Abstract
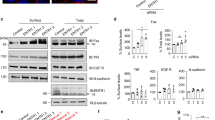
We generated a comprehensive picture of protease substrates in anti-Fas–treated apoptotic human Jurkat T lymphocytes. We used combined fractional diagonal chromatography (COFRADIC) sorting of protein amino-terminal peptides coupled to oxygen-16 or oxygen-18 differential labeling. We identified protease substrates and located the exact cleavage sites within processed proteins. Our analysis yielded 1,834 protein identifications and located 93 cleavage sites in 71 proteins. Indirect evidence of apoptosis-specific cleavage within 21 additional proteins increased the total number of processed proteins to 92. Most cleavages were at caspase consensus sites; however, other cleavage specificities suggest activation of other proteases. We validated several new processing events by immunodetection and by an in vitro assay using recombinant caspases and synthetic peptides containing presumed cleavage sites. The spliceosome complex appeared a preferred target, as 14 of its members were processed. Differential isotopic labeling further revealed specific release of nucleosomal components from apoptotic nuclei.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Dewachter, I. & Van Leuven, F. Secretases as targets for the treatment of Alzheimer's disease: the prospects. Lancet Neurol. 1, 409–416 (2002).
Berdowska, I. Cysteine proteases as disease markers. Clin. Chim. Acta 342, 41–69 (2004).
Docherty, A.J., Crabbe, T., O'Connell, J.P. & Groom, C.R. Proteases as drug targets. Biochem. Soc. Symp. 70, 147–161 (2003).
Lamkanfi, M., Declercq, W., Kalai, M., Saelens, X. & Vandenabeele, P. Alice in caspase land. A phylogenetic analysis of caspases from worm to man. Cell Death Differ. 9, 358–361 (2002).
Fischer, U., Janicke, R.U. & Schulze-Osthoff, K. Many cuts to ruin: a comprehensive update of caspase substrates. Cell Death Differ. 10, 76–100 (2003).
Cryns, V.L. et al. Specific proteolysis of the kinase protein kinase C-related kinase 2 by caspase-3 during apoptosis. Identification by a novel, small pool expression cloning strategy. J. Biol. Chem. 272, 29449–29453 (1997).
Kamada, S. et al. A cloning method for caspase substrates that uses the yeast two-hybrid system: cloning of the antiapoptotic gene gelsolin. Proc. Natl. Acad. Sci. USA 95, 8532–8537 (1998).
Kuida, K. et al. Altered cytokine export and apoptosis in mice deficient in interleukin-1β converting enzyme. Science 267, 2000–2003 (1995).
Li, P. et al. Mice deficient in IL-1β-converting enzyme are defective in production of mature IL-1β and resistant to endotoxic shock. Cell 80, 401–411 (1995).
Gerner, C. et al. The Fas-induced apoptosis analyzed by high throughput proteome analysis. J. Biol. Chem. 275, 39018–39026 (2000).
Thiede, B., Dimmler, C., Siejak, F. & Rudel, T. Predominant identification of RNA-binding proteins in Fas-induced apoptosis by proteome analysis. J. Biol. Chem. 276, 26044–26050 (2001).
Gerner, C. et al. Proteome analysis of nuclear matrix proteins during apoptotic chromatin condensation. Cell Death Differ. 9, 671–681 (2002).
Thiede, B., Siejak, F., Dimmler, C. & Rudel, T. Prediction of translocation and cleavage of heterogeneous ribonuclear proteins and Rho guanine nucleotide dissociation inhibitor 2 during apoptosis by subcellular proteome analysis. Proteomics 2, 996–1006 (2002).
Wajant, H. The Fas signalling pathway: more than a paradigm. Science 296, 1635–1636 (2002).
Gevaert, K. et al. Exploring proteomes and analyzing protein processing by mass spectrometric identification of sorted N-terminal peptides. Nat. Biotechnol. 21, 566–569 (2003).
Staes, A. et al. Global differential non-gel proteomics by quantitative and stable labelling of tryptic peptides with oxygen-18. J. Proteome. Res. 3, 786–791 (2004).
Thornberry, N.A. et al. A combinatorial approach defines specificities of members of the caspase family and granzyme B. Functional relationships established for key mediators of apoptosis. J. Biol. Chem. 272, 17907–17911 (1997).
Garcia-Calvo, M. et al. Inhibition of human caspases by peptide-based and macromolecular inhibitors. J. Biol. Chem. 273, 32608–32613 (1998).
Nicholson, D.W. et al. Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376, 37–43 (1995).
Krippner-Heidenreich, A. et al. Targeting of the transcription factor Max during apoptosis: phosphorylation-regulated cleavage by caspase-5 at an unusual glutamic acid residue in position P1. Biochem. J. 358, 705–715 (2001).
