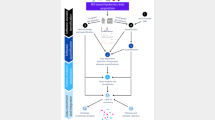
Abstract
We achieve automated and reliable annotation of lipid species and their molecular structures in high-throughput data from chromatography-coupled tandem mass spectrometry using decision rule sets embedded in Lipid Data Analyzer (LDA; http://genome.tugraz.at/lda2). Using various low- and high-resolution mass spectrometry instruments with several collision energies, we proved the method's platform independence. We propose that the software's reliability, flexibility, and ability to identify novel lipid molecular species may now render current state-of-the-art lipid libraries obsolete.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Wenk, M.R. Cell 143, 888–895 (2010).
Quehenberger, O. & Dennis, E.A. N. Engl. J. Med. 365, 1812–1823 (2011).
Dove, A. Science 347, 788–790 (2015).
Liebisch, G. et al. J. Lipid Res. 54, 1523–1530 (2013).
Song, H., Hsu, F.F., Ladenson, J. & Turk, J. J. Am. Soc. Mass Spectrom. 18, 1848–1858 (2007).
Taguchi, R. & Ishikawa, M. J. Chromatogr. A 1217, 4229–4239 (2010).
Kind, T. et al. Nat. Methods 10, 755–758 (2013).
Hsu, F.F. & Turk, J. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 877, 2673–2695 (2009).
Sud, M. et al. Nucleic Acids Res. 35, D527–D532 (2007).
Degtyarenko, K. et al. Nucleic Acids Res. 36, D344–D350 (2008).
Wishart, D.S. et al. Nucleic Acids Res. 37, D603–D610 (2009).
Jewison, T. et al. Nucleic Acids Res. 40, D815–D820 (2012).
Hartler, J. et al. Bioinformatics 27, 572–577 (2011).
Holčapek, M., Jandera, P., Zderadička, P. & Hrubá, L. J. Chromatogr. A 1010, 195–215 (2003).
Han, X., Yang, K. & Gross, R.W. Mass Spectrom. Rev. 31, 134–178 (2012).
Herzog, R. et al. PLoS One 7, e29851 (2012).
Yang, K., Cheng, H., Gross, R.W. & Han, X. Anal. Chem. 81, 4356–4368 (2009).
Husen, P. et al. PLoS One 8, e79736 (2013).
Cajka, T. & Fiehn, O. Trends Analyt. Chem. 61, 192–206 (2014).
Liebisch, G., Ejsing, C.S. & Ekroos, K. Clin. Chem. 61, 1542–1544 (2015).
Murphy, R.C. & Axelsen, P.H. Mass Spectrom. Rev. 30, 579–599 (2011).
Stein, S.E. & Scott, D.R. J. Am. Soc. Mass Spectrom. 5, 859–866 (1994).
Fauland, A. et al. J. Lipid Res. 52, 2314–2322 (2011).
Pedrioli, P.G. et al. Nat. Biotechnol. 22, 1459–1466 (2004).
Chambers, M.C. et al. Nat. Biotechnol. 30, 918–920 (2012).
Haug, K. et al. Nucleic Acids Res. 41, D781–D786 (2013).
Acknowledgements
Support by the Austrian Science Fund (FWF Project Grant P26148 to G.G.T.) and the Austrian Ministry for Science, Research and Economy (HSRSM Grant Omics Center Graz, BioTechMed-Graz to G.G.T.) is gratefully acknowledged. M.J.O.W. and Q.Z. were funded by the BBSRC (UK; Grant BBS/E/B/000C0415). We thank R. Salek for his extensive help in MetaboLights upload. Furthermore, we thank AB Sciex, Agilent Technologies, Bruker Daltonics, and Thermo Fisher Scientific for providing permission to distribute the WiffReader SDK, the MassHunter DAC, the CompassXtract, and the MSFileReader libraries in the software.
