Abstract
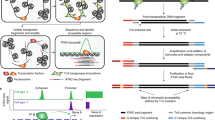
We present Omni-ATAC, an improved ATAC-seq protocol for chromatin accessibility profiling that works across multiple applications with substantial improvement of signal-to-background ratio and information content. The Omni-ATAC protocol generates chromatin accessibility profiles from archival frozen tissue samples and 50-μm sections, revealing the activities of disease-associated DNA elements in distinct human brain structures. The Omni-ATAC protocol enables the interrogation of personal regulomes in tissue context and translational studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Sequence Read Archive
Referenced accessions
Gene Expression Omnibus
References
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Nat. Methods 10, 1213–1218 (2013).
Jin, W. et al. Nature 528, 142–146 (2015).
Roadmap Epigenetics Consortium. et al. Nature 518, 317–330 (2015).
Koues, O.I. et al. Cell 165, 1134–1146 (2016).
Shih, H.Y. et al. Cell 165, 1120–1133 (2016).
Wu, J. et al. Nature 534, 652–657 (2016).
Gray, L.T. et al. eLife 6, e21883 (2017).
Scott, L.J. et al. Nat. Commun. 7, 11764 (2016).
Xu, J. et al. Nat. Genet. 49, 377–386 (2017).
Bao, X. et al. Genome Biol. 16, 284 (2015).
Corces, M.R. et al. Nat. Genet. 48, 1193–1203 (2016).
The ENCODE Project Consortium. Nature 489, 57–74 (2012).
Konopka, G. et al. Neuron 75, 601–617 (2012).
Henneman, W.J.P. et al. Neurology 72, 999–1007 (2009).
Filippini, N. et al. Neurology 75, 1645–1652 (2010).
Jones, A.R. et al. Neurobiol. Aging 34, 2234.e1–2234.e7 (2013).
Corces, M.R. et al. Protocol Exchange https://doi.org/10.1038/protex.2017.096 (2017).
Buenrostro, J.D., Wu, B., Chang, H.Y. & Greenleaf, W.J. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Mo, A. et al. Neuron 86, 1369–1384 (2015).
Piper, J. et al. Nucleic Acids Res. 41, e201 (2013).
Quinlan, A.R. & Hall, I.M. Bioinformatics 26, 841–842 (2010).
Leslie, R., O'Donnell, C.J. & Johnson, A.D. Bioinformatics 30, i185–i194 (2014).
Maurano, M.T. et al. Science 337, 1190–1195 (2012).
Love, M.I., Huber, W. & Anders, S. Genome Biol. 15, 550 (2014).
Schep, A.N., Wu, B., Buenrostro, J.D. & Greenleaf, W.J. bioRxiv 110346 (2017).
Picelli, S. et al. Genome Res. 24, 2033–2040 (2014).
Acknowledgements
We thank the Stanford Alzheimer's Disease Research Center (NIH P50 AG047366; to V. Henderson), the Pacific Udall Center for Excellence in Parkinson's Disease Research (NIH P50 NS062684; T.J.M.), and their participants for donating samples for research. We also thank J. Coller and X. Ji for sequencing assistance, E. Plowey and D. Channappa for tissue preparation, and P. Chu and A. Grewall for histology assistance. This work was supported by a grant from the Leukemia & Lymphoma Society Career Development Program (M.R.C.), US National Institutes of Health (NIH) training grant R25CA180993 (M.R.C.), NIH grants P50-HG007735 (H.Y.C. and W.J.G.), UM1HG00943 (W.J.G.), and U19AI057266 (W.J.G.), National Institute on aging grant RF1 AG053959 (T.J.M.), the Rita Allen Foundation (W.J.G.), the Human Frontier Science Program (W.J.G.), the National Science Foundation Graduate Research Fellowship Program (A.E.T.), and a US Department of Defense National Defense Science and Engineering Graduate (NDSEG) Fellowship (N.A.S.-A.).
