Abstract
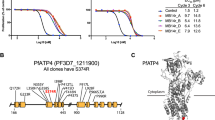
Malaria control is heavily dependent on chemotherapeutic agents for disease prevention and drug treatment. Defining the mechanism of action for licensed drugs, for which no target is characterized, is critical to the development of their second-generation derivatives to improve drug potency towards inhibition of their molecular targets. Mefloquine is a widely used antimalarial without a known mode of action. Here, we demonstrate that mefloquine is a protein synthesis inhibitor. We solved a 3.2 Å cryo-electron microscopy structure of the Plasmodium falciparum 80S ribosome with the (+)-mefloquine enantiomer bound to the ribosome GTPase-associated centre. Mutagenesis of mefloquine-binding residues generates parasites with increased resistance, confirming the parasite-killing mechanism. Furthermore, structure-guided derivatives with an altered piperidine group, predicted to improve binding, show enhanced parasiticidal effect. These data reveal one possible mode of action for mefloquine and demonstrate the vast potential of cryo-electron microscopy to guide the development of mefloquine derivatives to inhibit parasite protein synthesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
World Malaria Report (WHO, 2016); http://www.who.int/malaria/publications/world-malaria-report-2016/en
White, N. J. et al. Malaria. Lancet 383, 723–735 (2014).
Wells, T. N., Hooft van Huijsduijnen, R. & Van Voorhis, W. C. Malaria medicines: a glass half full? Nat. Rev. Drug Discov. 14, 424–442 (2015).
Nevin, R. L. & Byrd, A. M. Neuropsychiatric adverse reactions to mefloquine: a systematic comparison of prescribing and patient safety guidance in the US, UK, Ireland, Australia, New Zealand, and Canada. Neurol. Ther. 5, 69–83 (2016).
Lim, P. et al. Decreasing pfmdr1 copy number suggests that Plasmodium falciparum in western Cambodia is regaining in vitro susceptibility to mefloquine. Antimicrob. Agents Chemother. 59, 2934–2937 (2015).
Roberts, L. Malaria wars. Sci. Transl. Med. 352, 398–405 (2016).
Sanchez, C. P., Rotmann, A., Stein, W. D. & Lanzer, M. Polymorphisms within PfMDR1 alter the substrate specificity for anti-malarial drugs in Plasmodium falciparum. Mol. Microbiol. 70, 786–798 (2008).
Cowman, A. F., Galatis, D. & Thompson, J. K. Selection for mefloquine resistance in Plasmodium falciparum is linked to amplification of the pfmdr1 gene and cross-resistance to halofantrine and quinine. Proc. Natl Acad. Sci. USA 91, 1143–1147 (1994).
Reed, M. B., Saliba, K. J., Caruana, S. R., Kirk, K . & Cowman, A. F. Pgh1 modulates sensitivity and resistance to multiple antimalarials in Plasmodium falciparum. Nature 403, 906–909 (2000).
Sanchez, C. P., Dave, A., Stein, W. D. & Lanzer, M. Transporters as mediators of drug resistance in Plasmodium falciparum. Int. J. Parasitol. 40, 1109–1118 (2010).
Gamo, F.-J. et al. Thousands of chemical starting points for antimalarial lead identification. Nature 465, 305–310 (2010).
Dahl, E. L. et al. Tetracyclines specifically target the apicoplast of the malaria parasite Plasmodium falciparum. Antimicrob. Agents Chemother. 50, 3124–3131 (2006).
Goodman, C. D., Su, V. & McFadden, G. I. The effects of anti-bacterials on the malaria parasite Plasmodium falciparum. Mol. Biochem. Parasitol. 152, 181–191 (2007).
Wong, W. et al. Cryo-EM structure of the Plasmodium falciparum 80S ribosome bound to the anti-protozoan drug emetine. eLife 3, e03080 (2014).
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011).
Spahn, C. M. et al. Domain movements of elongation factor eEF2 and the eukaryotic 80S ribosome facilitate tRNA translocation. EMBO J. 23, 1008–1019 (2004).
