & Gilbert, H. J. H. The conserved noncatalytic 40-residue sequence in cellulases and hemicellulases from anaerobic fungi functions as a protein docking domain. J. Biol. Chem. 270, 29314–29322 (1995).
Nagy, T. T. et al. Characterization of a double dockerin from the cellulosome of the anaerobic fungus Piromyces equi. J. Mol. Biol. 373, 612–622 (2007).
Haitjema, C. H., Solomon, K. V., Henske, J. K., Theodorou, M. K. & O'Malley, M. A. Anaerobic gut fungi: advances in isolation, culture, and cellulolytic enzyme discovery for biofuel production. Biotechnol. Bioeng. 111, 1471–1482 (2014).
Solomon, K. V. et al. Early-branching gut fungi possess a large, comprehensive array of biomass-degrading enzymes. Science 351, 1192–1195 (2016).
Anderson, T. D. et al. Assembly of minicellulosomes on the surface of Bacillus subtilis. Appl. Environ. Microb. 77, 4849–4858 (2011).
Tsai, S.-L., Oh, J., Singh, S., Chen, R. & Chen, W. Functional assembly of minicellulosomes on the Saccharomyces cerevisiae cell surface for cellulose hydrolysis and ethanol production. Appl. Environ. Microb. 75, 6087–6093 (2009).
Tsai, S.-L., Dasilva, N. A. & Chen, W. Functional display of complex cellulosomes on the yeast surface via adaptive assembly. ACS Synth. Biol. 2, 14–21 (2013).
Wilson, C. A. & Wood, T. M. The anaerobic fungus Neocallimastix frontalis—isolation and properties of a cellulosome-type enzyme fraction with the capacity to solubilize hydrogen-bond-ordered cellulose. Appl. Microbiol. Biotechnol. 37, 125–129 (1992).
Nguyen, K. B. et al. Phosphorylation of spore coat proteins by a family of atypical protein kinases. Proc. Natl Acad. Sci. USA 113, E3482–E3491 (2016).
Steenbakkers, P. J. M. et al. beta-Glucosidase in cellulosome of the anaerobic fungus Piromyces sp. strain E2 is a family 3 glycoside hydrolase. Biochem. J. 370, 963–970 (2003).
Youssef, N. H. et al. The genome of the anaerobic fungus Orpinomyces sp. strain C1A reveals the unique evolutionary history of a remarkable plant biomass degrader. Appl. Environ. Microb. 79, 4620–4634 (2013).
Shah, D. S., Joucla, G., Remaud-Simeon, M. & Russell, R. R. B. Conserved repeat motifs and glucan binding by glucansucrases of oral streptococci and Leuconostoc mesenteroides. J. Bacteriol. 186, 8301–8308 (2004).
Raghothama, S. et al. Characterization of a cellulosome dockerin domain from the anaerobic fungus Piromyces equi. Nat. Struct. Biol. 8, 775–778 (2001).
Bayer, E. A., Belaich, J.-P., Shoham, Y. & Lamed, R. The cellulosomes: multienzyme machines for degradation of plant cell wall polysaccharides. Annu. Rev. Microbiol. 58, 521–554 (2004).
Garcia-Vallve, S., Romeu, A. & Palau, J. Horizontal gene transfer of glycosyl hydrolases of the rumen fungi. Mol. Biol. Evol. 17, 352–361 (2000).
Grigoriev, I. V. et al. Mycocosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42, D699–D704 (2014).
Markowitz, V. M. et al. IMG/m 4 version of the integrated metagenome comparative analysis system. Nucleic Acids Res. 42, D568–D573 (2014).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Steenbakkers, P. J. P. et al. Noncatalytic docking domains of cellulosomes of anaerobic fungi. J. Bacteriol. 183, 5325–5333 (2001).
Gilmore, S. P., Henske, J. K. & O'Malley, M. A. Driving biomass breakdown through engineered cellulosomes. Bioengineered 6, 204–208 (2015).
Martin, J. et al. Rnnotator: an automated de novo transcriptome assembly pipeline from stranded RNA-Seq reads. BMC Genomics 11, 663 (2010).
Solomon, K. V., Henske, J. K., Theodorou, M. K. & O'Malley, M. A. Robust and effective methodologies for cryopreservation and DNA extraction from anaerobic gut fungi. Anaerobe 38, 39–46 (2016).
Lam, K. K., LaButti, K., Khalak, A. & Tse, D. FinisherSC: a repeat-aware tool for upgrading de novo assembly using long reads. Bioinformatics 31, 3207–3209 (2015).