Author information
Authors and Affiliations
Contributions
M.R.C. and H.Y.C. conceived the project; M.R.C., E.G.H., and A.T.S. performed all of the experiments, with help from M.R.M. and A.C.C.; S.W.C. produced the Tn5 transposase complex that was used in all of the experiments; P.G.G. performed all of the GWAS analysis, with guidance and supervision from A. Kundaje; M.R.C. and A.J.R. performed all other data analysis; M.R.C., A.E.T., S.V., and N.A.S.-A., developed methods for the isolation of nuclei from frozen tissues; B.W., A. Kathiria, V.I.R., and W.J.G. provided protocol expertise and recommendations; K.S.M. and T.J.M. oversaw all brain tissue acquisition and processing; M.K. and L.A.O. oversaw acquisition and processing of thyroid cancer tissue; A.J.R. and P.A.K. oversaw acquisition and processing of primary foreskin keratinocytes; and M.R.C., E.G.H., W.J.G., and H.Y.C. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
H.Y.C. and W.J.G. are co-founders and consultants of Epinomics. A. Kundaje is a member of the scientific advisory board of Epinomics.
Integrated supplementary information
Supplementary Figure 1 Comparison of technical replicate consistency between and across the standard ATAC-seq method, DNase-seq, and Omni-ATAC.
(a) Heatmap-based representation of ATAC-seq quality control metrics including library size (purple), percent of reads mapping to mitochondrial DNA (blue), and enrichment of signal at TSSs (orange) for each optimization made to produce the Omni-ATAC protocol. Deeper color is used to depict the most desirable value of each statistic. All values were determined from 100,000 random aligned reads. Data represents the average of two technical replicates per square. (b-d) Correlation of technical replicates from GM12878 ATAC-seq data generated using (b) the Omni-ATAC protocol, (c) the standard ATAC-seq protocol, and (d) DNase-seq. Data for standard ATAC-seq and DNase-seq were obtained from publicly available data. Each dot represents an individual peak. Peaks analyzed were derived from a union peak set using data from all three methods. Only 50,000 random peaks with more than 5 reads are shown. R value indicated at the top of each plot. (e,f) Correlation of GM12878 ATAC-seq data generated using (e) Omni-ATAC protocol and (f) the standard ATAC-seq protocol with DNase-seq as in (b-d). (g) Pearson correlation heatmap showing sample by sample unsupervised clustering on all peaks identified across all technical replicates from each method. Each sample represents an individual technical replicate. (h,i) CTCF meta-footprint across the top 20,000 ATAC-seq peaks containing a CTCF motif in data from GM12878 cells derived using (h) the standard ATAC-seq protocol and (i) the Omni-ATAC protocol. Data for standard ATAC-seq method derived from published data.
Supplementary Figure 2 Comparison of Omni-ATAC to the Fast-ATAC method in primary human T cells.
(a-c) Correlation of technical replicates from primary human CD4+ T cells cells using (a) the Omni-ATAC protocol, (b) the Fast-ATAC protocol, and (c) the standard ATAC-seq protocol. All data shown was generated in this study from the same sample of T cells. Each dot represents an individual peak. Peaks analyzed were derived from a union peak set using data from all three methods. Only 50,000 random peaks with more than 5 reads are shown. R value indicated at the top of each plot. (d,e) Correlation of ATAC-seq data generated using the Omni-ATAC protocol with data generated using (d) the Fast-ATAC protocol or (e) the standard ATAC-seq protocol in CD4+ T cells as in (a-c). (f) Pearson correlation heatmap showing sample by sample unsupervised clustering on all peaks identified across all technical replicates of CD4+ T cells from each method.
Supplementary Figure 3 Comparison of Omni-ATAC to standard ATAC-seq and DNase-seq at the level of peaks.