Brodersen, D. E. et al. The structural basis for the action of the antibiotics tetracycline, pactamycin, and hygromycin B on the 30S ribosomal subunit. Cell 103, 1143–1154 (2000).
Lutz, R. E., Ohnmacht, C. J. & Patel, A. R. Antimalarials. 7. Bis(trifluoromethyl)-α-(2-piperidyl)-4-quinolinemethanols. J. Med. Chem. 14, 926–928 (1971).
Harms, J. M. et al. Translational regulation via L11: molecular switches on the ribosome turned on and off by thiostrepton and micrococcin. Mol. Cell 30, 26–38 (2008).
Ghorbal, M. et al. Genome editing in the human malaria parasite Plasmodium falciparum using the CRISPR–Cas9 system. Nat. Biotechnol. 32, 819–821 (2014).
Lelievre, J. et al. Activity of clinically relevant antimalarial drugs on Plasmodium falciparum mature gametocytes in an ATP bioluminescence ‘transmission blocking’ assay. PLoS ONE 7, e35019 (2012).
Delves, M. J. et al. Male and female Plasmodium falciparum mature gametocytes show different responses to antimalarial drugs. Antimicrob. Agents Chemother. 57, 3268–3274 (2013).
Waters, A. P., Syin, C. & McCutchan, T. F. Developmental regulation of stage-specific ribosome populations in Plasmodium. Nature 342, 438–440 (1989).
Aguiar, A. C., Pereira, D. B., Amaral, N. S., De Marco, L. & Krettli, A. U. Plasmodium vivax and Plasmodium falciparum ex vivo susceptibility to anti-malarials and gene characterization in Rondonia, west Amazon, Brazil. Malar. J. 13, 73 (2014).
Otigbuo, I. N. & Onabanjo, A. O. The in vitro and in vivo effects of mefloquine on Trypanosoma brucei brucei. J. Hyg. Epidemiol. Microbiol. Immunol. 36, 191–199 (1992).
Holfels, E., McAuley, J., Mack, D., Milhous, W. K. & McLeod, R. In vitro effects of artemisinin ether, cycloguanil hydrochloride (alone and in combination with sulfadiazine), quinine sulfate, mefloquine, primaquine phosphate, trifluoperazine hydrochloride, and verapamil on toxoplasma gondii. Antimicrob. Agents Chemother. 38, 1392–1396 (1994).
Khatter, H., Myasnikov, A. G., Natchiar, S. K. & Klaholz, B. P. Structure of the human 80S ribosome. Nature 520, 640–645 (2015).
Brown, A. et al. Structure of the large ribosomal subunit from human mitochondria. Science 346, 718–722 (2014).
Milner, E. et al. Structure–activity relationships amongst 4-position quinoline methanol antimalarials that inhibit the growth of drug sensitive and resistant strains of Plasmodium falciparum. Bioorg. Med. Chem. Lett. 20, 1347–1351 (2010).
Milner, E. et al. Anti-malarial activity of a non-piperidine library of next-generation quinoline methanols. Malar. J. 9, 51 (2010).
Na-Bangchang, K., Muhamad, P., Ruaengweerayut, R., Chaijaroenkul, W. & Karbwang, J. Identification of resistance of Plasmodium falciparum to artesunate–mefloquine combination in an area along the Thai–Myanmar border: integration of clinico-parasitological response, systemic drug exposure, and in vitro parasite sensitivity. Malar. J. 12, 263 (2013).
Garreau de Loubresse, N. et al. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014).
Sun, M . et al. Dynamical features of the Plasmodium falciparum ribosome during translation. Nucleic Acids Res. 43, 10515–10524 (2015).
Ahyong, V. et al. Identification of Plasmodium falciparum specific translation inhibitors from the MMV malaria Box using a high throughput in vitro translation screen. Malar. J. 15, 173 (2016).
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Bai, X. C., Fernandez, I. S., McMullan, G. & Scheres, S. H. Ribosome structures to near-atomic resolution from thirty thousand cryo-EM particles. eLife 2, e00461 (2013).