Author information
Authors and Affiliations
Contributions
J.H., A.T., M.T., F.S., G.H., H.C.K., and G.G.T. designed the study. J.H., A.T., M.T., G.N.R., and H.C.K. designed the experiments. K.A.Z. and G.H. provided the biological samples. A.T., M.T., G.N.R., F.T., A.C.-G., M.R.W., A.F., C.E.W., A.M.A., O.Q., Q.Z., and M.J.O.W. designed and performed the mass spectrometric experiments. J.H. and A.Z. implemented the algorithm and the software. J.H., A.T., O.A.Z., and H.C.K. developed the decision rule sets. J.H. and A.T. benchmarked the algorithm in comparison to LipidBlast and prepared the spectral evidence for the novel species. J.H. and H.C.K. prepared and uploaded the data in MetaboLights. J.H., A.T., F.S., and G.G.T. wrote the manuscript in cooperation with all contributing authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Notes 1–5 (PDF 7548 kb)
Supplementary Table 1
Overview of experiments performed and mass spectrometric platforms used. (XLSX 9 kb)
Supplementary Table 2
Lipid standards for control experiment 1. (XLSX 271 kb)
Supplementary Table 3
Isomeric pairs of lipid standards for different lipid subclasses applied in control experiment 2. (XLSX 29 kb)
Supplementary Table 4
Data evaluation of isomeric subclasses/adducts (control experiment 2). (XLSX 8 kb)
Supplementary Table 5
Pairs of structurally isomeric lipid standards for control experiment 3. (XLSX 48 kb)
Supplementary Table 6
Data evaluation of structurally isomeric pairs (described in Supplementary Table 5) measured in positive ion mode (control experiment 3). (XLSX 13 kb)
Supplementary Table 7
Data evaluation of structurally isomeric pairs (described in Supplementary Table 5) measured in negative ion mode (control experiment 3). (XLSX 12 kb)
Supplementary Table 8
Novel lipid molecular species identified in murine liver. (XLSX 14 kb)
Supplementary Table 9
Lipid species and lipid molecular species from murine liver detected by the seven MS/MS platforms and correctly identified by LDA. (XLSX 570 kb)
Supplementary Table 10
Overview on cross-platform species detection of seven MS/MS platforms based on murine liver samples. (XLSX 211 kb)
Supplementary Table 11
Sensitivity and positive predictive value (PPV) of LDA and LipidBlast (LB) in negative ion mode based on data acquired on Orbitrap Velos Pro in CID mode. (XLSX 9 kb)
Supplementary Table 12
Sensitivity and positive predictive value (PPV) of LDA and LipidBlast (LB) in positive ion mode based on data acquired on 4000 QTRAP. (XLSX 9 kb)
Supplementary Table 13
Sensitivity and positive predictive value (PPV) of LDA and LipidBlast (LB) in negative ion mode based on data acquired on 4000 QTRAP. (XLSX 9 kb)
Supplementary Table 14
Data evaluation of regio-isomeric pairs of chromatographically separated sn-1,2- and sn-1,3-diacylglycerols (DG). (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Hartler, J., Triebl, A., Ziegl, A. et al. Deciphering lipid structures based on platform-independent decision rules. Nat Methods 14, 1171–1174 (2017). https://doi.org/10.1038/nmeth.4470
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4470
This article is cited by
-
Ultraviolet exposure regulates skin metabolome based on the microbiome
Scientific Reports (2023)
-
Guiding the choice of informatics software and tools for lipidomics research applications
Nature Methods (2023)
-
Computational mass spectrometry accelerates C = C position-resolved untargeted lipidomics using oxygen attachment dissociation
Communications Chemistry (2022)
-
Utilizing Skyline to analyze lipidomics data containing liquid chromatography, ion mobility spectrometry and mass spectrometry dimensions
Nature Protocols (2022)
-
Phospholipid dynamics in ex vivo lung cancer and normal lung explants
Experimental & Molecular Medicine (2021)