(a) The number of PCR cycles required to amplify ATAC-seq libraries to an absolute yield of 80 femtomoles (20 ul at 4 nM) using the standard ATAC-seq protocol, the Fast-ATAC protocol, and the Omni-ATAC protocol. N=28 for each box chart, representing 2 technical replicates from 14 distinct cell types and cell contexts (Supplementary Table 1). (b) Fraction of reads in peaks mapping to TSSs (+/- 500 bp of TSS) and distal elements (>500 bp from TSS) from GM12878 libraries generated in this study and from public repositories using the standard ATAC-seq protocol (“Standard”), the Fast-ATAC-seq protocol (“Fast”), the Omni-ATAC-seq protocol (“Omni-ATAC”), and DNase-seq (“DNase”). Each bar represents the mean of 2 technical replicates with the exception of the previously published ATAC-seq data which had 4 technical replicates. Error bars represent standard deviation. **p < 0.01, ***p < 0.001 by two-tailed unpaired students t-test comparing the fraction of reads in peaks to reads outside of peaks. All values were determined from 5 million random aligned de-duplicated reads. (c) Histogram comparing the distribution of macs2 peak scores (-log10(pvalue of peak call)) from peaks called from 20 million randomly selected aligned de-duplicated reads in GM12878 cells as in (d). (d,e) In silico simulation of peak calling from variable read depth using data derived from (d) GM12878 cells with the standard ATAC-seq protocol, DNase-seq, and the Omni-ATAC protocol or (e) CD4+ T cells with the standard ATAC-seq protocol, the Fast-ATAC protocol, and the Omni-ATAC protocol. Input reads were randomly selected from a pool of properly aligned, non-duplicate reads. All input reads were trimmed to equal length (36 bp) prior to alignment to match DNase-seq data. GM12878 data from the standard ATAC-seq protocol and DNase-seq were obtained from publicly available sources. All data for CD4+ T cells was generated from a single donor. Data points represent the mean of three individual down-sampled technical replicates. (f-h) Signal at peaks called as “unique” in the analysis presented in Figure 1c for (f) the standard ATAC-seq protocol, (g) the Omni-ATAC protocol, and (h) DNase-seq. Peak score is calculated in the same way as the TSS enrichment score but for a different set of specified peaks. (i-l) Input read normalized sequencing tracks at a control locus (i,j) and a B cell-related locus (k,l) from GM12878 cells derived using standard ATAC-seq, DNase-seq, and Omni-ATAC. In each case, 60 million properly aligned, non-duplicate reads were used to allow for direct comparison. All tracks in each panel are shown on the same y-axis scale. Tracks shown in (j,l) depict the same regions shown in (i,k) but with a reduced y-axis scale to show the difference in signal-to-background in data generated using the Omni-ATAC protocol compared to the standard ATAC-seq protocol and DNase-seq. Peaks that pass above the 10 read y-axis limit are truncated at y=10. All input reads were trimmed to equal length (36 bp) prior to alignment to match DNase-seq. Regions shown represent (i,j) chr19:36,097,939-36,272,939, and (k,l) chr9:36,373,966-37,494,500.
Supplementary Figure 4 Omni-ATAC outperforms the standard ATAC-seq method in situations requiring low cell numbers.
(a,b) Sequencing tracks of ATAC-seq data at (a) a control locus and (b) the PAX5 locus derived from the standard ATAC-seq protocol and the Omni-ATAC protocol using 500 GM12878 cells. No normalization was performed. All available filtered reads are shown under the assumption that the libraries have been sequenced to relatively complete depth. Each track represents an individual technical replicate. Data for standard ATAC-seq from Buenrostro et al. 20131. Region shown represents (a) chr19:36,097,939-36,272,939 or (b) chr9:36,373,966-37,494,500. (c) The total raw reads, filtered reads, and the percent duplication (how completely the libraries have been sequenced) are shown for each library to the right. (d,e) Sequencing tracks shown in (a,b) but with a y-axis scale of 5 to show the difference in signal-to-background in data derived using the Omni-ATAC protocol compared to the standard ATAC-seq protocol. Peaks that pass above the 5 read y-axis limit are truncated at y=5. Each track represents an individual technical replicate. Region shown represents (d) chr19:36,097,939-36,272,939 or (e) chr9:36,373,966-37,494,500. (f) Fraction of reads in peaks mapping to TSSs (+/- 500 bp of TSS) and distal elements (>500 bp from TSS) from the libraries shown in (a-e). Each bar represents the mean of 3 technical replicates. Error bars represent standard deviation. ***p<0.001 by two-tailed unpaired students t-test. Data for standard ATAC-seq from Buenrostro et al. 20131. (g) Metaplot showing enrichment of reads at TSSs in data derived from 500 GM12878 cells using Omni-ATAC (N=3) compared to published data using standard ATAC-seq1 (N=3). (h) Overlap of 500 cell GM12878 peaks called from all available reads using the standard ATAC-seq protocol and the Omni-ATAC protocol. Numbers represent the mean of three individual technical replicates in each case. Overlap was performed with a union set of GM12878 peaks identified by DNase-seq, standard ATAC-seq, and Omni-ATAC. Data for standard ATAC-seq from Buenrostro et al. 20131. (i) Correlation of Omni-ATAC data from 500 GM12878 cells with Omni-ATAC data from 50,000 GM12878 cells. Each dot represents an individual peak. Peaks analyzed were derived from a union peak set using data from the standard ATAC-seq method, DNase-seq, and Omni-ATAC. Only 50,000 random peaks with more than 5 reads are shown. R value indicated at the top of the plot. (j) Pearson correlation heatmap showing sample by sample unsupervised clustering on all peaks identified across all technical replicates of 500 and 50,000 GM12878 cells. Data for standard ATAC-seq for both 500 cell and 50,000 cell GM12878 data from Buenrostro et al. 20131. DNase-seq was performed on an unknown number of cells.