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Nicholls, R. A., Long, F. & Murshudov, G. N. Low-resolution refinement tools in REFMAC5. Acta Crystallogr. D 68, 404–417 (2012).
Chou, F. C., Sripakdeevong, P., Dibrov, S. M., Hermann, T. & Das, R. Correcting pervasive errors in RNA crystallography through enumerative structure prediction. Nat. Methods 10, 74–76 (2013).
Chen, V. B. et al. Molprobity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Baum, J. et al. Reticulocyte-binding protein homologue 5—an essential adhesin involved in invasion of human erythrocytes by Plasmodium falciparum. Int. J. Parasitol. 39, 371–380 (2009).
Boyle, M. J. et al. Isolation of viable Plasmodium falciparum merozoites to define erythrocyte invasion events and advance vaccine and drug development. Proc. Natl Acad. Sci. USA 107, 14378–14383 (2010).
Gillespie, R. J. et al. 4-quinolinemethanol derivatives as purine receptor antagonists (II) US patent 6,608,085 (2003).
Acknowledgements
The authors thank I. Lucet, J. Boddey, S. Herrmann, G. McFadden, J. Rayner, A. Ruecker, M. Delves, H. Baumann, G. Murshudov and P. Emsley for discussions and experimental assistance, S. Chen and C. Savva for help with microscopy, and J. Grimmett and T. Darling for help with computing. The experimental data were made possible by Victorian State Government Operational Infrastructure Support and Australian Government NHMRC IRIISS. The research was directly supported by a National Health and Medical Research Council of Australia (NHMRC) Project Grant (APP1024678 to J.B. and W.W.), the Australian Cancer Research Foundation, a Human Frontier Science Program (HFSP) Young Investigator Program Grant (RGY0071/2011, to J.B.) and grants from the UK Medical Research Council (MC_UPA0251013, to S.H.W.S.). W.W. is an Early Career Development Awardee (APP1053801) from the NHMRC and was in receipt of a travel award from OzEMalaR to visit MRC–LMB UK to conduct experiments. X.-C.B. is supported by an EU FP7 Marie Curie Postdoctoral Fellowship. A.B. and I.S.F. are supported by grants to V. Ramakrishnan from the Wellcome Trust (WT096570) and the UK Medical Research council (MC_U105184332). J.B. was supported by a Future Fellowship (FT100100112) from the Australian Research Council (ARC) and is currently supported by an Investigator Award from the Wellcome Trust (100993/Z/13/Z). Additional support for this work came from a Pathfinder Award from the Wellcome Trust (105686).
Author information
Authors and Affiliations
Contributions
W.W., X.-C.B., B.E.S., K.E.J., T.T., D.S.M., S.A.R., S.H.W.S. and J.B. designed all experiments. W.W., X.-C.B., B.E.S., K.E.J., A.B., T.T., D.S.M., J.K.T., E.H. and I.S.F. performed experiments. W.W., X.-C.B., B.E.S., K.E.J., T.T., A.B., J.K.T., S.A.R., A.F.C., S.H.W.S. and J.B. contributed to manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-6, Supplementary Data Tables 1-4. (PDF 10094 kb)
Rights and permissions
About this article
Cite this article
Wong, W., Bai, XC., Sleebs, B. et al. Mefloquine targets the Plasmodium falciparum 80S ribosome to inhibit protein synthesis. Nat Microbiol 2, 17031 (2017). https://doi.org/10.1038/nmicrobiol.2017.31
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.31
This article is cited by
-
Deaggregation of mutant Plasmodium yoelii de-ubiquitinase UBP1 alters MDR1 localization to confer multidrug resistance
Nature Communications (2024)
-
Malaria therapeutics: are we close enough?
Parasites & Vectors (2023)
-
Nonclassical mechanisms to irreversibly suppress β-hematin crystal growth
Communications Biology (2023)
-
A Legionella toxin exhibits tRNA mimicry and glycosyl transferase activity to target the translation machinery and trigger a ribotoxic stress response
Nature Cell Biology (2023)
-
CRISPR-based oligo recombineering prioritizes apicomplexan cysteines for drug discovery
Nature Microbiology (2022)