Abstract
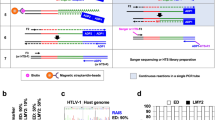
Integrating vector systems used in clinical gene therapy have proven their therapeutic potential in the long-term correction of immunodeficiencies1,2,3,4. The integration loci of such vectors in the cellular genome represent a molecular marker unique for each transduced cell and its clonal progeny. To gain insight into the physiology of gene-modified hematopoietic repopulation and vector-related influences on clonal contributions, we have previously introduced a technology—linear amplification–mediated (LAM) PCR—for detecting and sequencing unknown DNA flanking sequences down to the single cell level5 (Supplementary Note online). LAM-PCR analyses have enabled qualitative and quantitative measurements of the clonal kinetics of hematopoietic regeneration in gene transfer studies, and uncovered the clonal derivation of non-leukemogenic and leukemogenic insertional side effects in preclinical and clinical gene therapy studies4,6,7,8. The reliability and robustness of this method results from the initial preamplification of the vector-genome junctions preceding nontarget DNA removal via magnetic selection. Subsequent steps are carried out on a semisolid streptavidin phase, including synthesis of double complementary strands, restriction digest, ligation of a linker cassette onto the genomic end of the fragment and exponential PCR(s) with vector- and linker cassette–specific primers. LAM-PCR can be adjusted to all unknown DNA sequences adjacent to a known DNA sequence. Here we describe the use of LAM-PCR analyses to identify 5′ long terminal repeat (LTR) retroviral vector adjacent genomic sequences (Fig. 1 and Box 1).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Cavazzana-Calvo, M. et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 288, 669–672 (2000).
Aiuti, A. et al. Correction of ADA-SCID by stem cell gene therapy combined with nonmyeloablative conditioning. Science 296, 2410–2413 (2002).
Gaspar, H.B. et al. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet 364, 2181–2187 (2004).
Ott, M.G. et al. Correction of X-linked chronic granulomatous disease by gene therapy is augmented by insertional activation of MDS/EVI1, PRDM16 or SETBP1. Nat. Med. 12, 401–409 (2006).
Schmidt, M. et al. Polyclonal long-term repopulating stem cell clones in a primate model. Blood 100, 2737–2743 (2002).
Hacein-Bey-Abina, S. et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 302, 415–419 (2003).
Deichmann, A. et al. Vector integration is non-random, clustered and influences the in vivo fate of lymphopoiesis in SCID-X1 gene therapy. J. Clin. Invest. 117, 2225–2232 (2007).
Schwarzwaelder, K. et al. Gammaretrovirus-mediated correction of SCID-X1 is associated with skewed vector integration site distribution in vivo. J. Clin. Invest. 117, 2241–2249 (2007).
Mueller, P.R. & Wold, B. In vivo footprinting of a muscle specific enhancer by ligation mediated PCR. Science 246, 780–786 (1989).
Glimm, H. et al. Efficient marking of human cells with rapid but transient repopulating activity in autografted recipients. Blood 106, 893–898 (2005).
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft, the German Ministry of Education and Research, the European Union and the US National Institutes of Health. We thank all present and former members and collaborators of our laboratory who have contributed work or materials to the many experiments, time and efforts toward establishing and optimizing this protocol.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
LAM-PCR is filed as a US patent 6514706 that is licensed to Cinciannati Childrens Hospital Medical Center (CCHMC), USA.
Supplementary information
Supplementary Table, Methods and Note
Supplementary Tables 1–2, Supplementary Methods, Supplementary Note (PDF 201 kb)
Rights and permissions
About this article
Cite this article
Schmidt, M., Schwarzwaelder, K., Bartholomae, C. et al. High-resolution insertion-site analysis by linear amplification–mediated PCR (LAM-PCR). Nat Methods 4, 1051–1057 (2007). https://doi.org/10.1038/nmeth1103
Issue Date:
DOI: https://doi.org/10.1038/nmeth1103
This article is cited by
-
RAISING is a high-performance method for identifying random transgene integration sites
Communications Biology (2022)
-
Common clonal origin of conventional T cells and induced regulatory T cells in breast cancer patients
Nature Communications (2021)
-
Clonal tracking using embedded viral barcoding and high-throughput sequencing
Nature Protocols (2020)
-
Successful engraftment of gene-corrected hematopoietic stem cells in non-conditioned patients with Fanconi anemia
Nature Medicine (2019)
-
Targeted in vivo knock-in of human alpha-1-antitrypsin cDNA using adenoviral delivery of CRISPR/Cas9
Gene Therapy (2018)