Supplementary Figure 5 Omni-ATAC enables the use of less amounts of transposase without decreasing data quality.
(a) Heatmap-based representation of ATAC-seq quality control metrics including library size, percent of reads mapping to mitochondrial DNA, and enrichment of signal at TSSs. Deeper color is used to depict the most desirable value of each statistic. All values were determined from 5 million random aligned reads. Data represents two technical replicates per square. (b-d) Correlation of Omni-ATAC data generated with varying amounts of transposase enzyme. Technical replicates using (b) 100% Tn5 transposase input and (c) 20% transposase input are shown alongside (d) the correlation of 100% Tn5 transposase input with 20% Tn5 transposase input. Peaks analyzed were derived from DNase-seq data. Only 50,000 random peaks with more than 5 reads are shown. R value indicated at the top of each plot. (e) Fraction of reads in peaks mapping to TSSs (+/- 500 bp of TSS) and distal elements (>500 bp from TSS) for Omni-ATAC data performed with various amounts of Tn5 transposase input. Peak set used was derived from DNase-seq data. Each bar represents the mean of 2 technical replicates. Error bars represent standard deviation. NS = not significant.
Supplementary Figure 6 Application of Omni-ATAC to frozen post-mortem human brain samples yields robust data.
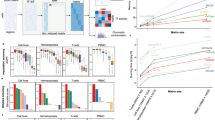
(a) Representative phase contrast image of nuclei isolated from frozen post-mortem human brain. Scale bar represents 10 μm. Experiment was repeated 5 times with 5 total images collected. (b) Representative DAPI staining of the nuclei shown in (a). Scale bar represents 10 μm. (c) Three-dimensional principal component analysis showing the first 3 principal components from ATAC-seq data generated with the Omni-ATAC protocol in 5 different brain regions from 2 different biological donors, each with 2 technical replicates. Different brain regions are indicated by color. CC = corpus callosum; CN = caudate nucleus; CB = cerebellum; HIP = hippocampus; MFG = middle frontal gyrus. (d) Differentially-accessible peaks defined as being at least 2 standard deviations away from the mean of all other regions. Transcription factors whose binding motifs are identified as enriched using the Hypergeometric Optimization of Motif Enrichment (Homer) algorithm in the accessible chromatin regions of each brain region are shown to the right. (e) Transcription factor deviation of all technical replicates and biological donors. TFs enriched in each cluster are shown to the right. Color scale represents the minimum and maximum of each row. Each row represents an individual TF motif. Each column represents an individual replicate of ATAC-seq using the Omni-ATAC protocol.
Supplementary Figure 7 GWAS analysis of ATAC-seq data from different brain regions provides key insights into the ontogeny of various brain diseases.
(a) Box plot of the minimum empirical p value from all brain regions for each GWAS SNP set assayed. *p < 0.05 by two-tailed unpaired students t-test. (b) Significance of enrichment of disease-specific GWAS polymorphisms in the uniquely-accessible regions of the 5 different brain regions shown in Fig. 2b. The empirical p value is depicted colorimetrically with reference to association-based permutations of the GWAS SNPs. (c) Heatmap representation of the enrichment (left) and significance (right) of the enrichment of GWAS SNPs in the uniquely-accessible regions of each brain region. For enrichment, 0 values are represented colorimetrically the same as a log2(enrichment) of -3. Regions without significant associations are shown as white in the p value heatmap.