Wu, D. et al. Apoptotic release of histones from nucleosomes. J. Biol. Chem. 277, 12001–12008 (2002).
Galande, S., Dickinson, L.A., Mian, I.S., Sikorska, M. & Kohwi-Shigematsu, T. SATB1 cleavage by caspase 6 disrupts PDZ domain-mediated dimerization, causing detachment from chromatin early in T-cell apoptosis. Mol. Cell. Biol. 21, 5591–5604 (2001).
Sahara, S. et al. Acinus is a caspase-3–activated protein required for apoptotic chromatin condensation. Nature 401, 168–173 (1999).
Thiede, B., Treumann, A., Kretschmer, A., Sohlke, J. & Rudel, T. Shotgun proteome analysis of protein cleavage in apoptotic cells. Proteomics 5, 2123–2130 (2005).
Buratti, E. & Baralle, F.E. Characterization and functional implications of the RNA binding properties of nuclear factor TDP-43, a novel splicing regulator of CFTR exon 9. J. Biol. Chem. 276, 36337–36343 (2001).
Lallena, M.J., Chalmers, K.J., Llamazares, S., Lamond, A.I. & Valcarcel, J. Splicing regulation at the second catalytic step by Sex-lethal involves 3′ splice site recognition by SPF45. Cell 109, 285–296 (2002).
Zhou, Z., Licklider, L.J., Gygi, S.P. & Reed, R. Comprehensive proteomic analysis of the human spliceosome. Nature 419, 182–185 (2002).
Schwerk, C. et al. ASAP, a novel protein complex involved in RNA processing and apoptosis. Mol. Cell. Biol. 23, 2981–2990 (2003).
Schultz, J., Milpetz, F., Bork, P. & Ponting, C.P. SMART, a simple modular architecture research tool: identification of signalling domains. Proc. Natl. Acad. Sci. USA 95, 5857–5864 (1998).
Jiang, Z.H. & Wu, J.Y. Alternative splicing and programmed cell death. Proc. Soc. Exp. Biol. Med. 220, 64–72 (1999).
Acknowledgements
The authors thank P. Vandenabeele for his critical reading of the manuscript, for the suggestions made and for providing the recombinant caspases. K.G. is a postdoctoral fellow and L.M. is a research assistant of the Fund for Scientific Research-Flanders (Belgium; F.W.O.-Vlaanderen). The project was supported by research grants from the Fund for Scientific Research-Flanders (Belgium; project number G.0008.03), the Inter University Attraction Poles (IUAP, project number P5/05), the GBOU research initiative (project number 20204) of the Flanders Institute of Science and Technology (IWT) and the European Union Interaction Proteome (6th Framework Program).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Histogram showing the distribution of ratios of isotopic peptide couples measured following COFRADIC isolation. (PDF 108 kb)
Supplementary Fig. 2
Analysis of the apoptotic-specific cleavage sites within known or predicted domains and motifs of the identified processed Jurkat proteins. (PDF 1648 kb)
Supplementary Table 1
N-termini of mitochondrial proteins. (PDF 89 kb)
Supplementary Table 2
Internally located α-N-acetylated peptides. (PDF 171 kb)
Supplementary Table 3
Identification features of peptides linked to proteins processed in apoptotic Jurkat cells. (PDF 149 kb)
Supplementary Table 4
Peptides only present in lysates of living cells. (PDF 128 kb)
Supplementary Table 5
Summary of proteins that are specifically cleaved in Fas-stimulated Jurkat T-lymphocytes. (PDF 113 kb)
Supplementary Table 6
Peptides exclusively found in the proteome digest of apoptotic cells. (PDF 102 kb)
Rights and permissions
About this article
Cite this article
Van Damme, P., Martens, L., Van Damme, J. et al. Caspase-specific and nonspecific in vivo protein processing during Fas-induced apoptosis. Nat Methods 2, 771–777 (2005). https://doi.org/10.1038/nmeth792
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth792
This article is cited by
-
Proteome integral solubility alteration high-throughput proteomics assay identifies Collectin-12 as a non-apoptotic microglial caspase-3 substrate
Cell Death & Disease (2023)
-
In situ self-assembly for cancer therapy and imaging
Nature Reviews Materials (2023)
-
Spatially Resolved Tagging of Proteolytic Neo-N termini with Subtiligase-TM
The Journal of Membrane Biology (2021)
-
SheddomeDB: the ectodomain shedding database for membrane-bound shed markers
BMC Bioinformatics (2017)
-
Highly sensitive and adaptable fluorescence-quenched pair discloses the substrate specificity profiles in diverse protease families
Scientific Reports (2017)