Chin, C. S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005); corrigendum 441, 120 (2006).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Trong, S. et al. Gap resolution: a software package for improving newbler genome assemblies. in Proceedings of the 4th Annual Meeting on Sequencing Finishing, Analysis in the Future 35 (2009).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Eddy, S. R. Multiple alignment using hidden Markov models. Proc. Int. Conf. Intell. Syst. Mol. Biol. 3, 114–120 (1995).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. Trimal: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Ali, B. R. S. et al. Cellulases and hemicellulases of the anaerobic fungus Piromyces constitute a multiprotein cellulose-binding complex and are encoded by multigene families. FEMS Microbiol. Lett. 125, 15–21 (1995).
Xiao, Z., Storms, R. & Tsang, A. Microplate-based carboxymethylcellulose assay for endoglucanase activity. Anal. Biochem. 342, 176–178 (2005).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
The authors acknowledge funding support from the Office of Science (BER), US Department of Energy (DE-SC0010352), the US Department of Agriculture (award no. 2011-67017-20459), the National Science Foundation (DGE 1144085) and the Institute for Collaborative Biotechnologies through grant no. W911NF-09-0001 from the US Army Research Office. A portion of this research was performed under the Facilities Integrating Collaborations for User Science (FICUS) exploratory effort and used resources at the DOE Joint Genome Institute and the Environmental Molecular Sciences Laboratory, which are DOE Office of Science User Facilities. Both facilities are sponsored by the Office of Biological and Environmental Research and operated under contract nos. DE-AC02-05CH11231 (JGI) and DE-AC05-76RL01830 (EMSL). The authors acknowledge support from the California NanoSystems Institute (CNSI), supported by the University of California, Santa Barbara, and the University of California, Office of the President. SPR data were generated in the UCSB and UCOP-supported Biological Nanostructures Laboratory within the California NanoSystems Institute. The authors thank P.J. Weimer (US Dairy Forage Research Center) for lignocellulosic substrates. B.H. acknowledges IDEX Aix-Marseille (Grant Microbio-E) and Agence Nationale de la Recherche (grant no. ANR-14-CE06-0020) for funding.
Author information
Authors and Affiliations
Contributions
C.H.H., S.P.G. and M.A.O. planned the experiments. C.H.H. and R.D. performed ELISA and S.P.G. performed SPR experiments. C.H.H., S.P.G., A.K. and M.A.O. wrote the manuscript. H.M.B., S.O.P. and A.T.W. performed proteomic analyses. K.V.S. and J.K.H. prepared and analysed genomic samples for N. californiae, P. finnis and A. robustus. B.B., T.v.A. and J.H.P.H. prepared and analysed genomic samples for Piromyces sp. E2. Z.Z. and J.C. sequenced, K.L. assembled, and A.K., S.J.M. and A.A.S. annotated and analysed genomes. B.H. and M.H. analysed and classified carbohydrate-active enzymes. M.A.O., S.E.B., K.B. and I.V.G. coordinated genome projects at JGI.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Description of Supplementary Datasets, Supplementary Tables 1–7 and Supplementary Figures 1–9 (PDF 2587 kb)
Supplementary Datasets
The supplementary datasets contain 3 types of files related to each of the 10 Pfam domains for which there is evidence of DDP HGT between Neocallimastigomycota and Bacteria. (ZIP 333 kb)
Rights and permissions
About this article
Cite this article
Haitjema, C., Gilmore, S., Henske, J. et al. A parts list for fungal cellulosomes revealed by comparative genomics. Nat Microbiol 2, 17087 (2017). https://doi.org/10.1038/nmicrobiol.2017.87
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.87
This article is cited by
-
A family of di-glutamate mucin-degrading enzymes that bridges glycan hydrolases and peptidases
Nature Catalysis (2024)
-
Anaerobic fungi in the tortoise alimentary tract illuminate early stages of host-fungal symbiosis and Neocallimastigomycota evolution
Nature Communications (2024)
-
Understanding the microbial fibre degrading communities & processes in the equine gut
Animal Microbiome (2023)
-
Genomes of fungi and relatives reveal delayed loss of ancestral gene families and evolution of key fungal traits
Nature Ecology & Evolution (2023)
-
Structure and enzymatic characterization of CelD endoglucanase from the anaerobic fungus Piromyces finnis
Applied Microbiology and Biotechnology (2023)