Supplementary Figure 8 The Omni-ATAC protocol enables generation of robust ATAC-seq data from 50-μm-thick frozen tissue sections.
(a) Depth-normalized sequencing tracks surrounding amyotrophic lateral sclerosis disease-associated SNP rs7175096. Tracks were normalized according to the total number of reads in peaks. Each individual replicate (N=4 technical replicates for each of 2 donors per tissue fragment, N=1 technical replicate for each of 2 donors for corpus callosum sections, and N=2 technical replicates for each of 2 donors for cerebellum sections) is displayed with 25% transparency to show consistency among technical replicates and biological donors. Region shown represents chr15:92,390,775-92,470,760. Data derived from 20 mg tissue fragments and 50 μm tissue sections are represented by a cartoon image to the right. (b-d) Depth normalized sequencing tracks from all replicates from all regions comparing ATAC-seq data generated using the Omni-ATAC protocol in 50 μm tissue sections and larger 20 mg tissue fragments. Each individual replicate (N=4 technical replicates for each of 2 donors per tissue fragment, N=1 technical replicate for each of 2 donors for corpus callosum sections, and N=2 technical replicates for each of 2 donors for cerebellum sections) is displayed with 25% transparency to show consistency among technical replicates and biological donors. Regions shown represent (c) chr2:182,378,211-182,585,768, (d) chr21:34,391,306-34,455,298, and (e) chr22:38,347,790-38,398,955. (e-f) Correlation of ATAC-seq data from 50 μm sections of (f) corpus callosum or (g) cerebellum with ATAC-seq data generated from a nearby 20 mg fragment of the same brain region from the same donor. Each dot represents an individual peak from a union set of peaks called from all technical replicates and biological donors of data derived from tissue fragments from the same brain region. Only 50,000 random peaks with more than 5 reads are shown. (g) Histology and immunohistochemistry of 5 μm frozen sections from cerebellum and corpus callosum immediately adjacent to the 50 μm section used for ATAC-seq shown in Figure 2d-e. Stains from left to right: hematoxylin and eosin (H&E), Nissl. Scale bar represents 100 μm in each image.
Supplementary Figure 9 Histology of tissue immediately adjacent to a 50-μm section used for ATAC-seq.
(a,e) Hematoxylin and eosin staining of corpus callosum and cerebellum. Scale bar represents 1 mm. (b,f) Nissl staining of corpus callosum and cerebellum. Scale bar represents 1 mm. (c,g) Anti-NEUN immunohistochemistry of corpus callosum and cerebellum. Scale bar represents 1 mm. (d,h) Anti-SOX10 immunohistochemistry of corpus callosum and cerebellum. Scale bar represents 100 μm. All images represent same tissue shown in Figure 2d.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Note 1.
Life Sciences Reporting Summary
Life Sciences Reporting Summary.
Supplementary Protocol 1
Omni-ATAC: Improved ATAC-seq protocol.
Supplementary Protocol 2
Isolation of nuclei from frozen tissues.
Supplementary Table 1
Sequencing statistics for all libraries generated in this work.
Supplementary Table 2
Clinical characteristics of brain donors.
Supplementary Table 3
Empirical p values and enrichment values for all GWAS data.
Rights and permissions
About this article
Cite this article
Corces, M., Trevino, A., Hamilton, E. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat Methods 14, 959–962 (2017). https://doi.org/10.1038/nmeth.4396
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4396
This article is cited by
-
txci-ATAC-seq: a massive-scale single-cell technique to profile chromatin accessibility
Genome Biology (2024)
-
Chromatin accessibility and cell cycle progression are controlled by the HDAC-associated Sin3B protein in murine hematopoietic stem cells
Epigenetics & Chromatin (2024)
-
The glucocorticoid receptor as a master regulator of the Müller cell response to diabetic conditions in mice
Journal of Neuroinflammation (2024)
-
The dynamic landscape of chromatin accessibility and active regulatory elements in the mediobasal hypothalamus influences the seasonal activation of the reproductive axis in the male quail under long light exposure
BMC Genomics (2024)
-
Chromatin accessibility and H3K9me3 landscapes reveal long-term epigenetic effects of fetal-neonatal iron deficiency in rat hippocampus
BMC Genomics (2024)